Search Count: 1,776
All
Selected
 |
Organism: Mycoplasmoides pneumoniae m129
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN |
 |
Crystal Structure Of Terpeniod Cyclase Spsods From Rhizobacterium Serratia Plymuthica
Organism: Serratia plymuthica 4rx13
Method: X-RAY DIFFRACTION Resolution:2.22 Å Release Date: 2026-03-18 Classification: LYASE Ligands: GOL |
 |
Organism: Serratia plymuthica 4rx13
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2026-03-18 Classification: LYASE Ligands: POP, MG, TRS |
 |
Organism: Serratia plymuthica 4rx13
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-03-18 Classification: LYASE Ligands: GPP, MG, TRS, POP |
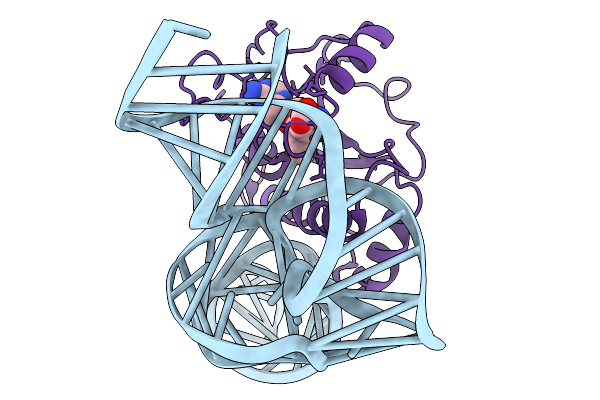 |
Organism: Saccharomyces cerevisiae bmn1-35, Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: RNA BINDING PROTEIN/RNA Ligands: AN6 |
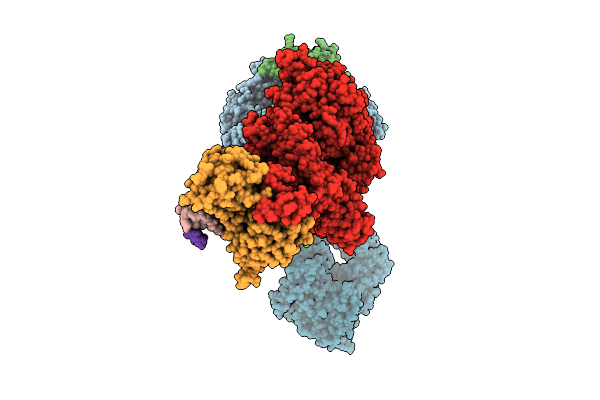 |
Organism: Human alphaherpesvirus 1 strain r-15
Method: ELECTRON MICROSCOPY Release Date: 2025-12-31 Classification: VIRAL PROTEIN Ligands: ZN, A1BXB |
 |
Organism: Human alphaherpesvirus 1 strain r-15
Method: ELECTRON MICROSCOPY Release Date: 2025-12-31 Classification: VIRAL PROTEIN Ligands: ZN, A1CHB |
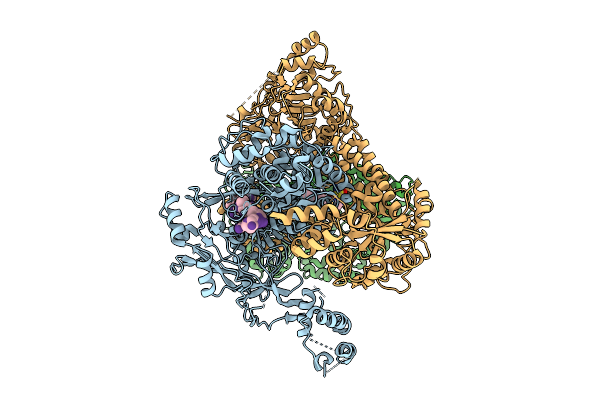 |
Organism: Human alphaherpesvirus 1 strain r-15
Method: ELECTRON MICROSCOPY Release Date: 2025-12-31 Classification: VIRAL PROTEIN Ligands: ZN, A1BXB |
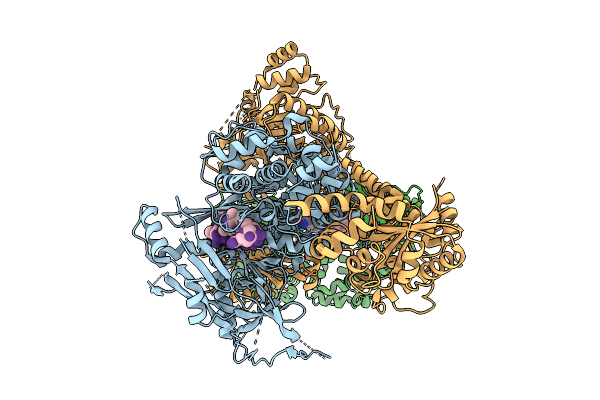 |
Organism: Human alphaherpesvirus 1 strain r-15
