Search Count: 711
All
Selected
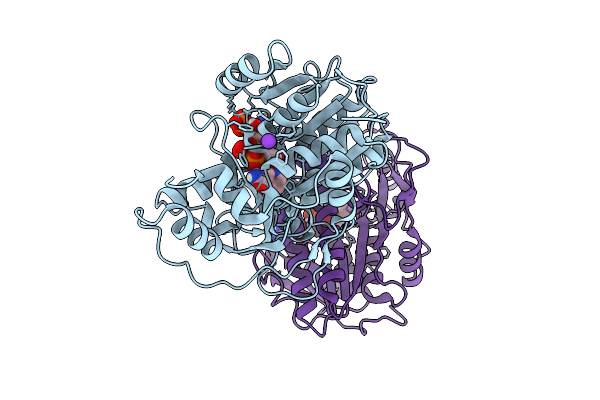 |
Ketoreductase Engineering For A Chemoenzymatic Fluorination And Dynamic Kinetic Reduction Cascade
Organism: Sporobolomyces salmonicolor
Method: X-RAY DIFFRACTION Resolution:1.48 Å Release Date: 2025-07-30 Classification: OXIDOREDUCTASE Ligands: NAP, K |
 |
Organism: Hansschlegelia zhihuaiae
Method: X-RAY DIFFRACTION Resolution:1.54 Å Release Date: 2024-05-01 Classification: HYDROLASE Ligands: 1SM, TLA, GOL |
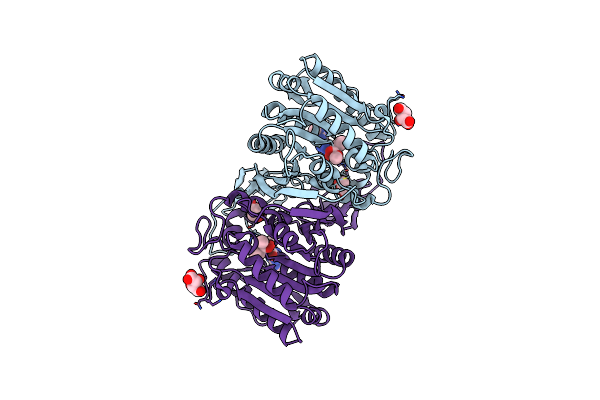 |
Organism: Hansschlegelia zhihuaiae
Method: X-RAY DIFFRACTION Resolution:1.59 Å Release Date: 2024-05-01 Classification: HYDROLASE Ligands: RXF, TLA, GOL |
 |
Organism: Hansschlegelia zhihuaiae
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2024-04-10 Classification: HYDROLASE Ligands: R4O, TLA, GOL |
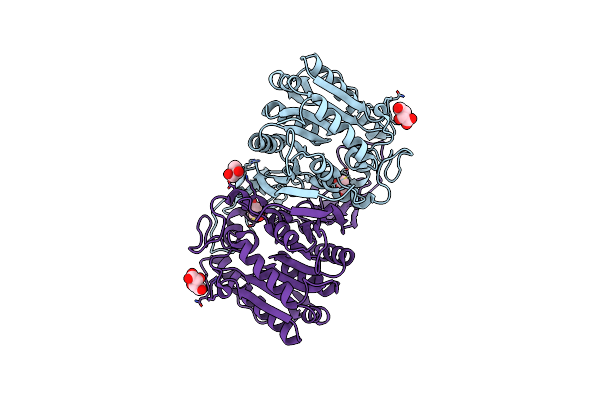 |
Organism: Hansschlegelia zhihuaiae
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2024-04-10 Classification: HYDROLASE Ligands: TLA, GOL |
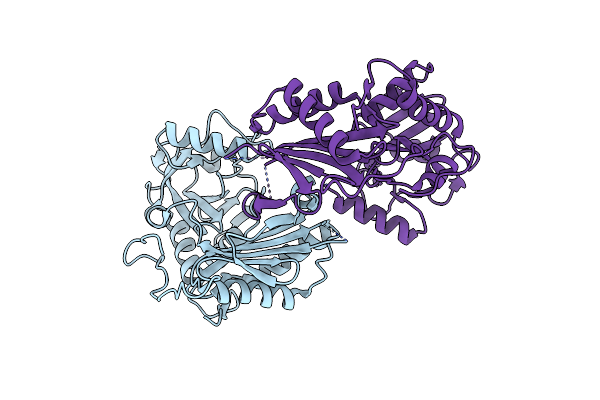 |
Organism: Mycolicibacterium vanbaalenii (strain dsm 7251 / jcm 13017 / bcrc 16820 / kctc 9966 / nrrl b-24157 / pyr-1)
