Search Count: 1,712
All
Selected
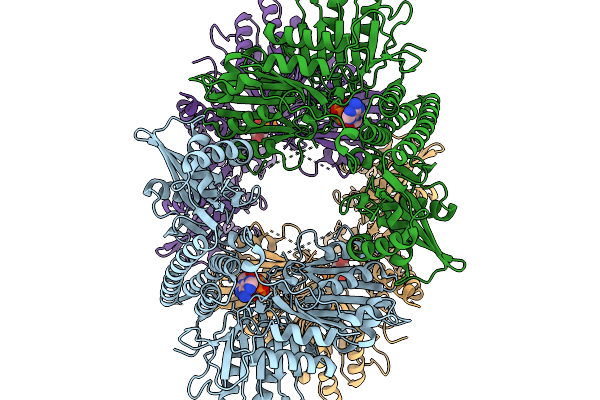 |
Organism: Citrobacter freundii
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: TRANSFERASE Ligands: ATP |
 |
Structure Of Mycobacterium Tuberculosis Inha In Complex With Pyridomycin Derivative Kv37A (Compound 9)
Organism: Mycobacterium tuberculosis '98-r604 inh-rif-em'
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: A1JG0 |
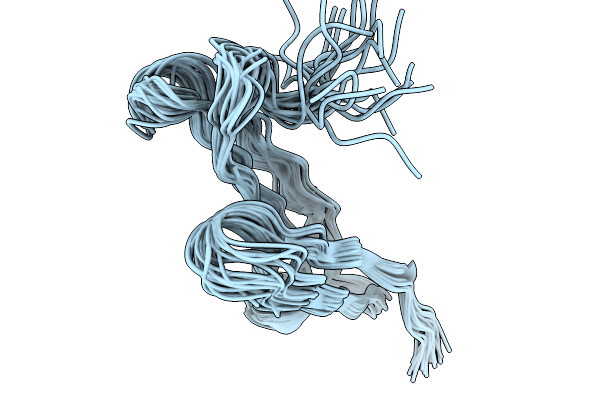 |
Organism: Colletotrichum orbiculare
Method: SOLUTION NMR Release Date: 2025-11-19 Classification: VIRAL PROTEIN |
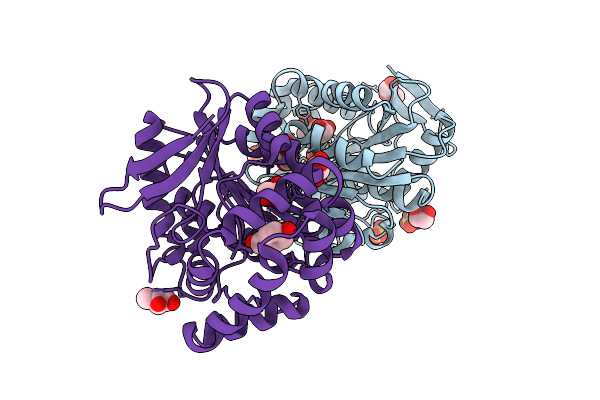 |
The Crystal Structure Of Alpha-Beta-Fold_Hydrolase From Microlunatus Sagamiharensis.
