Search Count: 1,254
 |
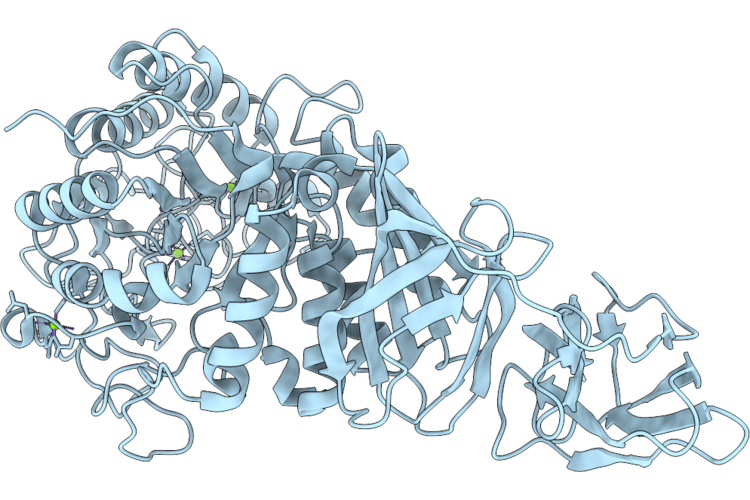
Crystal Structure Of The C-Terminal Of An Alpha-Amylase Family Glycosyl Hydrolase From Vibrio Parahaemolyticus
Organism: Vibrio parahaemolyticus
Method: X-RAY DIFFRACTION Resolution:1.42 Å Release Date: 2026-04-29 Classification: HYDROLASE Ligands: GOL, MG |
 |
Crystal Structure Of The N-Terminal Of An Alpha-Amylase Family Glycosyl Hydrolase From Vibrio Parahaemolyticus
Organism: Vibrio parahaemolyticus
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2026-04-29 Classification: HYDROLASE Ligands: MG |
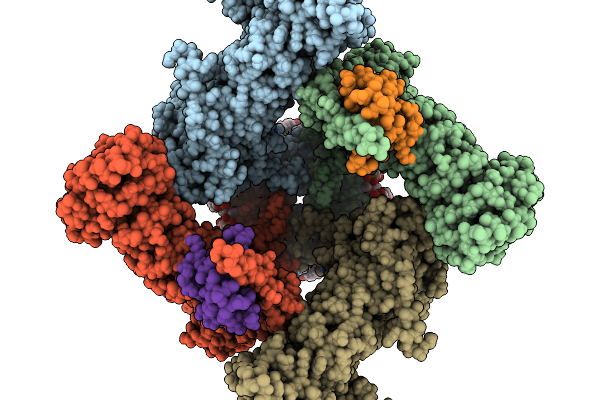 |
Organism: Halyomorpha halys, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN Ligands: 6OU, LBN, CA, D39 |
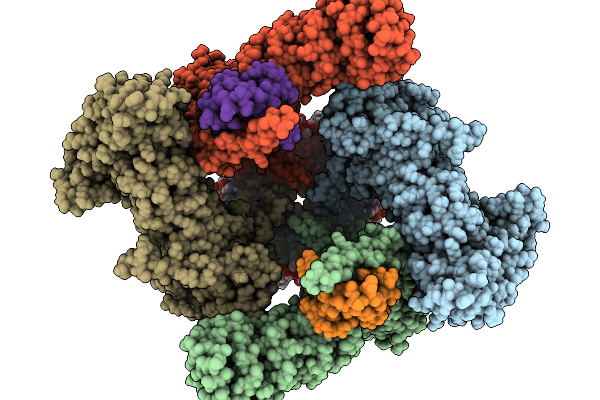 |
Organism: Halyomorpha halys, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.49 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN Ligands: 6OU, LBN, NCA, CA, D39 |
 |
Structure Of Nanchung-Inactive-Calmodulin In Complex With Nicotinamide, Edta
Organism: Halyomorpha halys, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.85 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN Ligands: 6OU, LBN, NCA, D39 |
 |
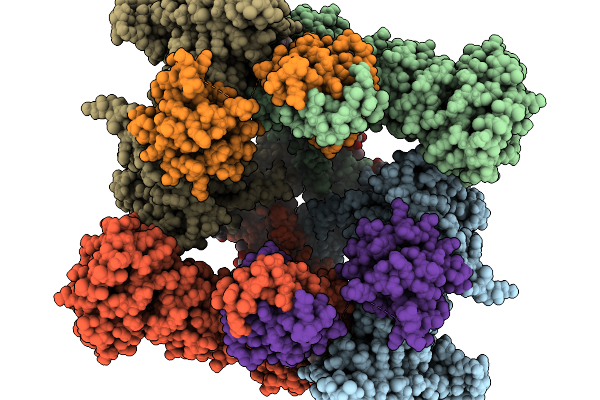
Structure Of Nanchung-Inactive-Calmodulin In Complex With Afidopyropen And Calcium
Organism: Halyomorpha halys, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.01 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN Ligands: 6OU, LBN, A1B32, CA, D39 |