Method: ELECTRON MICROSCOPY Release Date: 2025-12-31 Classification: VIRAL PROTEIN Ligands: ZN, A1BXD |
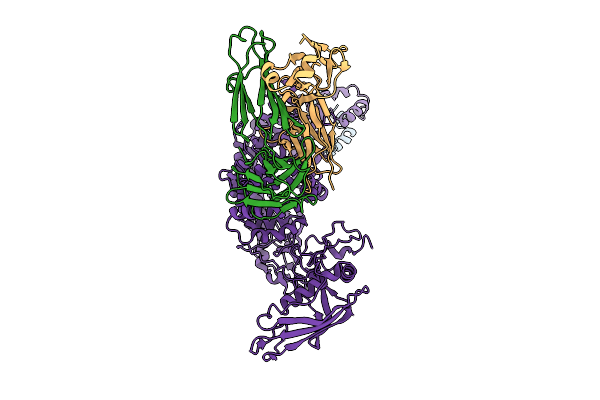 |
P116 From Mycoplasma Pneumoniae In Complex With Mild Growth Suppressor Monoclonal Antibody
Organism: Mus musculus, Mycoplasmoides pneumoniae m129
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: PROTEIN TRANSPORT |
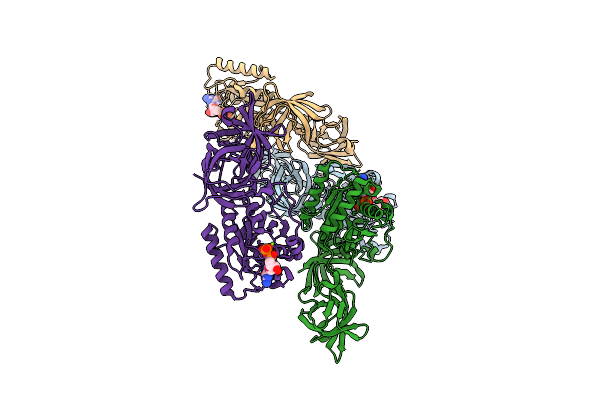 |
Organism: Mycoplasma pneumoniae (strain atcc 29342 / m129 / subtype 1)
Method: X-RAY DIFFRACTION Resolution:2.24 Å Release Date: 2025-10-29 Classification: BIOSYNTHETIC PROTEIN Ligands: GDP, MG |
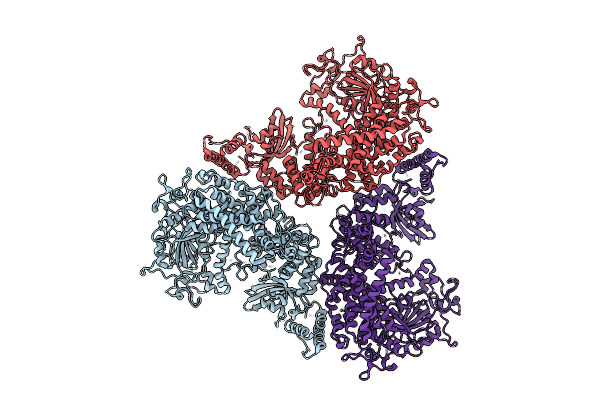 |
Cryo-Em Structure Of Essential Mycoplasma Pneumoniae Lipoprotein Mpn444 Homotrimer
Organism: Mycoplasmoides pneumoniae m129
Method: ELECTRON MICROSCOPY Release Date: 2025-09-17 Classification: UNKNOWN FUNCTION |
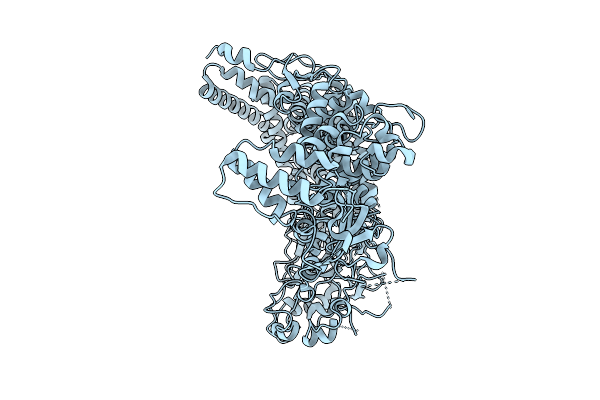 |
Organism: Mycoplasmoides pneumoniae m129
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: UNKNOWN FUNCTION |
 |
Organism: Methanobrevibacter ruminantium m1
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-04-23 Classification: HYDROLASE |
 |
Organism: Methanobrevibacter ruminantium m1
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-04-23 Classification: HYDROLASE |
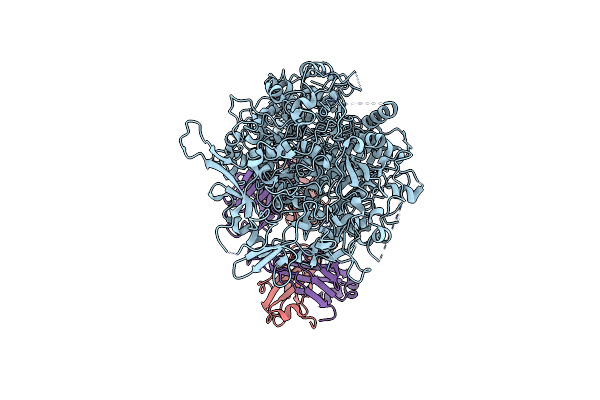 |
Single-Particle Cryo-Em Of Mycoplasma Pneumoniae Adhesin P1 Complexed With The Anti-Adhesive Fab Fragment.