Organism: Microlunatus sagamiharensis
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-10-29 Classification: HYDROLASE Ligands: SO4, CL, 4HP, GOL, 3HR |
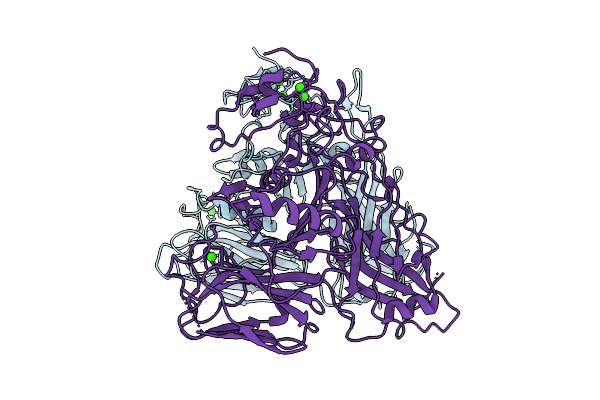 |
Organism: Clostridioides difficile r20291
Method: ELECTRON MICROSCOPY Resolution:3.56 Å Release Date: 2025-07-16 Classification: TOXIN Ligands: CA |
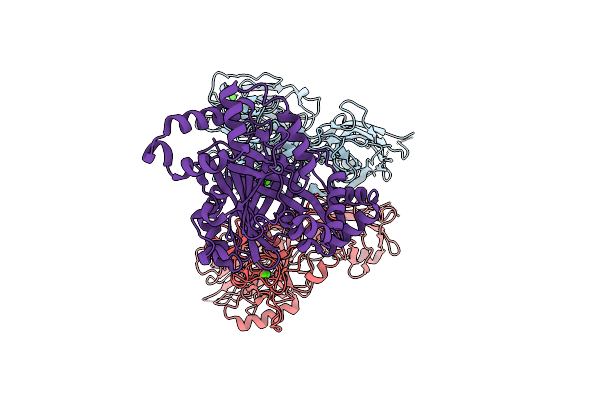 |
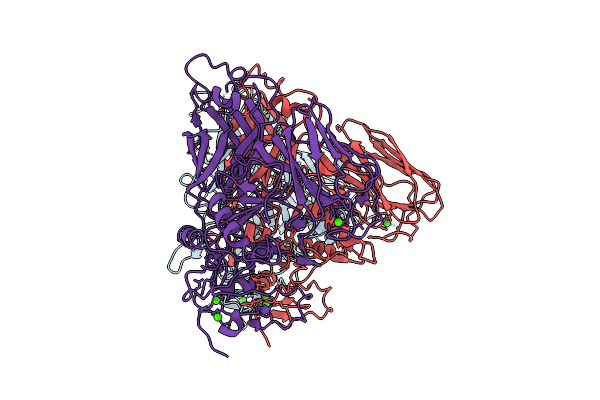
Clostridioides Difficile Transferase B Component Dimer In Complex With The A Component
Organism: Clostridioides difficile r20291
Method: ELECTRON MICROSCOPY Release Date: 2025-07-16 Classification: TOXIN Ligands: CA |
 |
Organism: Clostridioides difficile r20291
Method: ELECTRON MICROSCOPY Resolution:4.19 Å Release Date: 2025-07-16 Classification: TOXIN Ligands: CA |
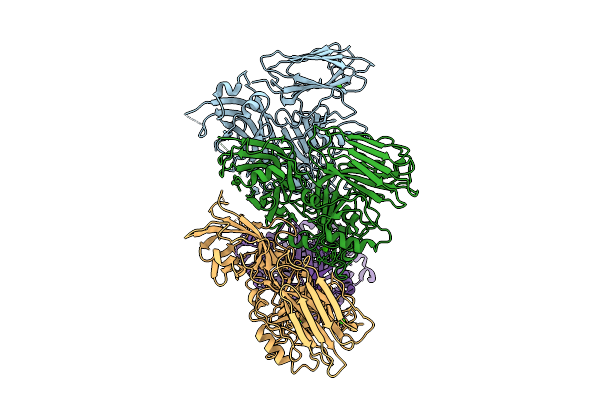 |
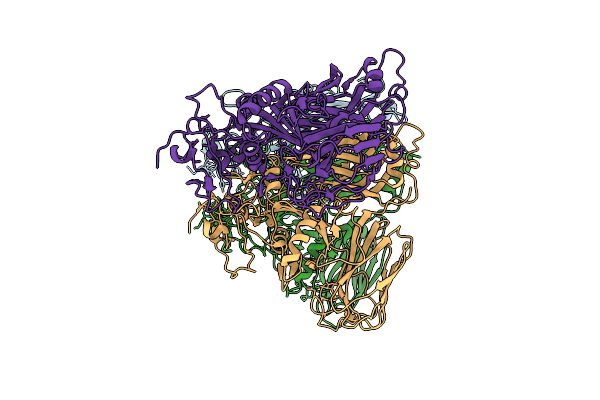
Clostridioides Difficile Transferase B Component Trimer In Complex With The A Component