 |
Structure Of Nanchung-Inactive-Calmodulin In Complex With Afidopyropen, Edta
Organism: Halyomorpha halys, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.13 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN Ligands: 6OU, LBN, A1B32, D39 |
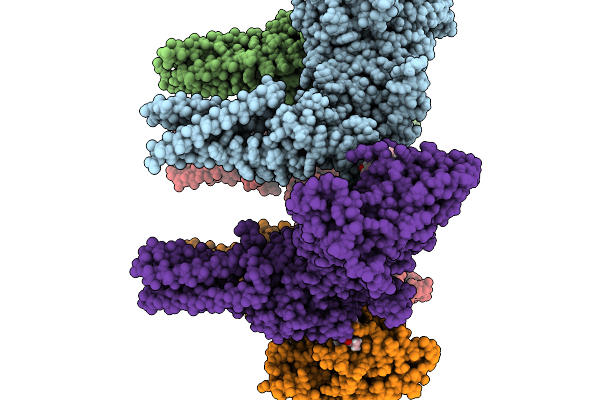 |
Organism: Halyomorpha halys
Method: ELECTRON MICROSCOPY Resolution:3.28 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN Ligands: A1B32 |
 |
Atomic Structure Of Vibrio Effector Fragment Vopv Bound To Beta-Cytoplasmic/Gamma1-Cytoplasmic F-Actin
Organism: Homo sapiens, Vibrio parahaemolyticus
Method: ELECTRON MICROSCOPY Release Date: 2025-11-19 Classification: STRUCTURAL PROTEIN Ligands: MG, ANP |
 |
Organism: Vibrio parahaemolyticus
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2025-08-06 Classification: HYDROLASE |
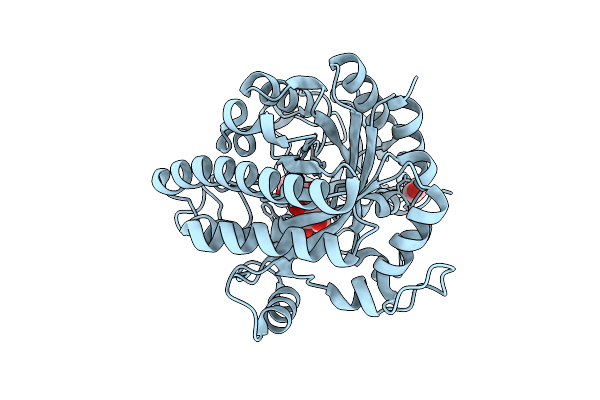 |
Organism: Vibrio parahaemolyticus
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-07-30 Classification: HYDROLASE Ligands: EDO, B3P |
 |
Crystal Structure Of The Type Iii Secretion Chaperone Veca From Vibrio Parahaemolyticus
Organism: Vibrio parahaemolyticus
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-06-25 Classification: CHAPERONE Ligands: PO4, TBR |
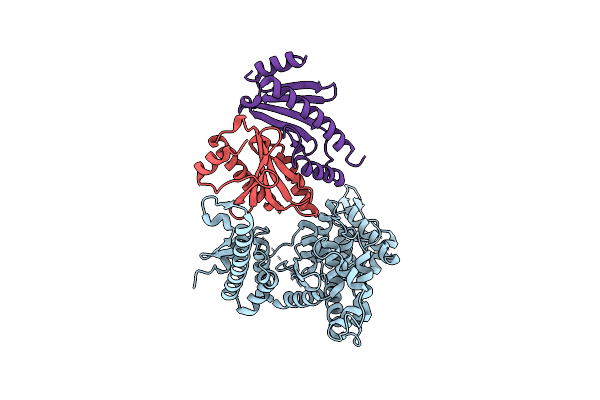 |
Crystal Structure Of The Virulence Effector Vepa In Complex With Its Secretion Chaperone Veca
