Search Count: 2,009
All
Selected
 |
Organism: Aspergillus nidulans fgsc a4
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: FE, SO4, IP1 |
 |
Organism: Aspergillus nidulans fgsc a4
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: FE, SO4, IP1 |
 |
Organism: Aspergillus nidulans fgsc a4
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: SO4, FE, ACV |
 |
Organism: Aspergillus nidulans fgsc a4
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: FE, SO4, IP1 |
 |
Organism: Nocardia brasiliensis
Method: X-RAY DIFFRACTION Resolution:3.15 Å Release Date: 2026-03-11 Classification: TRANSFERASE Ligands: MG, TON, A1BYH, TRT |
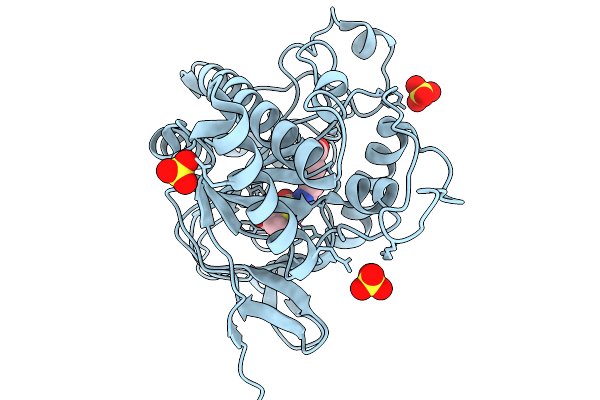 |
Isopenicillin N Synthase N252E Variant In Complex With Fe And Ipn After O2 Exposure
Organism: Aspergillus nidulans fgsc a4
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: FE, SO4, IP1 |
 |
Organism: Ruminococcus bromii l2-63
Method: ELECTRON MICROSCOPY Release Date: 2025-10-22 Classification: HYDROLASE |
 |
Organism: Ruminococcus bromii l2-63
Method: ELECTRON MICROSCOPY Release Date: 2025-10-22 Classification: HYDROLASE |
 |
Organism: Ruminococcus bromii l2-63
Method: ELECTRON MICROSCOPY Release Date: 2025-10-22 Classification: HYDROLASE |
 |
Organism: Ruminococcus bromii l2-63
Method: ELECTRON MICROSCOPY Release Date: 2025-10-22 Classification: HYDROLASE |
 |
Structural Basis For The Polymer-Protein Binding Mechanism Of Polyvinyl Alcohol Esterase
Organism: Comamonas sp. nyz500
Method: X-RAY DIFFRACTION Resolution:1.92 Å Release Date: 2025-10-08 Classification: HYDROLASE Ligands: SO4, ACT, GOL, CL, NA |
 |
Structural Basis For The Polymer-Protein Binding Mechanism Of Polyvinyl Alcohol Esterase
Organism: Comamonas sp. nyz500
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-10-08 Classification: HYDROLASE Ligands: GOL |
 |
