Search Count: 5,934
All
Selected
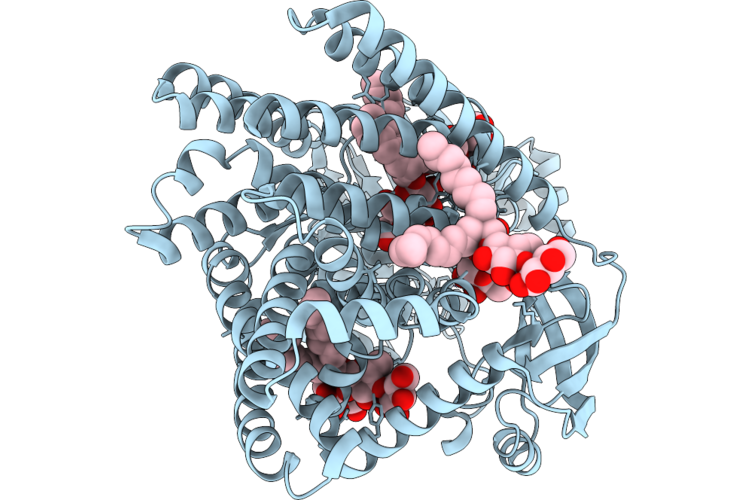 |
Organism: Halalkalibacterium halodurans c-125
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: MEMBRANE PROTEIN Ligands: AV0, ZN |
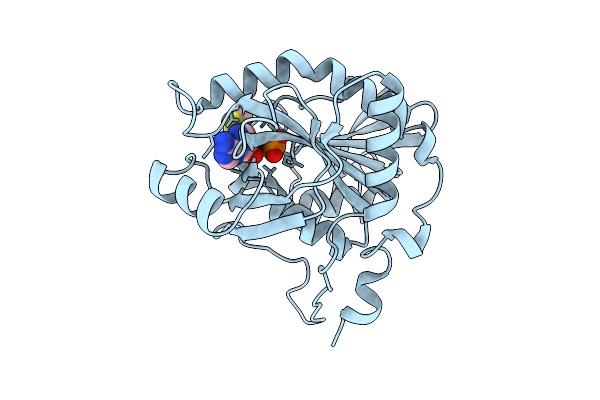 |
Crystal Structure Of Mtap From Aeropyrum Pernix Complex With Mta At 343K, Soaked In Potassium Phosphate Ph7.0 With Mta
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: PO4, MTA |
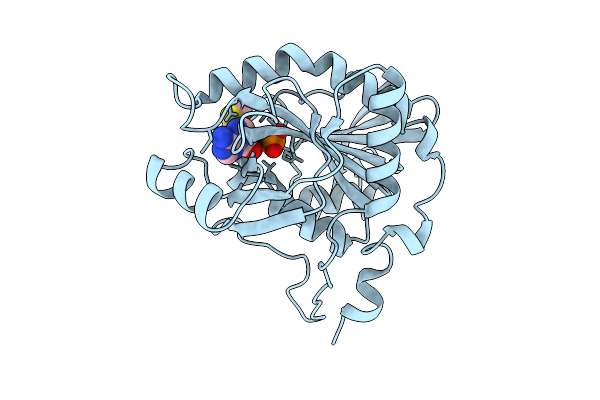 |
Crystal Structure Of Mtap From Aeropyrum Pernix Complex With Mta At 323K, Soaked In Potassium Phosphate Ph7.0 With Mta
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.59 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: PO4, MTA |
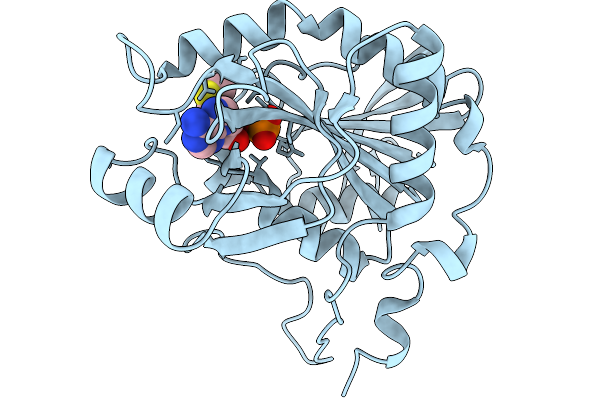 |
Crystal Structure Of Mtap From Aeropyrum Pernix Complex With Mta At 333K, Soaked In Potassium Phosphate Ph7.0 With Mta
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2026-03-25 Classification: TRANSFERASE Ligands: PO4, MTA |
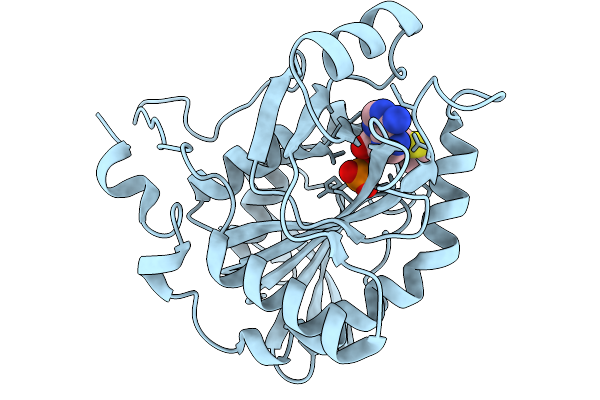 |
Crystal Structure Of Mtap From Aeropyrum Pernix Complex With Mta At 293K, Soaked In Potassium Phosphate Ph7.0 With Mta
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2026-03-25 Classification: TRANSFERASE Ligands: PO4, MTA |
