Search Count: 3,278
All
Selected
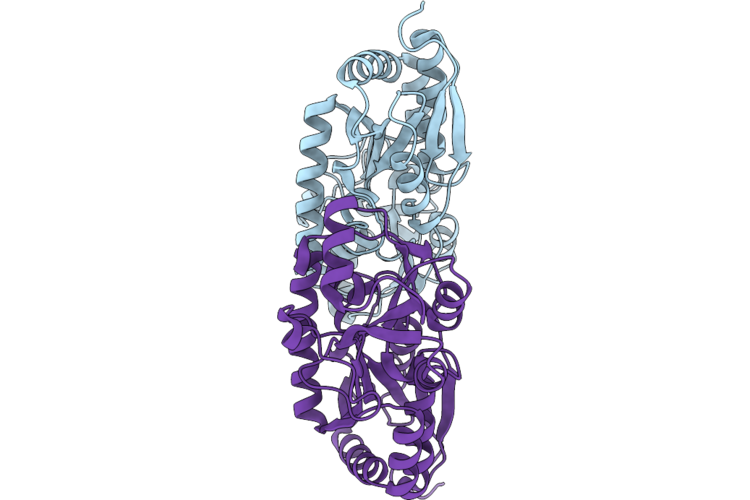 |
Crystal Structure Of A Tctc Solute Binding Protein From Vibrio Sp. C42, No Ligand
Organism: Vibrio atlanticus
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING |
 |
Aminoacyl-Trna-Dependent Peptide Synthase, Sba18, Complexed With Streptothrisamine
Organism: Streptomyces sp.
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-01 Classification: TRANSFERASE Ligands: A1L3W, TRS |
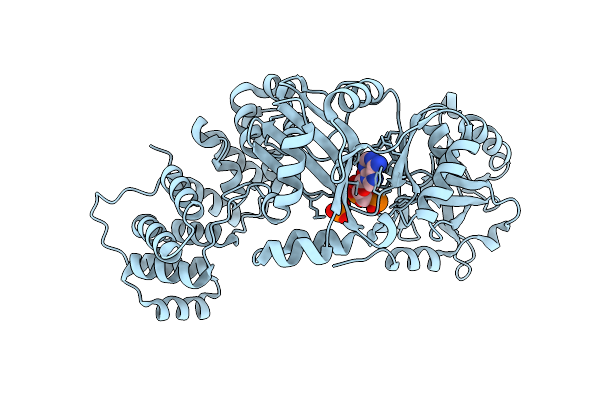 |
Organism: Thermococcus gammatolerans
Method: X-RAY DIFFRACTION Resolution:2.17 Å Release Date: 2026-03-18 Classification: DNA BINDING PROTEIN,LIGASE Ligands: PO4, AMP |
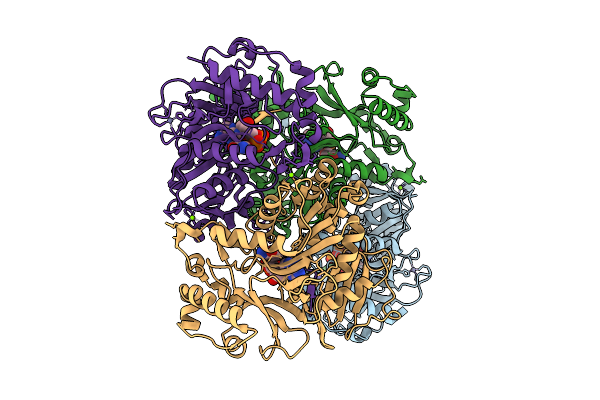 |
Clxa From Clostridium Cavendishii In Complex With 3-Amino,4-Hydroxybenzoic Acid And Its Adenylate
Organism: Clostridium cavendishii
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-03-18 Classification: LIGASE Ligands: MG, ZN, AMP, A1JLN, A1JOG |
 |
Pr1 Phage Heterodimeric Dna Ligase In Complex With 21-Mer Nicked Dna (Random Sequence)
Organism: Providencia phage vb_pres_pr1