Structural Basis For The Polymer-Protein Binding Mechanism Of Polyvinyl Alcohol Esterase
Organism: Comamonas sp. nyz500
Method: X-RAY DIFFRACTION Resolution:2.84 Å Release Date: 2025-10-08 Classification: HYDROLASE Ligands: CL, GOL, ACT |
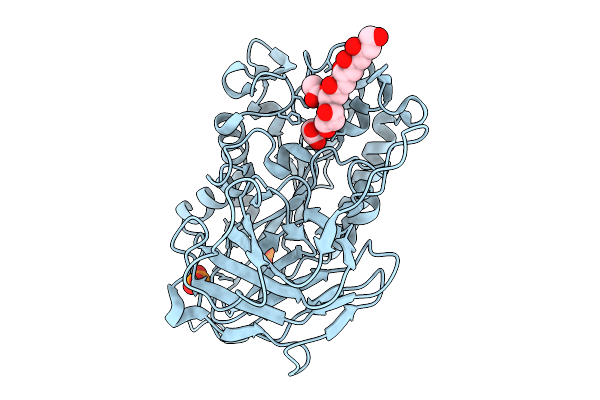 |
Structural Basis For The Polymer-Protein Binding Mechanism Of Polyvinyl Alcohol Esterase
Organism: Comamonas sp. nyz500
Method: X-RAY DIFFRACTION Resolution:2.13 Å Release Date: 2025-10-08 Classification: HYDROLASE Ligands: CL, A1EGH, ACT, PO4 |
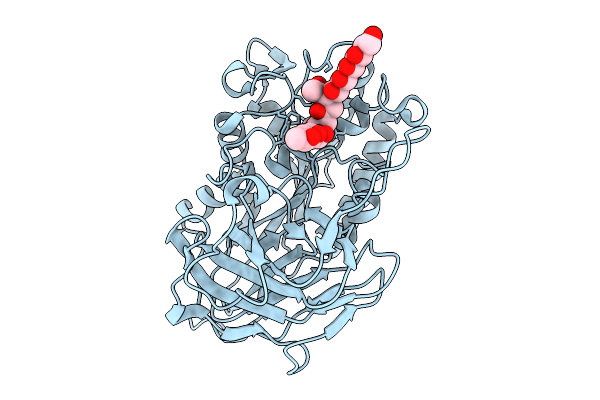 |
Structural Basis For The Polymer-Protein Binding Mechanism Of Polyvinyl Alcohol Esterase
Organism: Comamonas sp. nyz500
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2025-10-08 Classification: HYDROLASE Ligands: A1EGL, CL |
 |
Ensemble Refined Structure Of Cotb2 In Complex With 2-Fluoro-3,7,18-Dolabellatriene
Organism: Streptomyces melanosporofaciens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-09-17 Classification: HYDROLASE Ligands: MG, PPV, EXW, MPD, CL, CCN, 2PE, K |
 |
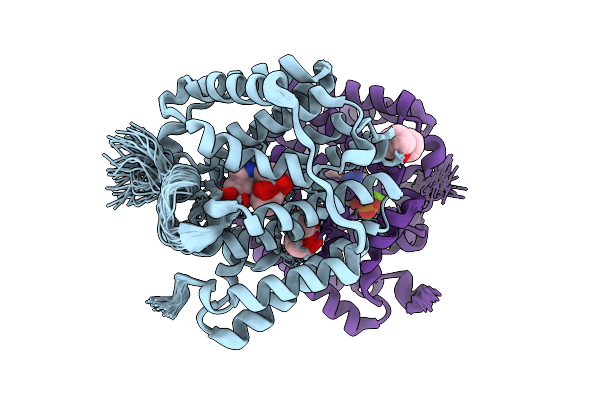
Organism: Streptomyces melanosporofaciens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-09-17 Classification: HYDROLASE Ligands: AHD, MG |
 |
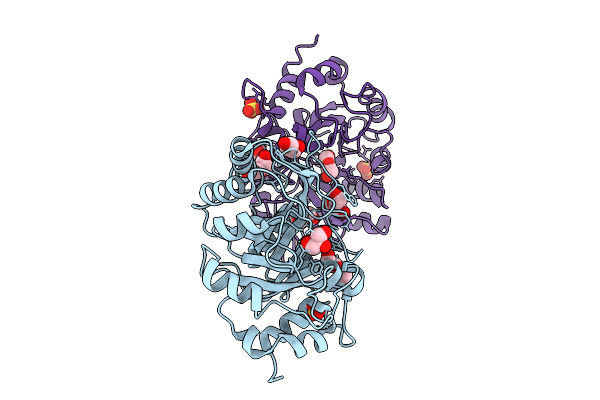
Organism: Streptomyces melanosporofaciens
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2025-09-17 Classification: HYDROLASE Ligands: MPD, AHD, MG, CL |
 |
Organism: Metschnikowia pulcherrima
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2025-08-27 Classification: HYDROLASE Ligands: PEG, GOL, SO4, NAG |
 |
The Glycoside Hydrolase Family 71 (Gh71) Member Angh71C From Aspergillus Nidulans.
Organism: Aspergillus nidulans fgsc a4
Method: X-RAY DIFFRACTION Resolution:1.33 Å Release Date: 2025-06-25 Classification: CARBOHYDRATE Ligands: SO4, BTB, NA |

