Search Count: 1,742
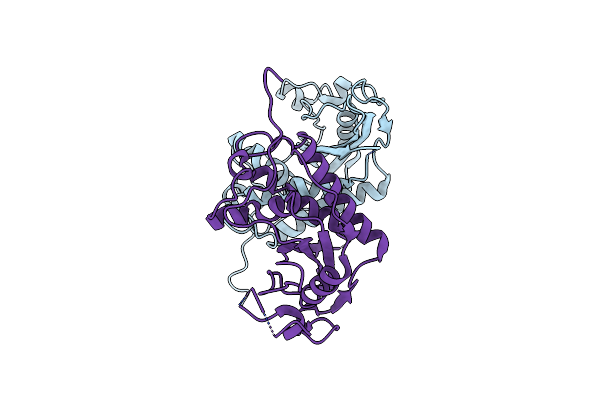 |
Organism: Solenopsis invicta
Method: X-RAY DIFFRACTION Resolution:3.05 Å Release Date: 2008-09-09 Classification: ALLERGEN |
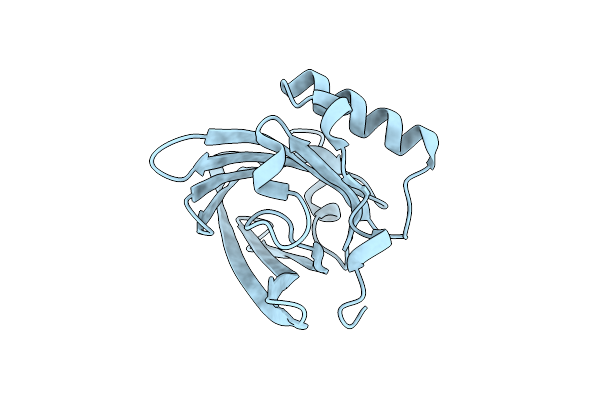 |
Organism: Bos taurus
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2015-09-16 Classification: ALLERGEN |
 |
Organism: Bos taurus
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2015-09-16 Classification: ALLERGEN |
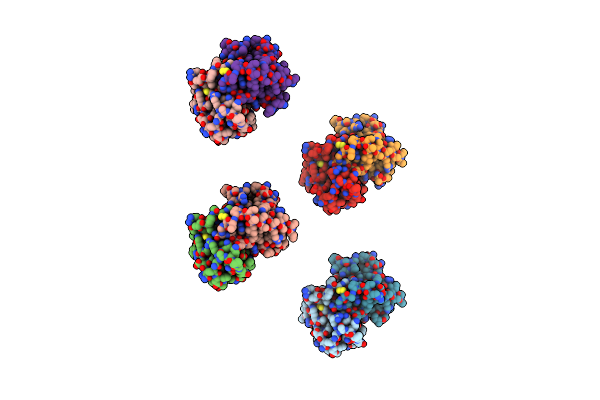 |
Organism: Solenopsis invicta
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2011-11-23 Classification: ALLERGEN Ligands: HP6 |
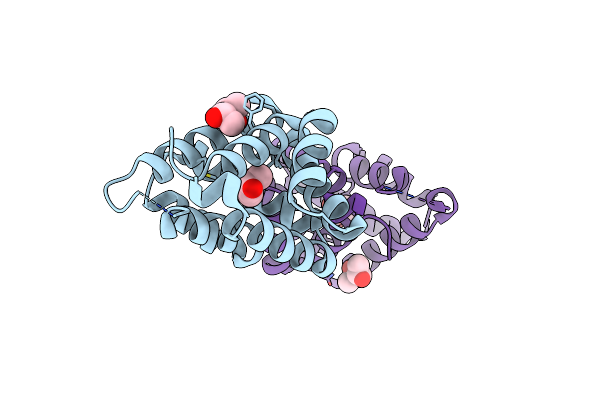 |
Organism: Felis catus
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2003-10-14 Classification: ALLERGEN Ligands: MPD |
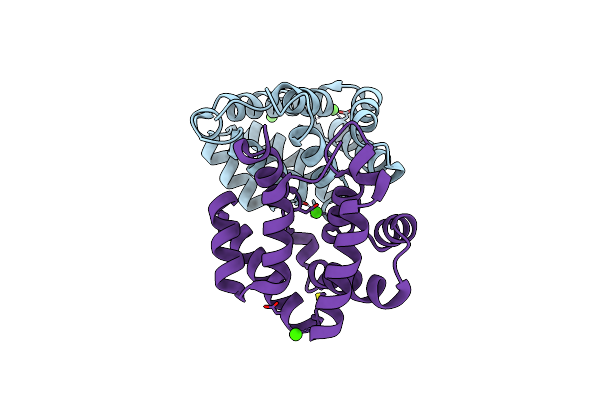 |
Structural Characterization Of The Tetrameric Form Of The Major Cat Allergen Fel D 1
Organism: Felis catus
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2007-04-17 Classification: ALLERGEN Ligands: CA |