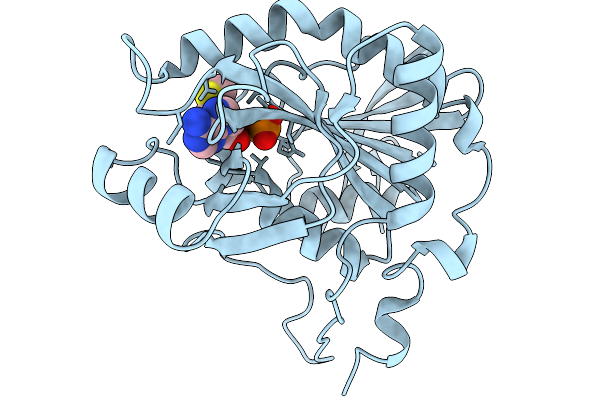 |
Crystal Structure Of 5'-Deoxy-5'-Methylthioadenosine Phosphorylase From Aeropyrum Pernix Complex With 5'-Deoxy-5'-Methylthioadenosine 353K 1.77 Angstrom
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2026-03-25 Classification: TRANSFERASE Ligands: PO4, MTA |
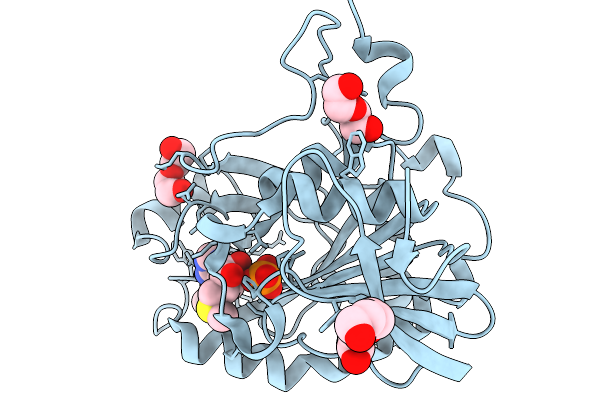 |
Crystal Structure Of Mtap From Aeropyrum Pernix Complex With Mta At 100K, Soaked In Potassium Phosphate Ph7.0 With Mta
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.19 Å Release Date: 2026-03-25 Classification: TRANSFERASE Ligands: PO4, MTA, PEG, PGE |
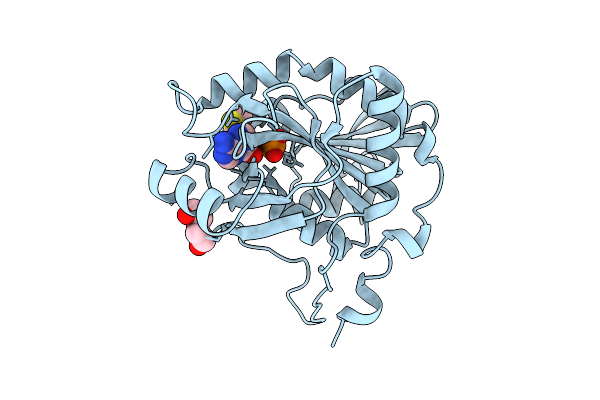 |
Crystal Structure Of 5'-Deoxy-5'-Methylthioadenosine Phosphorylase From Aeropyrum Pernix Complex With 5'-Deoxy-5'-Methylthioadenosine 343K 1.58 Angstrom
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2026-03-11 Classification: TRANSFERASE Ligands: PO4, MTA, PEG |
 |
Crystal Structure Of 5'-Deoxy-5'-Methylthioadenosine Phosphorylase From Aeropyrum Pernix Complex With 5'-Deoxy-5'-Methylthioadenosine 100K
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.37 Å Release Date: 2026-03-04 Classification: TRANSFERASE Ligands: PO4, MTA, PEG, PGE |
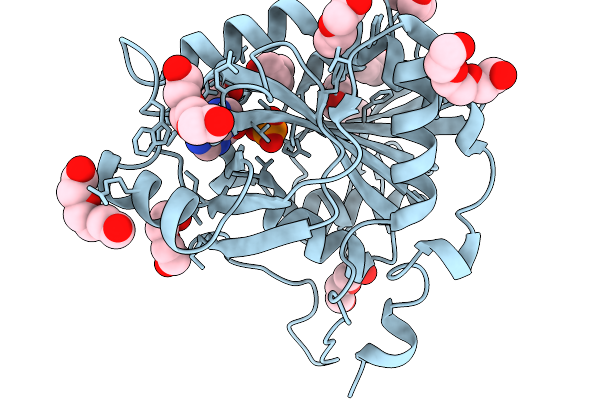 |
