Search Count: 19,985
All
Selected
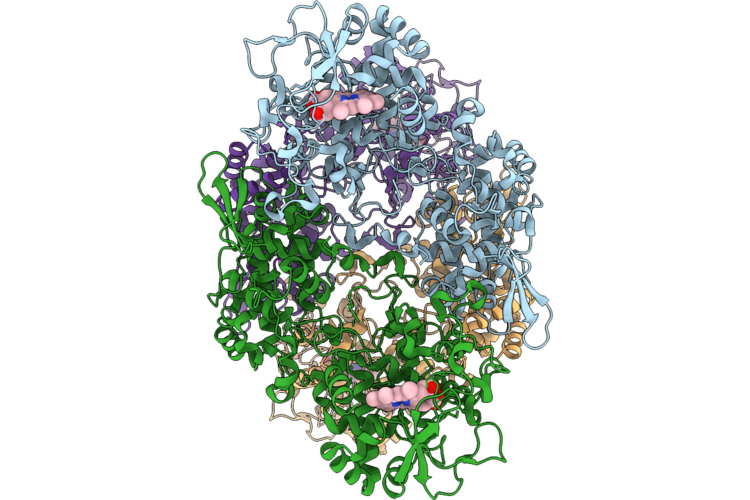 |
Cryoem Structure Of Eckatg S-Trp105 At 2.22 Angstrom Resolution Revealing An Asymmetric Sulfur Center In O=S-Trp
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM |
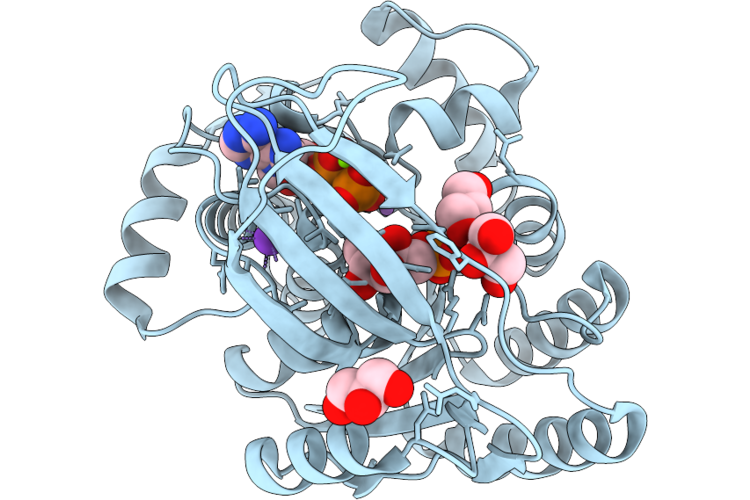 |
1-Phosphofructokinase (Fruk) From E. Coli With Bound Fructose 1-Phosphate And Adp
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: MG, F1X, ADP, LI, K, GOL |
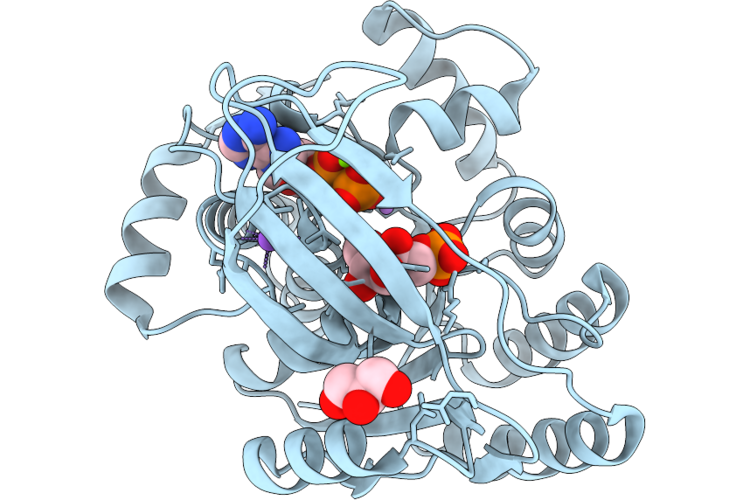 |
1-Phosphofructokinase Mutant K95H From E. Coli With Bound Fructose 1-Phosphate And Adp
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.54 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: MG, LI, NA, ADP, F1X, GOL |
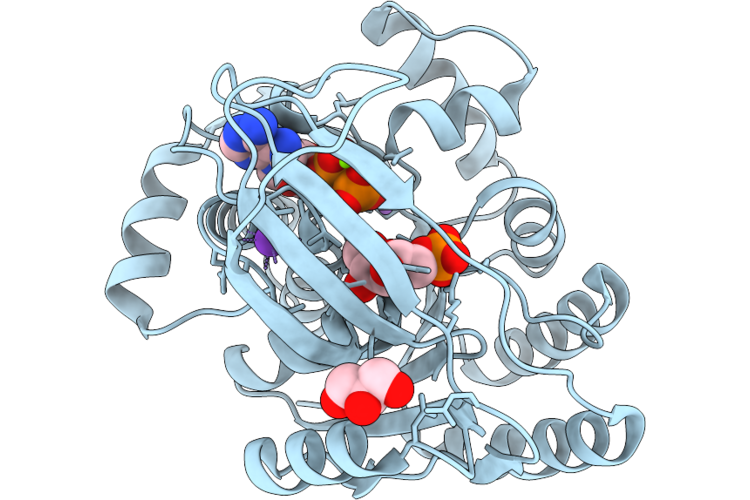 |
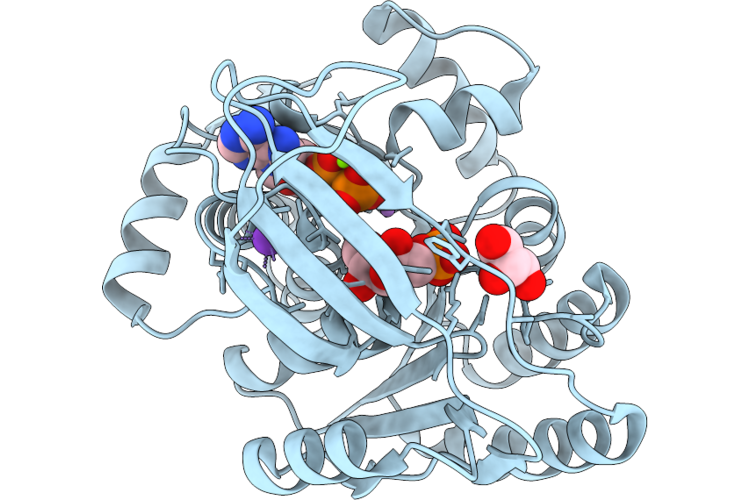