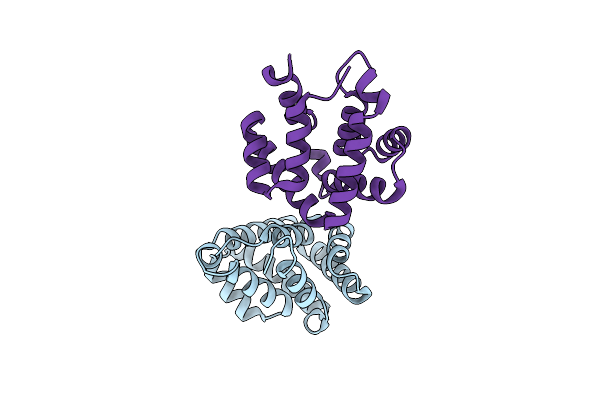 |
Organism: Felis catus
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2006-05-16 Classification: ALLERGEN |
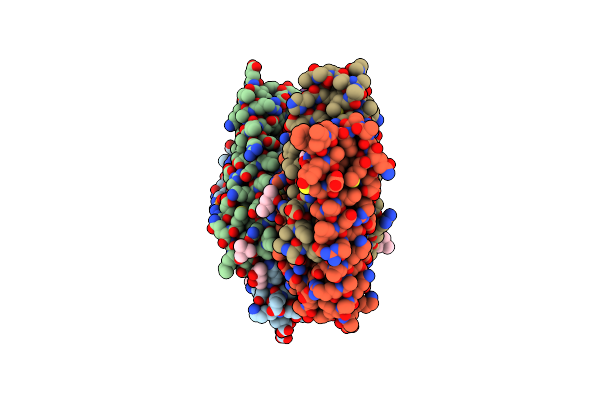 |
Organism: Dermatophagoides pteronyssinus
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2010-06-02 Classification: ALLERGEN Ligands: MPD, MRD, PO4 |
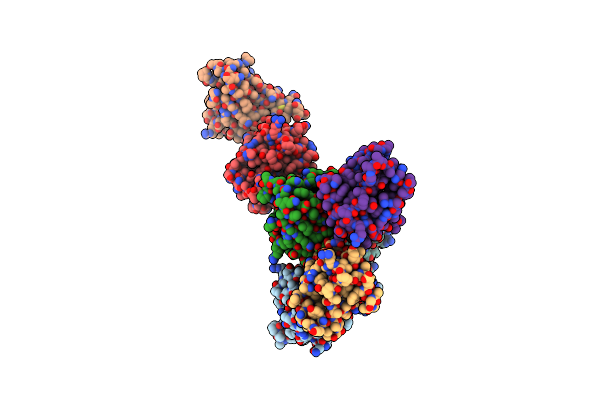 |
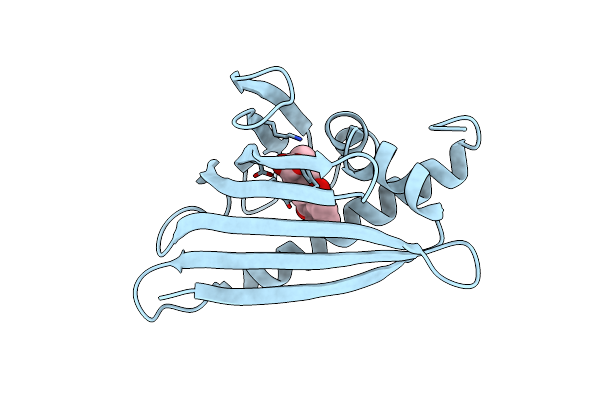
Organism: Canis lupus familiaris
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2021-11-10 Classification: ALLERGEN |
 |
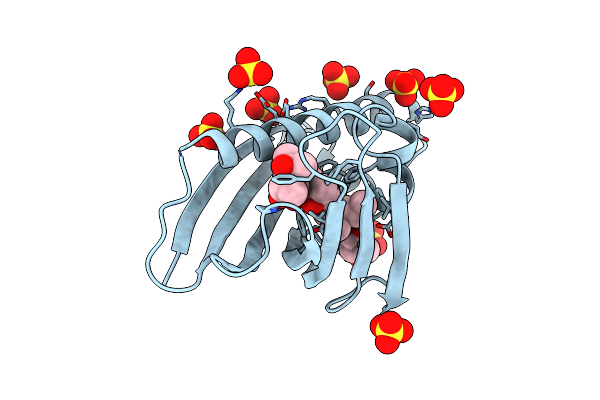
Crystal Structure Of The Major Pollen Allergen Bet V 1-A In Complex With P303
Organism: Betula pendula
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2015-03-11 Classification: ALLERGEN Ligands: 2AX |
 |
Organism: Betula pendula
Method: X-RAY DIFFRACTION Resolution:1.16 Å Release Date: 2012-05-30 Classification: ALLERGEN Ligands: MPD, SO4, TRS, MRD |
 |
 |
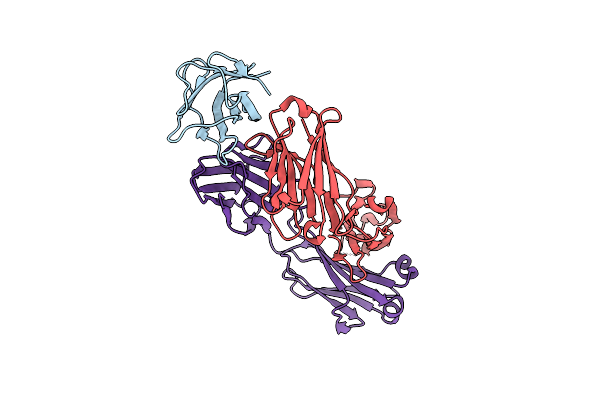 |