Organism: Vibrio parahaemolyticus
Method: X-RAY DIFFRACTION Resolution:2.49 Å Release Date: 2025-06-25 Classification: TOXIN |
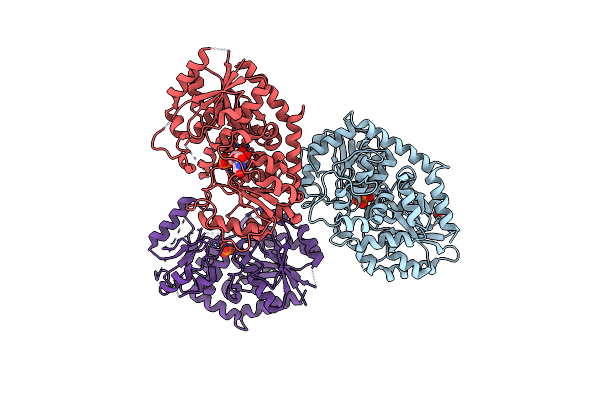 |
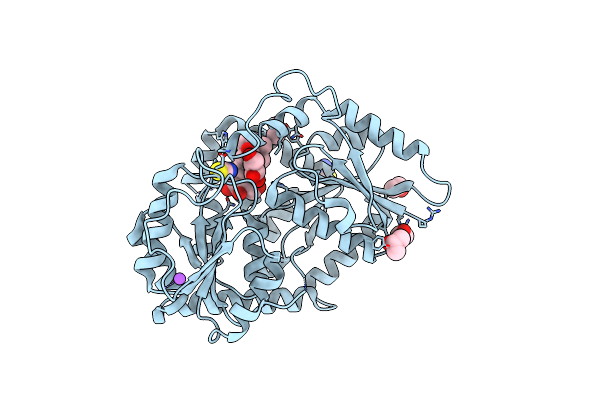
Organism: Stevia rebaudiana
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2025-05-21 Classification: TRANSFERASE Ligands: UPG, IPA, PO4 |
 |
Organism: Stevia rebaudiana
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-05-21 Classification: TRANSFERASE |
 |
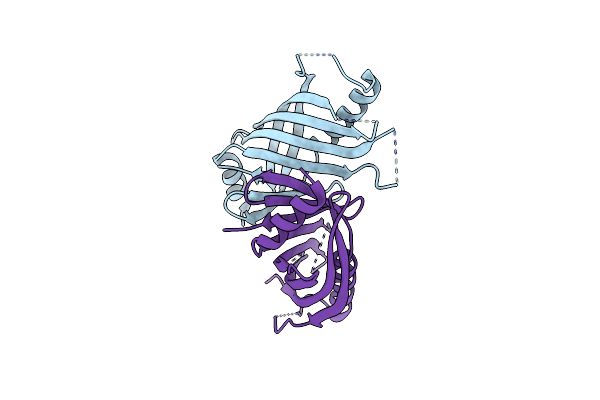
Organism: Vibrio parahaemolyticus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2024-09-25 Classification: HYDROLASE |
 |
Organism: Vibrio parahaemolyticus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2024-09-11 Classification: MEMBRANE PROTEIN |
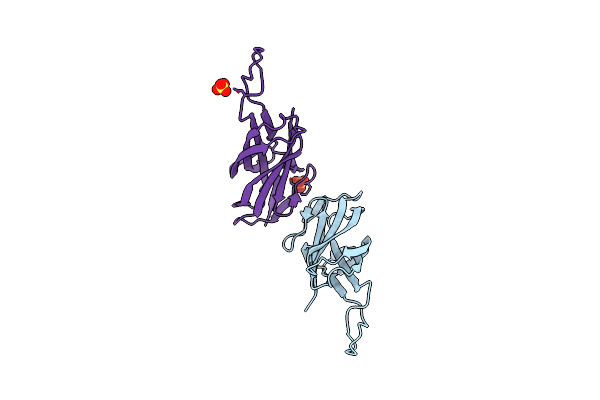 |
Intramolecular Ester Bond-Containing Repeat Domain From Gemella Massiliensis Adhesin
Organism: Gemella massiliensis
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2024-06-19 Classification: CELL ADHESION Ligands: SO4 |
 |
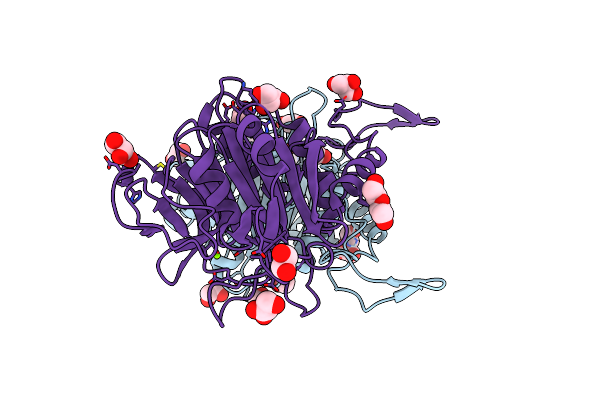
Organism: Vibrio parahaemolyticus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2024-05-22 Classification: UNKNOWN FUNCTION Ligands: ZN |
 |
Organism: Arthrobacter citreus
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2024-02-07 Classification: HYDROLASE Ligands: MG, EDO, TRS, PEG, GOL |