Organism: Mycoplasmoides pneumoniae m129, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-03-05 Classification: CELL ADHESION |
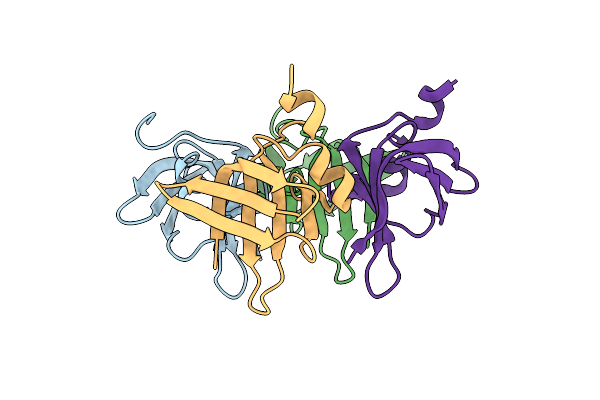 |
Self Assembly Domain Of The Surface Layer Protein Of Viridibacillus Arvi (Aa 765-844)
Organism: Viridibacillus arvi
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2024-08-28 Classification: STRUCTURAL PROTEIN Ligands: BR |
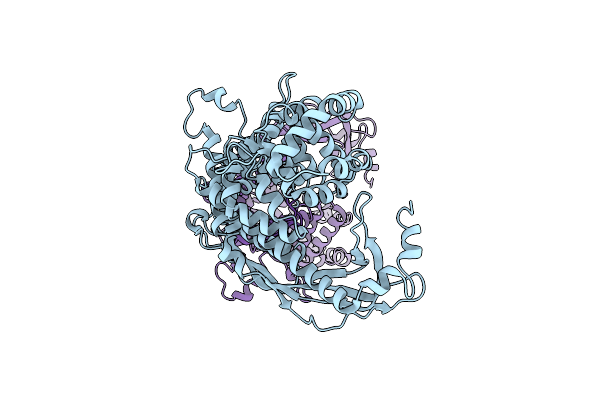 |
P116 Dimer In The Full State (Pdb Structure Of The Full-Length Ectodomain Truncated To Amino Acids 246-818)
Organism: Mycoplasmoides pneumoniae m129
Method: ELECTRON MICROSCOPY Release Date: 2024-05-29 Classification: LIPID BINDING PROTEIN |
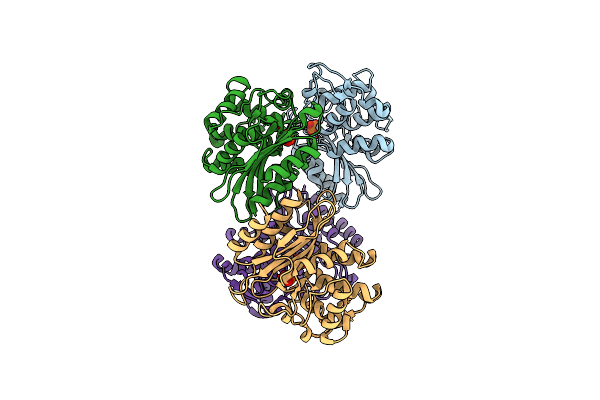 |
Xfel Structure Of Mycobacterium Tuberculosis Beta Lactamase Microcrystals Mixed With Sulbactam For 3 Ms
Organism: Mycobacterium tuberculosis str. beijing/w bt1
Method: X-RAY DIFFRACTION Resolution:2.62 Å Release Date: 2023-09-20 Classification: HYDROLASE/Inhibitor Ligands: PO4 |
 |
Crystal Structure Of Duf371 Domain-Containing Protein From Methanobrevibacter Ruminantium
Organism: Methanobrevibacter ruminantium
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2023-07-12 Classification: STRUCTURAL GENOMICS |