Crystal Structure Of The Major Grass Pollen Allergen Phl P 2 In Complex With Its Specific Ige-Fab
Organism: Phleum pratense, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2009-02-17 Classification: ALLERGEN |
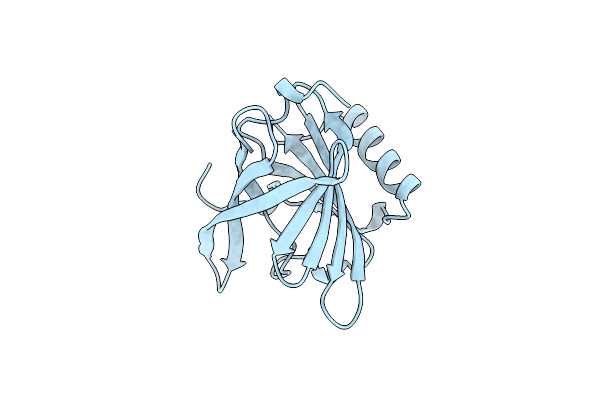 |
Organism: Bos taurus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 1999-05-11 Classification: ALLERGEN |
 |
 |
Organism: Arachis hypogaea
Method: X-RAY DIFFRACTION Resolution:2.43 Å Release Date: 2012-02-15 Classification: ALLERGEN |
 |
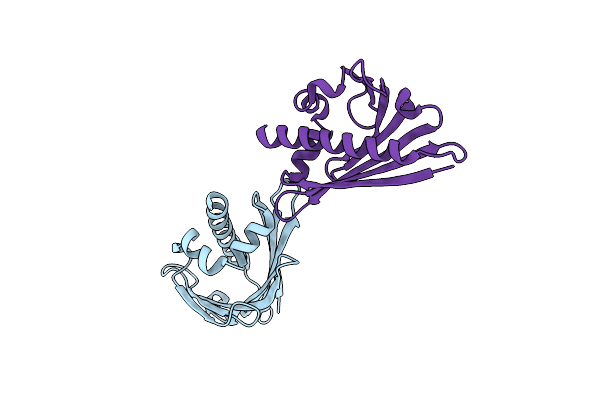
Crystal Structure Of Birch Pollen Allergen Bet V 1 Mutant G26L, D69I, P90L, K97I
Organism: Betula pendula
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2015-11-25 Classification: ALLERGEN Ligands: SO4 |
 |
Organism: Apium graveolens
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2005-06-13 Classification: ALLERGEN |
 |
1.6 A Crystal Structure Of The Lipocalin Dog Allergen Can F 1 With The C118S Mutation
Organism: Canis lupus familiaris
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2023-04-05 Classification: ALLERGEN |