1-Phosphofructokinase Mutant K95H From E. Coli With Bound Fructose 6-Phosphate And Adp
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: GOL, ADP, F6P, MG, K, LI |
 |
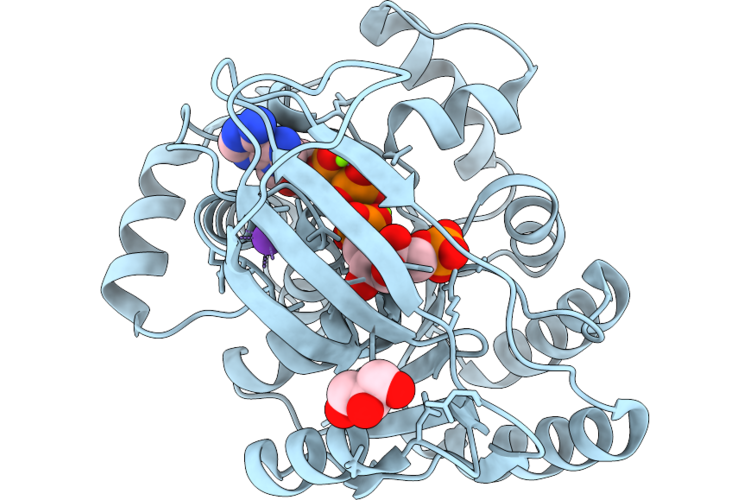
1-Phosphofructokinase Mutant K95T/G110S From E. Coli With Bound Fructose 1-Phosphate And Adp
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.41 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: MG, F1X, ADP, K, LI, GOL |
 |
1-Phosphofructokinase Mutant K95T/G110S From E. Coli With Bound Fructose-1,6-Bisphosphate And Adp
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: MG, ADP, FBP, GOL, K |
 |
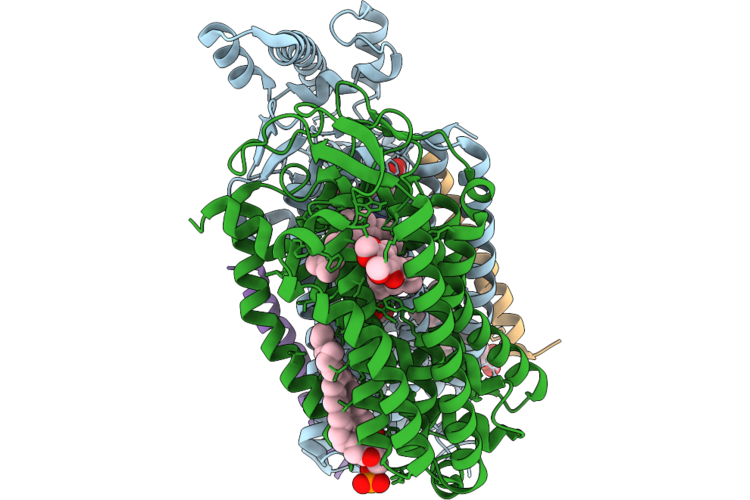
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM, A1JN4, MQ9, LPP, UQ8, OXY, PGT, POV |
 |
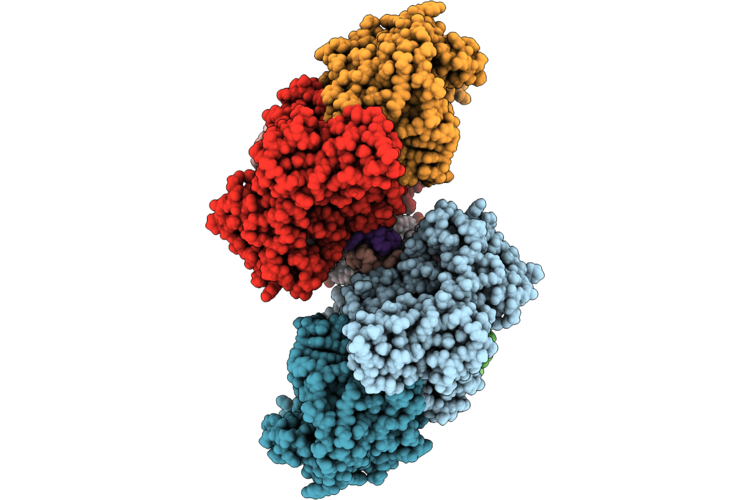