Crystal Structure Of 5'-Deoxy-5'-Methylthioadenosine Phosphorylase From Aeropyrum Pernix 100K
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.37 Å Release Date: 2026-03-04 Classification: TRANSFERASE Ligands: PO4, ADE, PGE, PEG |
 |
Crystal Structure Of 5'-Deoxy-5'-Methylthioadenosine Phosphorylase From Aeropyrum Pernix 323K
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.59 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: PO4, ADE, PEG |
 |
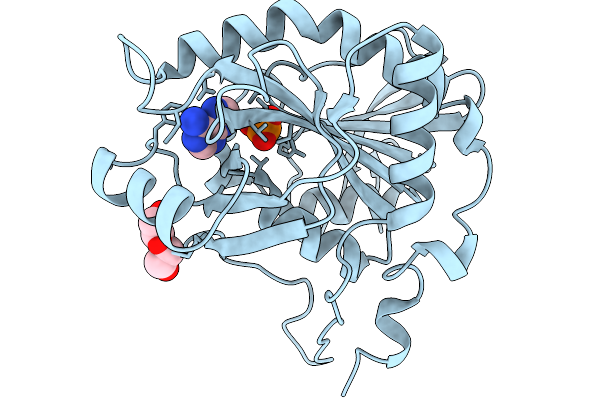
Crystal Structure Of 5'-Deoxy-5'-Methylthioadenosine Phosphorylase From Aeropyrum Pernix 293K
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: PO4, PEG, ADE |
 |
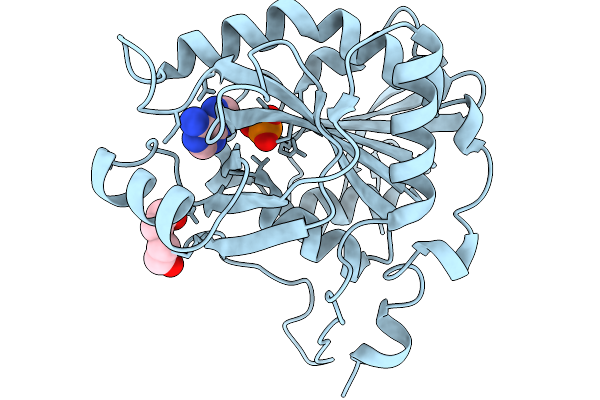
Crystal Structure Of 5'-Deoxy-5'-Methylthioadenosine Phosphorylase From Aeropyrum Pernix 343K
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-02-11 Classification: TRANSFERASE Ligands: PO4, PEG, ADE |
 |
Crystal Structure Of 5'-Deoxy-5'-Methylthioadenosine Phosphorylase From Aeropyrum Pernix 333K
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2026-02-11 Classification: TRANSFERASE Ligands: PO4, PEG, ADE |
 |
Crystal Structure Of A Leaf-Branch Compost Cutinase Variant, Iccg-H218S/F222I
Organism: Unidentified prokaryotic organism