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2026-02-18 Classification: DNA BINDING PROTEIN Ligands: NMN, AMP, VO4, ZN |
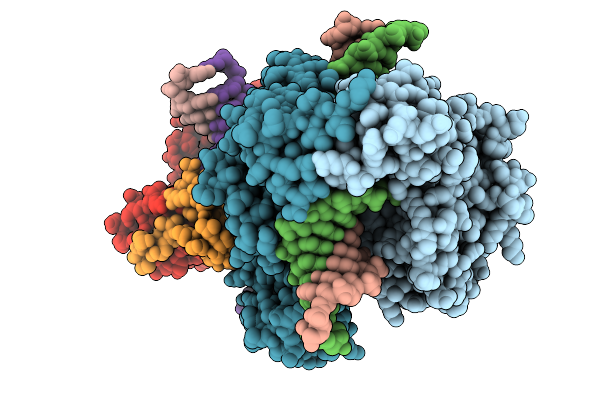 |
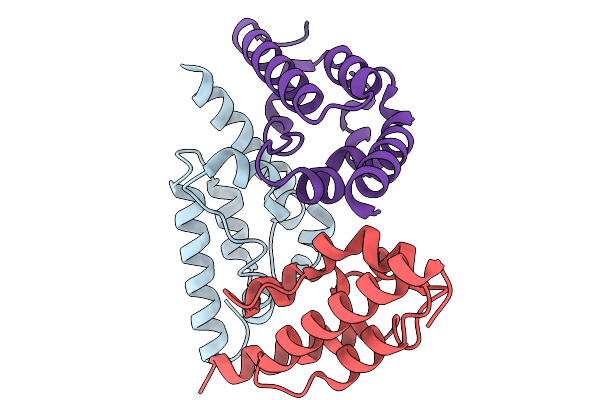
Pr1 Phage Heterodimeric Dna Ligase In Complex With 21-Mer Nicked Dna Containing Phage Nick Sequence
Organism: Providencia phage vb_pres_pr1
Method: X-RAY DIFFRACTION Resolution:3.16 Å Release Date: 2026-01-14 Classification: DNA BINDING PROTEIN Ligands: NMN, AMP, ZN |
 |
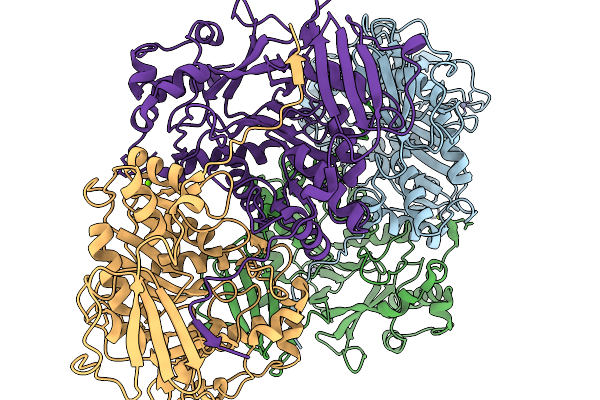
Organism: Streptococcus macedonicus
Method: X-RAY DIFFRACTION Resolution:2.68 Å Release Date: 2026-01-07 Classification: UNKNOWN FUNCTION |
 |
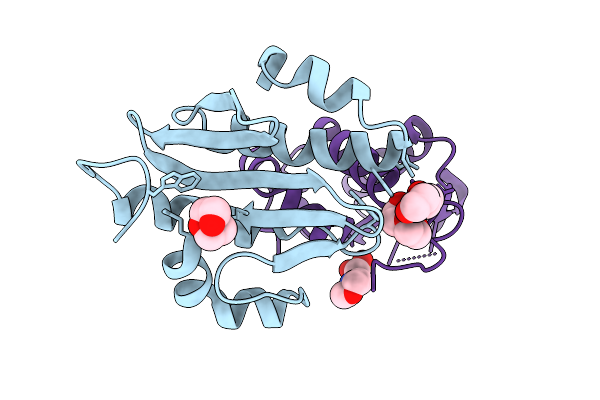
Organism: Clostridium cavendishii
Method: X-RAY DIFFRACTION Resolution:2.57 Å Release Date: 2025-11-19 Classification: LIGASE Ligands: ZN, MG |
 |
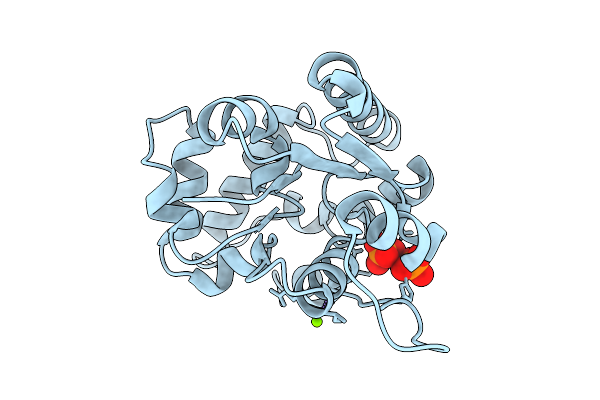
Organism: Listeria monocytogenes 10403s
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2025-11-19 Classification: METAL BINDING PROTEIN Ligands: PEG, MES |
 |
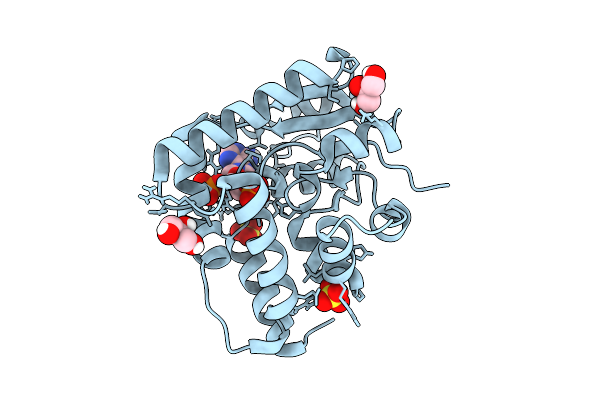
Crystal Structure Of Streptomyces Sp. Sulfotransferase Cpz8 Involved In Caprazamycins Synthesis
Organism: Streptomyces sp.