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: A1JN4, HEM, LPP, OXY, UQ8 |
 |
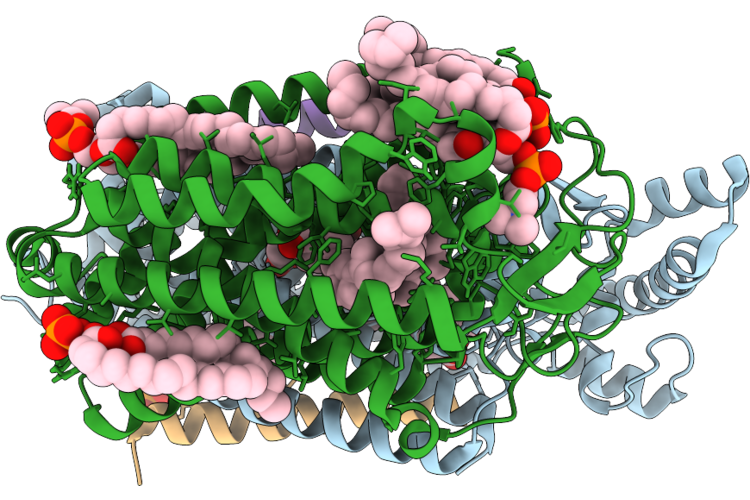
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM, LPP, UQ8, 0NI, OXY, A1JN4, PGT, POV |
 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: A1JN4, HEM, LPP, OXY, POV, MQ9 |
 |
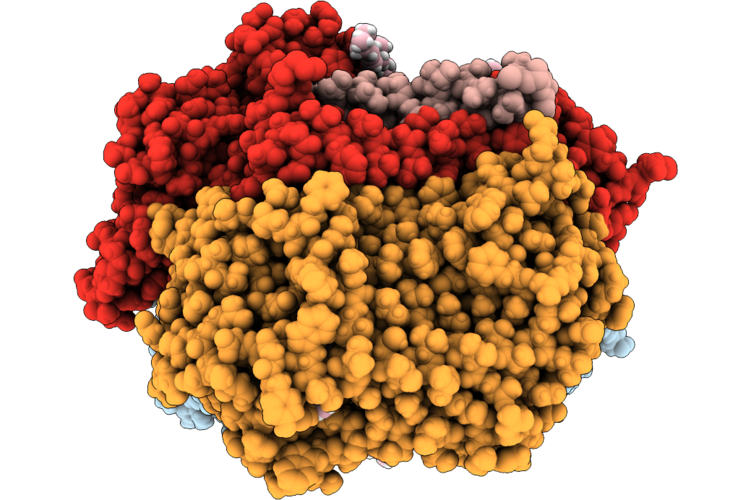
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: A1JN4, HEM, LPP, OXY, UQ8 |
 |
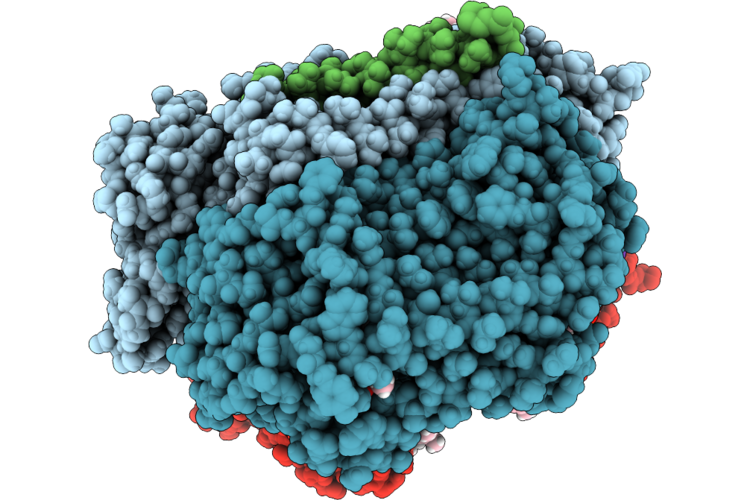
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM, A1JN4, LPP, UQ8, MQ8, OXY |
 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: LPP, UQ8, HEM, MQ8, A1JN4, OXY |
 |
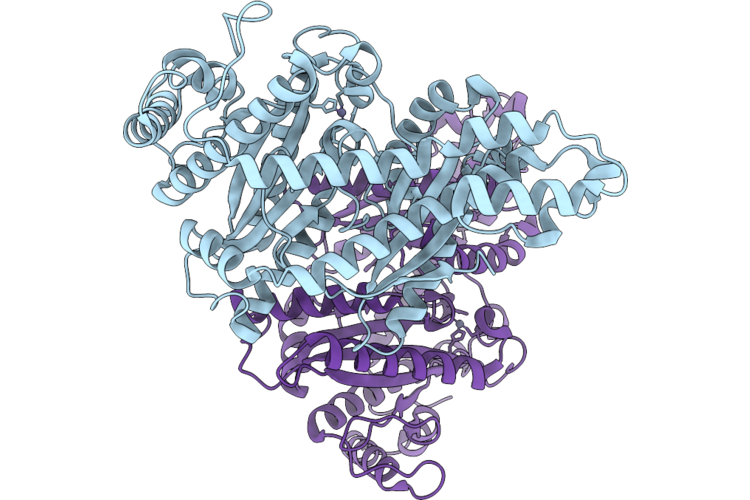
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.13 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Dimer, Major State, Active Conformation
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.15 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Dimer, Minor State
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.45 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Reducing Conditions, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.06 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Reducing Conditions, Dimer, Major State, Inactive Conformation
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.12 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Reducing Conditions, Dimer, Minor State
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.27 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca C283A/C284A Inactive Mutant, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.08 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |

