Search Count: 24
All
Selected
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.50 Å Release Date: 2026-03-18 Classification: CELL CYCLE Ligands: NK3 |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.40 Å Release Date: 2026-03-18 Classification: CELL CYCLE Ligands: NK3 |
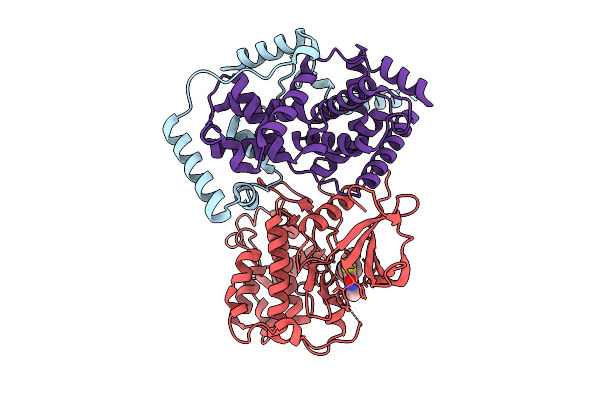 |
Cryo-Em Structure Of The Cdk11B-Cyclin L2-Sap30Bp Bound To Ots964 (Conformation 1)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.40 Å Release Date: 2026-03-18 Classification: SPLICING Ligands: NK3 |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.30 Å Release Date: 2026-03-18 Classification: SPLICING Ligands: ANP, MG |
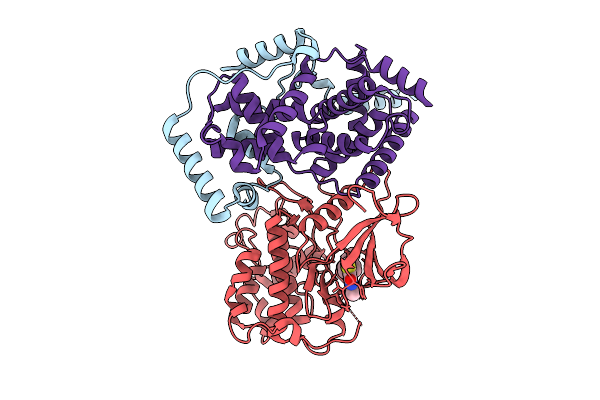 |
Cryo-Em Structure Of The Cdk11B-Cyclin L2-Sap30Bp Bound To Ots964 (Conformation 2)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.40 Å Release Date: 2026-03-18 Classification: SPLICING Ligands: NK3 |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-07-24 Classification: CELL CYCLE |
 |
Organism: Homo sapiens, Xenopus laevis
Method: ELECTRON MICROSCOPY Release Date: 2024-07-24 Classification: CELL CYCLE |
 |
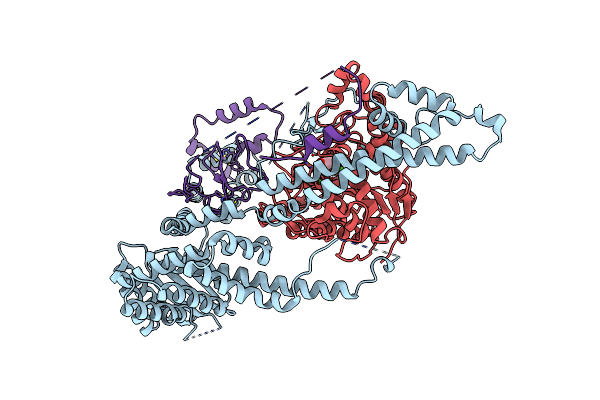
Cryo-Em Structure Of The Human Sin3B Histone Deacetylase Complex At 3.7 Angstrom
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-05-10 Classification: GENE REGULATION Ligands: ZN, CA |
 |
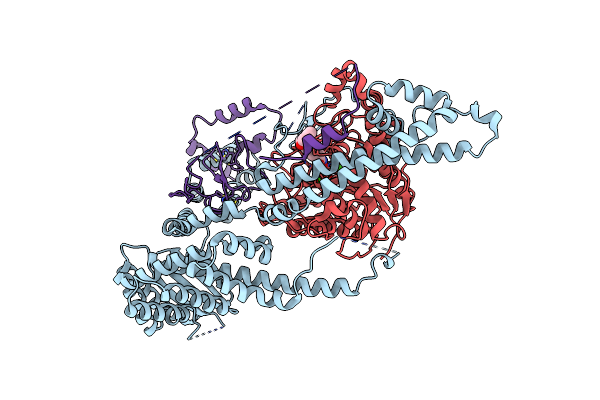
Cryo-Em Structure Of The Human Sin3B Histone Deacetylase Core Complex At 2.8 Angstrom
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-05-10 Classification: GENE REGULATION Ligands: ZN, CA, ACT |
 |
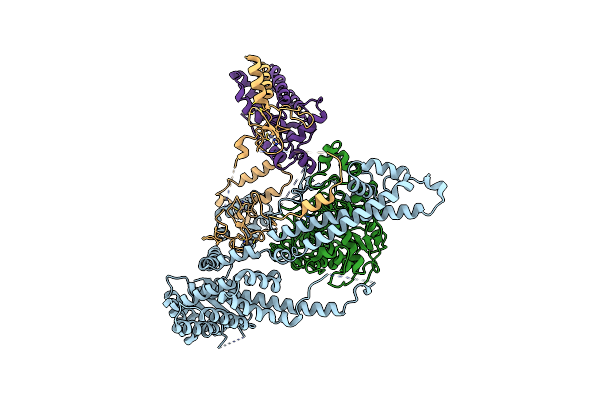
Cryo-Em Structure Of The Human Sin3B Histone Deacetylase Core Complex With Saha At 2.8 Angstrom
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-05-10 Classification: GENE REGULATION Ligands: SHH, ZN, CA |
 |