Method: X-RAY DIFFRACTION Resolution:1.38 Å Release Date: 2025-11-05 Classification: BIOSYNTHETIC PROTEIN Ligands: PO4, MG |
 |
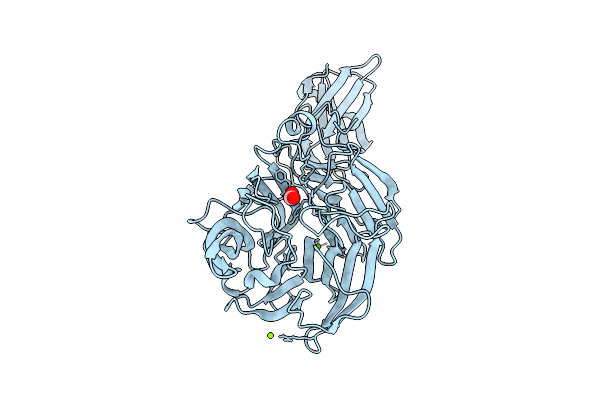
Crystal Structure Of Streptomyces Sp. Sulfotransferase Cpz8 In Complex With Pap, Involved In Caprazamycin Synthesis
Organism: Streptomyces sp.
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2025-11-05 Classification: BIOSYNTHETIC PROTEIN Ligands: GOL, A3P, SO4 |
 |
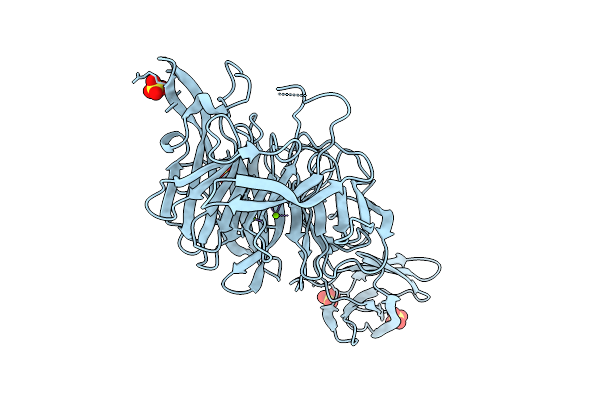
Crystal Structure Of Streptomyces Sp. Sulfotransferase Cpz4 Involved In Caprazamycins Synthesis
Organism: Streptomyces sp.
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2025-11-05 Classification: BIOSYNTHETIC PROTEIN Ligands: FMT, MG |
 |
Crystal Structure Of The Reaction Intermediate Of Streptomyces Sp. Sulfotransferase Cpz4 Involved In Caprazamycins Synthesis
Organism: Streptomyces sp.
Method: X-RAY DIFFRACTION Resolution:2.11 Å Release Date: 2025-11-05 Classification: BIOSYNTHETIC PROTEIN Ligands: SO4, MG |
 |
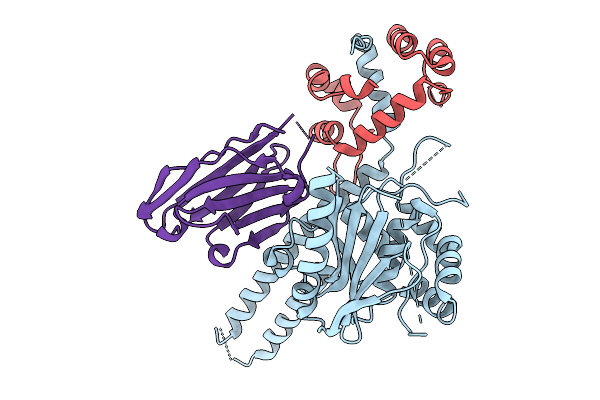
Organism: Streptomyces sp.
