Search Count: 59
All
Selected
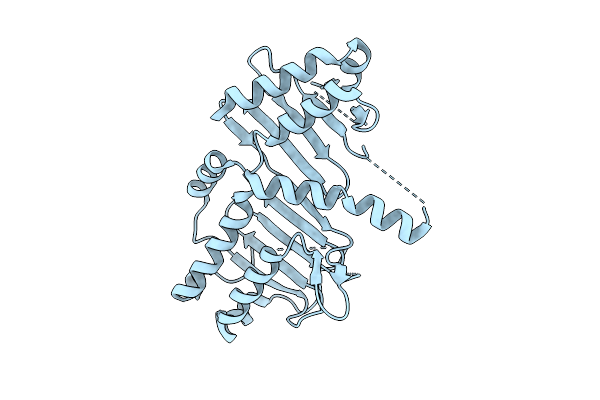 |
Organism: Streptomyces novoguineensis
Method: X-RAY DIFFRACTION Resolution:3.50 Å Release Date: 2026-04-08 Classification: LYASE Ligands: HOH |
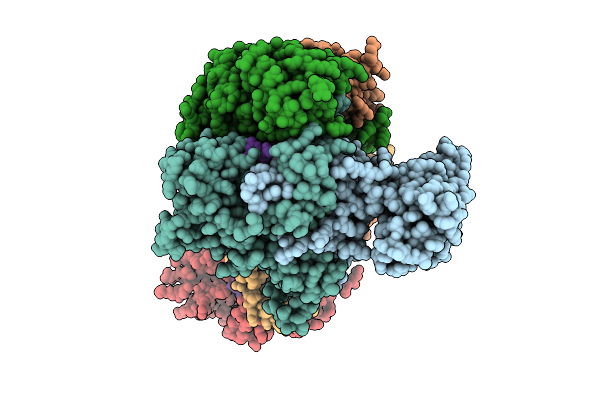 |
Organism: Pseudomonas aeruginosa ucbpp-pa14
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: RNA BINDING PROTEIN/RNA |
 |
Organism: Pseudomonas aeruginosa ucbpp-pa14
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: RNA BINDING PROTEIN/RNA |
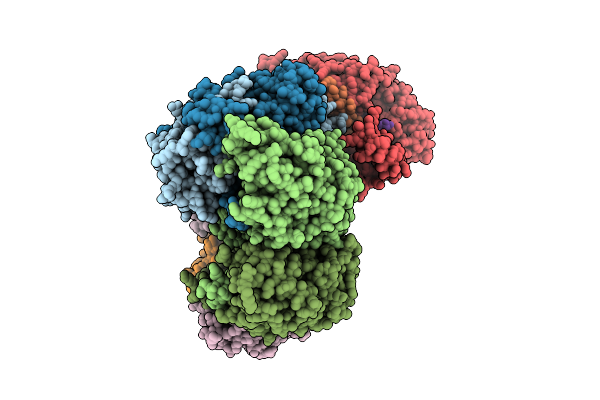 |
Organism: Pseudomonas aeruginosa ucbpp-pa14
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: RNA BINDING PROTEIN/RNA |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-10-15 Classification: MEMBRANE PROTEIN Ligands: PC1 |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-06-11 Classification: TRANSLOCASE Ligands: FUD, PC1 |
 |
Organism: Dragon grouper nervous necrosis virus
Method: SOLUTION NMR Release Date: 2024-10-30 Classification: VIRAL PROTEIN |
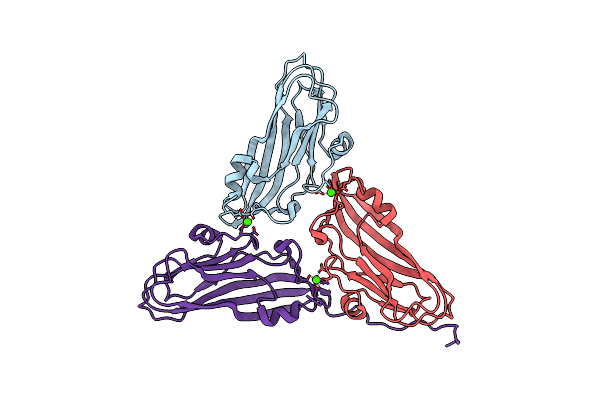 |
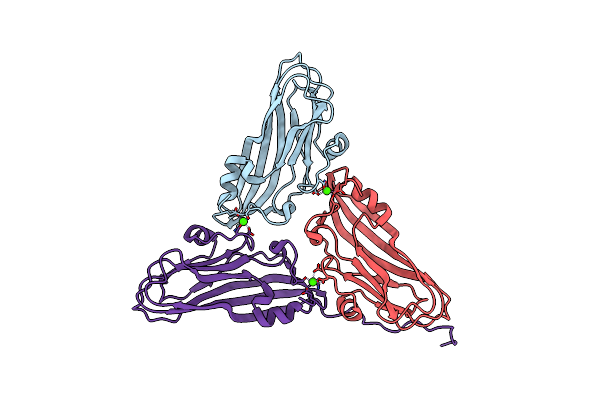
Cryo-Em Structure Of Dragon Grouper Nervous Necrosis Virus-Like Particle At Ph8.0 (3.23A)
Organism: Dragon grouper nervous necrosis virus
Method: ELECTRON MICROSCOPY Release Date: 2024-08-14 Classification: VIRUS Ligands: CA |
 |
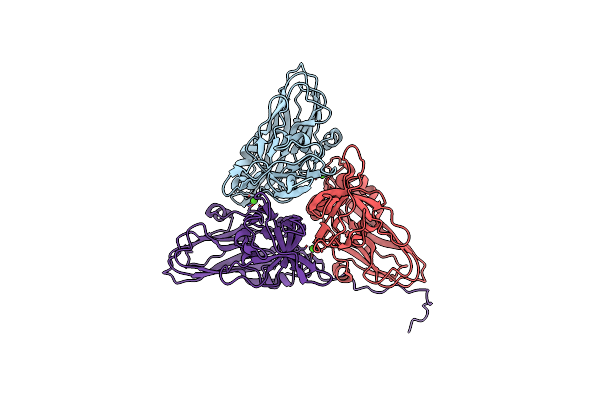
Cryo-Em Structure Of Dragon Grouper Nervous Necrosis Virus-Like Particle At Ph6.5 (2.82A)
Organism: Dragon grouper nervous necrosis virus
Method: ELECTRON MICROSCOPY Release Date: 2024-08-14 Classification: VIRUS Ligands: CA |
 |
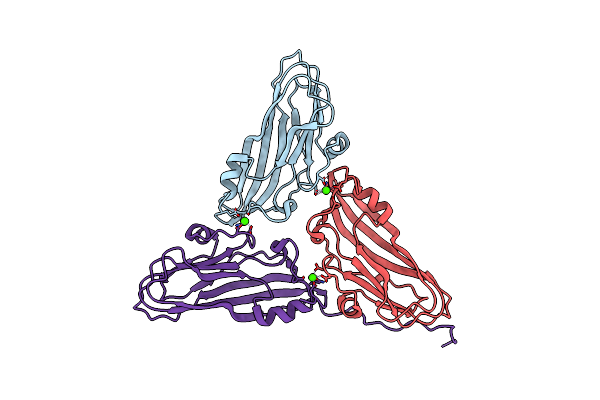
Cryo-Em Structure Of Dragon Grouper Nervous Necrosis Virus-Like Particle At Ph5.0 (3.52A)
Organism: Dragon grouper nervous necrosis virus