Organism: Clostridioides difficile r20291
Method: ELECTRON MICROSCOPY Resolution:7.86 Å Release Date: 2025-07-16 Classification: TOXIN Ligands: CA |
 |
Organism: Clostridioides difficile r20291
Method: ELECTRON MICROSCOPY Resolution:7.01 Å Release Date: 2025-07-16 Classification: TOXIN Ligands: CA |
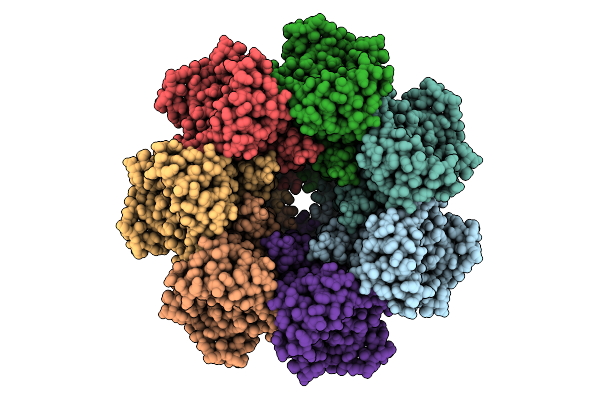 |
Organism: Clostridioides difficile r20291
Method: ELECTRON MICROSCOPY Resolution:3.33 Å Release Date: 2025-07-16 Classification: TOXIN Ligands: CA |
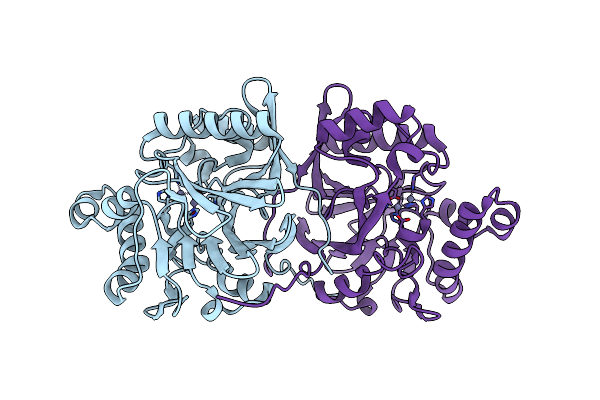 |
Crystal Structure Of Methyl Parathion Hydrolase Mutant A127L/I145L/Q272D/S279A/S304P
Organism: Pseudomonas sp. wbc-3
Method: X-RAY DIFFRACTION Resolution:1.39 Å Release Date: 2025-06-11 Classification: HYDROLASE Ligands: ZN |
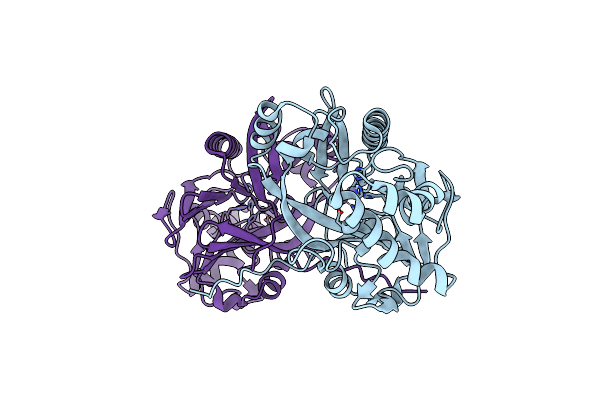 |
Crystal Structure Of Methyl Parathion Hydrolase Mutant A66D/I143V/I145L/Q272D/S279A/
Organism: Pseudomonas sp. wbc-3