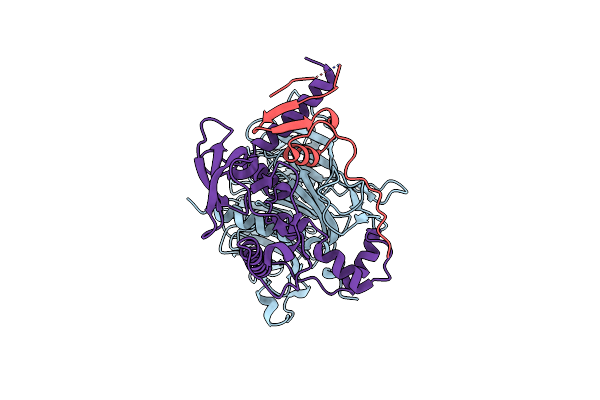
Cryo-Em Structure Of The Human Sin3B Full-Length Complex At 3.4 Angstrom Resolution
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-05-10 Classification: HYDROLASE Ligands: ZN, CA |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2022-08-10 Classification: CELL CYCLE |
 |
Organism: Homo sapiens, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2021-01-13 Classification: CELL CYCLE Ligands: ZN |
 |
Cryo-Em Structure Of The Anaphase-Promoting Complex/Cyclosome, In Complex With The Mitotic Checkpoint Complex (Apc/C-Mcc) At 3.8 Angstrom Resolution
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2020-02-19 Classification: CELL CYCLE |
 |
Cryo-Em Structure Of The Anaphase-Promoting Complex/Cyclosome, In Complex With The Nek2A Substrate At 3.9 Angstrom Resolution
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.90 Å Release Date: 2020-02-19 Classification: CELL CYCLE Ligands: ZN |
 |
Cryo-Em Structure Of Trip13 In Complex With Atp Gamma S, P31Comet, C-Mad2 And Cdc20
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2018-05-02 Classification: CELL CYCLE Ligands: AGS |
 |
Cryo-Em Structure Of The Anaphase-Promoting Complex/Cyclosome, In Complex With The Mitotic Checkpoint Complex (Apc/C-Mcc) At 4.2 Angstrom Resolution
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:4.20 Å Release Date: 2016-08-10 Classification: CELL CYCLE |
 |
Organism: Homo sapiens, Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Resolution:3.90 Å Release Date: 2016-05-25 Classification: CELL CYCLE Ligands: ZN |
 |
Organism: Homo sapiens, Bos taurus, Unidentified
Method: ELECTRON MICROSCOPY Resolution:3.40 Å Release Date: 2016-05-25 Classification: CELL CYCLE Ligands: ZN |
 |
Organism: Drosophila melanogaster
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2013-11-13 Classification: TRANSCRIPTION |