Method: ELECTRON MICROSCOPY Release Date: 2024-08-14 Classification: VIRUS Ligands: CA |
 |
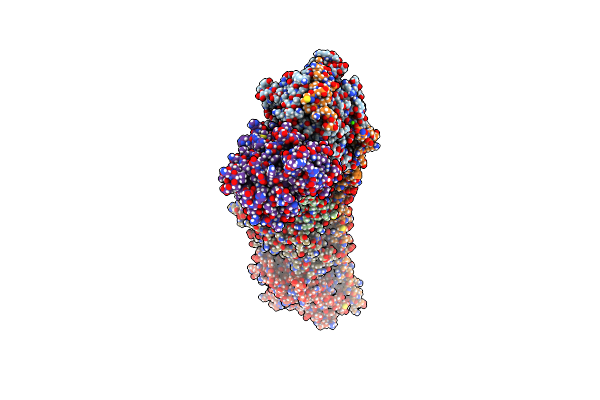
Cryo-Em Structure Of Dragon Grouper Nervous Necrosis Virion At Ph6.5 (3.12A)
Organism: Dragon grouper nervous necrosis virus
Method: ELECTRON MICROSCOPY Release Date: 2024-08-14 Classification: VIRUS Ligands: CA |
 |
Organism: Rattus norvegicus, Gallus gallus, Bos taurus
Method: X-RAY DIFFRACTION Resolution:2.78 Å Release Date: 2024-03-27 Classification: CELL CYCLE Ligands: GTP, MG, CA, GDP, MES, A1H01, IMD, ACP |
 |
Organism: Rattus norvegicus, Gallus gallus, Bos taurus
Method: X-RAY DIFFRACTION Resolution:2.57 Å Release Date: 2024-03-27 Classification: CELL CYCLE Ligands: GTP, MG, CA, IMD, GDP, MES, A1H00, GOL, ACP |
 |
Organism: Homo sapiens, Mus musculus, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2023-10-11 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens, Mus musculus, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2023-10-11 Classification: MEMBRANE PROTEIN Ligands: 140 |
 |
Organism: Homo sapiens, Mus musculus, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2023-10-11 Classification: MEMBRANE PROTEIN Ligands: 9HO |
 |
Organism: Escherichia coli, Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2023-10-11 Classification: MEMBRANE PROTEIN Ligands: NFI |
 |
Structural Basis Of Transcriptional Activation By The Ompr/Phob-Family Response Regulator Pmra
Organism: Escherichia coli bl21(de3), Klebsiella pneumoniae jm45, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2023-08-30 Classification: TRANSCRIPTION |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2022-05-11 Classification: HYDROLASE |
 |
Organism: Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2022-04-06 Classification: BIOSYNTHETIC PROTEIN Ligands: NI |