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2025-06-11 Classification: HYDROLASE Ligands: ZN |
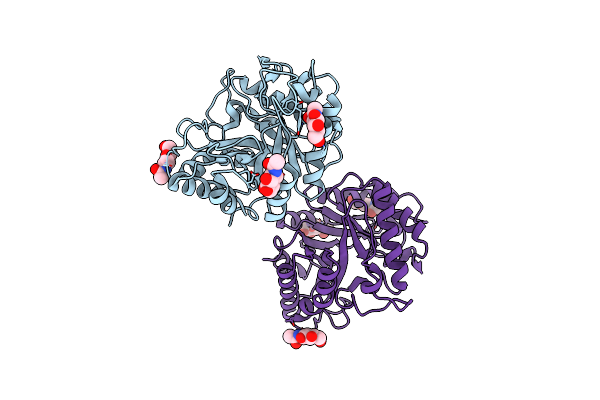 |
Organism: Bispora sp. mey-1
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2025-05-28 Classification: HYDROLASE Ligands: NAG |
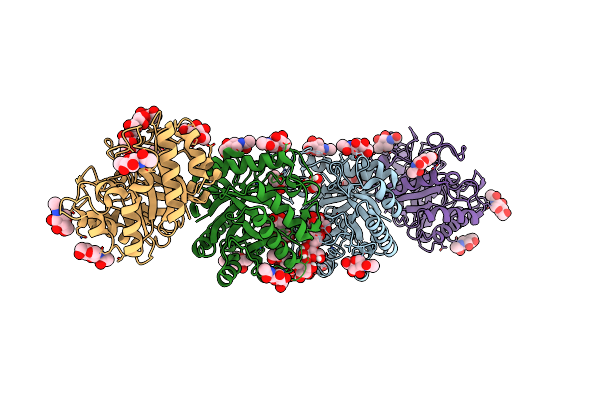 |
Crystal Structure Of A Beta-1,4-Endoglucanase From Bispora Sp. Mey-1 In Complex With Mannotetrose
Organism: Bispora sp. mey-1
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2025-05-21 Classification: HYDROLASE Ligands: NAG, PEG |
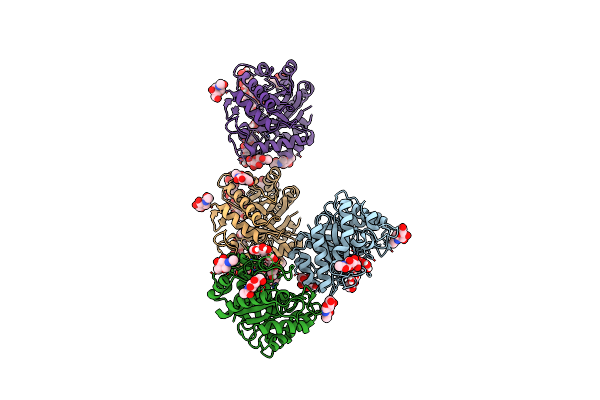 |
Crystal Structure Of A Beta-1,4-Endoglucanase From Bispora Sp. Mey-1 In Complex With Cellotetraose
Organism: Bispora sp. mey-1
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2025-05-21 Classification: HYDROLASE Ligands: NAG |
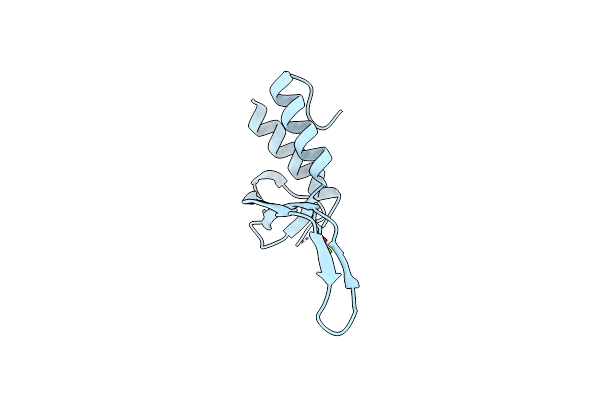 |
Organism: Mycobacterium phage phaedrus
Method: X-RAY DIFFRACTION Resolution:1.21 Å Release Date: 2024-12-18 Classification: VIRAL PROTEIN Ligands: MG |