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2026-01-21 Classification: HYDROLASE Ligands: ACT, CA |
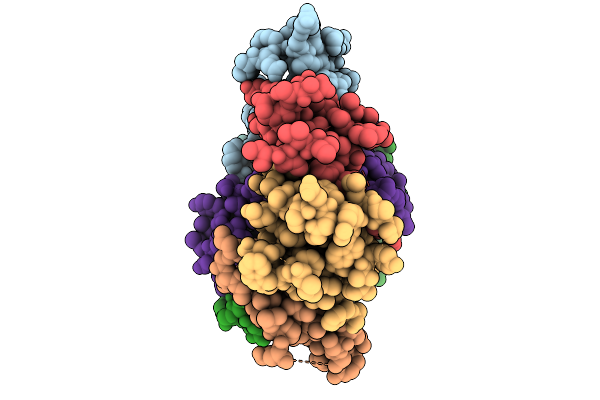 |
Structure Of The Bacteroides Fragilis Nctc9343 T6Ss Hcp2-Hcp3 Heterohexamer In Complex With The Effector Bte1
Organism: Bacteroides fragilis nctc 9343
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2026-01-21 Classification: STRUCTURAL PROTEIN |
 |
Organism: Bacteroides fragilis nctc 9343
Method: ELECTRON MICROSCOPY Resolution:3.11 Å Release Date: 2026-01-21 Classification: STRUCTURAL PROTEIN |
 |
Structure Of Clim-Stalled Bacillus Subtilis 70S Ribosome With Release Factor Bound In The A-Site
Organism: Clostridioides difficile 630, Bacillus subtilis subsp. subtilis str. 168
Method: ELECTRON MICROSCOPY Resolution:2.30 Å Release Date: 2026-01-14 Classification: RIBOSOME Ligands: ZN |
 |
Organism: Clostridioides difficile 630, Bacillus subtilis subsp. subtilis str. 168
Method: ELECTRON MICROSCOPY Resolution:2.30 Å Release Date: 2026-01-14 Classification: RIBOSOME |
 |
Structure Of Clim-Stalled Bacillus Subtilis 70S Ribosome With Trna-Tyr In The A-Site
Organism: Clostridioides difficile 630, Bacillus subtilis subsp. subtilis str. 168
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-01-14 Classification: RIBOSOME Ligands: ZN |