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-10-08 Classification: OXIDOREDUCTASE Ligands: NAD |
 |
Crystal Structure Of The Uch37 Rpn13 Deubad Complex Bound To An Inhibitory Nanobody In The Canonical Ubiquitin Binding Site
Organism: Homo sapiens, Camelidae mixed library
Method: X-RAY DIFFRACTION Resolution:2.41 Å Release Date: 2025-09-03 Classification: HYDROLASE Ligands: CL |
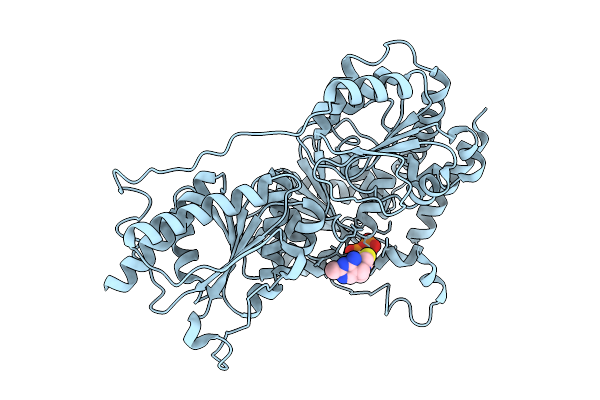 |
Organism: Listeria monocytogenes 10403s
Method: X-RAY DIFFRACTION Resolution:2.61 Å Release Date: 2025-07-30 Classification: TRANSFERASE Ligands: TPP, MG |
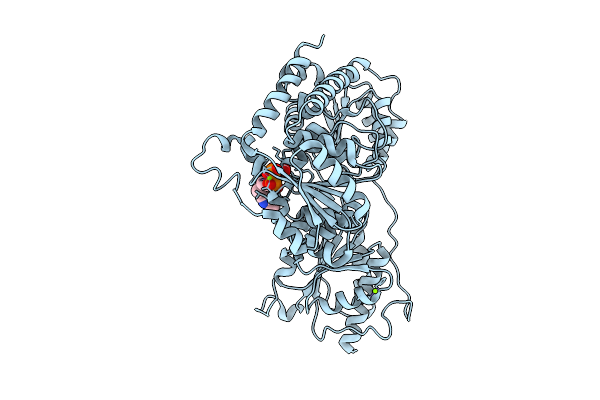 |
Crystal Structure Of L. Monocytogenes Mend With Mg2+ And Intermediate I Bound
Organism: Listeria monocytogenes 10403s
Method: X-RAY DIFFRACTION Resolution:2.79 Å Release Date: 2025-07-30 Classification: TRANSFERASE Ligands: TD6, MG |
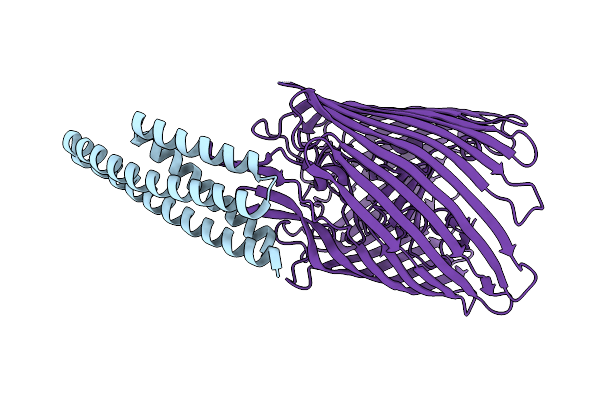 |
Cryo-Em Structure Of The Heme/Hemoglobin Transporter Chua, In Complex With De Novo Designed Binder G7
Organism: Synthetic construct, Escherichia coli cft073
Method: ELECTRON MICROSCOPY Release Date: 2025-05-21 Classification: TRANSPORT PROTEIN |
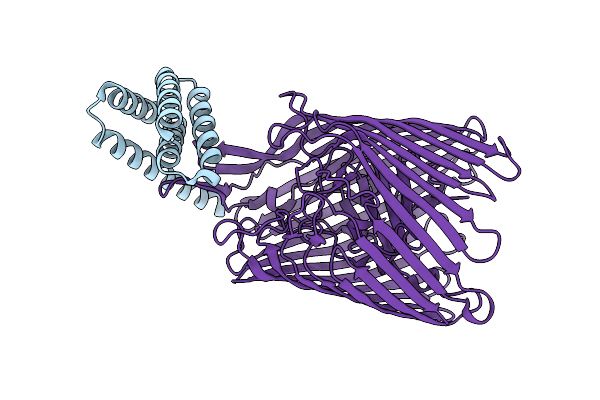 |
Cryo-Em Structure Of The Heme/Hemoglobin Transporter Chua, In Complex With De Novo Designed Binder H3
Organism: Synthetic construct, Escherichia coli cft073
Method: ELECTRON MICROSCOPY Release Date: 2025-05-21 Classification: TRANSPORT PROTEIN |
 |
Organism: Streptomyces sp.
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2025-05-21 Classification: SUGAR BINDING PROTEIN Ligands: BTB, EDO |