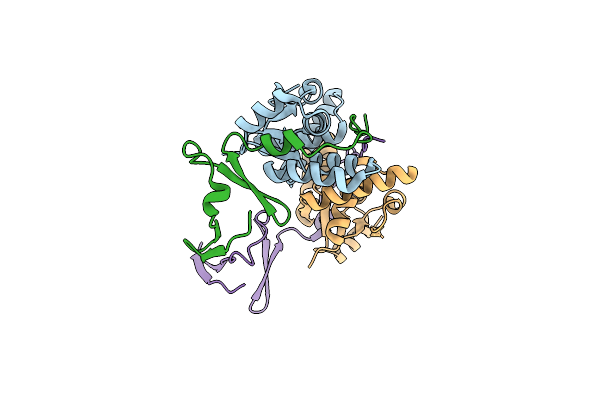 |
Organism: Escherichia coli kly
Method: X-RAY DIFFRACTION Resolution:2.86 Å Release Date: 2024-12-18 Classification: TOXIN |
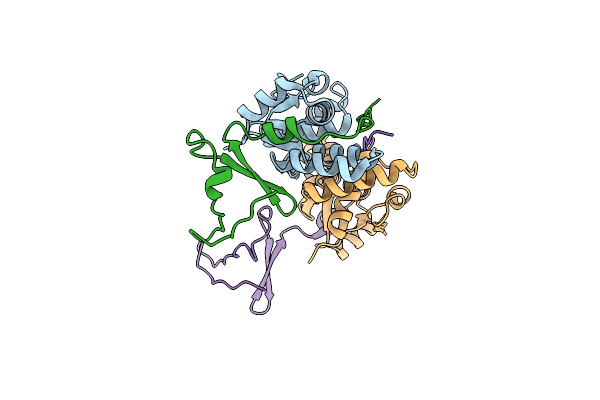 |
Crystal Structure Of The E. Coli F-Plasmid Vapbc Toxin-Antitoxin Complex (Vapb T3N, A13P, L16R)
Organism: Escherichia coli kly
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2024-12-18 Classification: TOXIN |
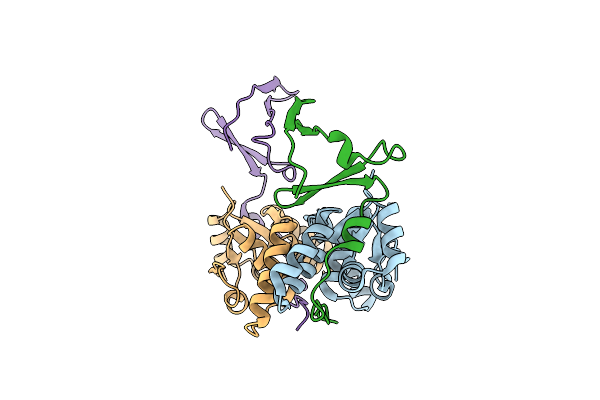 |
Crystal Structure Of The E. Coli F-Plasmid Vapbc Toxin-Antitoxin Complex (Vapb T3N)
Organism: Escherichia coli kly
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2024-12-18 Classification: TOXIN |
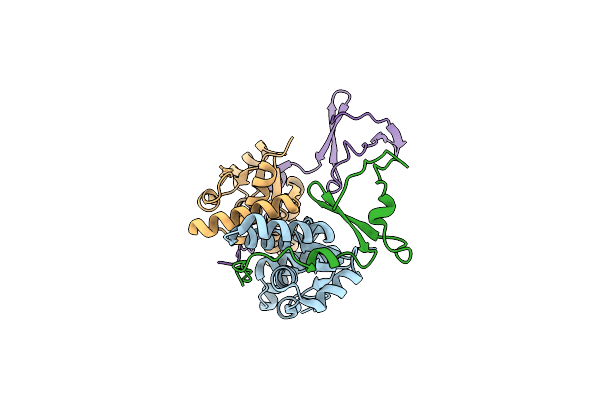 |
Crystal Structure Of The E. Coli F-Plasmid Vapbc Toxin-Antitoxin Complex (Vapb V5E)
Organism: Escherichia coli kly
Method: X-RAY DIFFRACTION Resolution:3.15 Å Release Date: 2024-12-18 Classification: TOXIN |