Method: X-RAY DIFFRACTION Resolution:2.33 Å Release Date: 2024-01-24 Classification: TRANSFERASE |
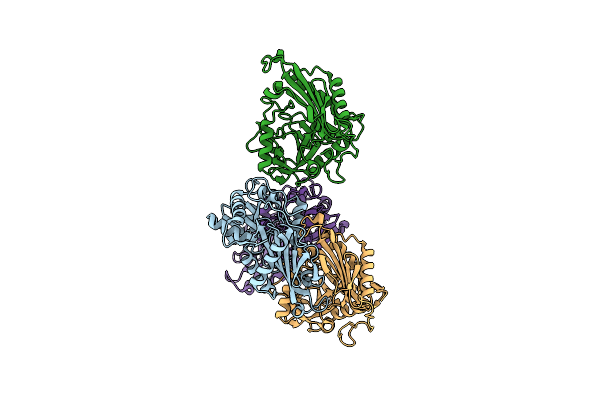 |
Organism: Mycolicibacterium vanbaalenii (strain dsm 7251 / jcm 13017 / bcrc 16820 / kctc 9966 / nrrl b-24157 / pyr-1)
Method: X-RAY DIFFRACTION Resolution:2.27 Å Release Date: 2024-01-24 Classification: TRANSFERASE |
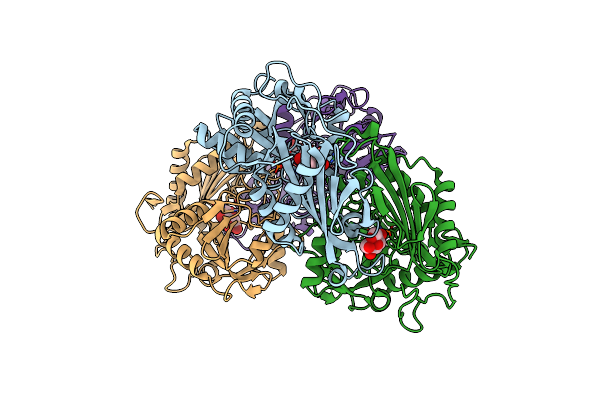 |
Crystal Structure Of Mv In Complex With Llp And Fru From Mycobacterium Vanbaalenii
Organism: Mycolicibacterium vanbaalenii (strain dsm 7251 / jcm 13017 / bcrc 16820 / kctc 9966 / nrrl b-24157 / pyr-1)
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2024-01-24 Classification: TRANSFERASE Ligands: FUD |
 |
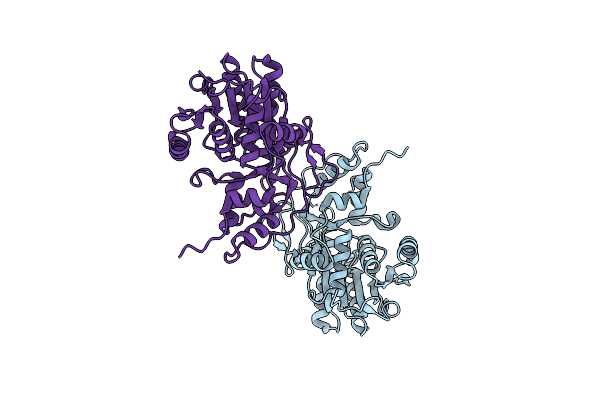
Crystal Structure Of Carbonyl Reductase Sscr Mutant 1 From Sporobolomyces Salmonicolor
Organism: Sporobolomyces salmonicolor
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2023-12-27 Classification: OXIDOREDUCTASE |
 |
Crystal Structure Of A Carbonyl Reductase Sscr Mutant From Sporobolomyces Salmonicolor
Organism: Sporobolomyces salmonicolor
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2023-12-27 Classification: OXIDOREDUCTASE |
 |
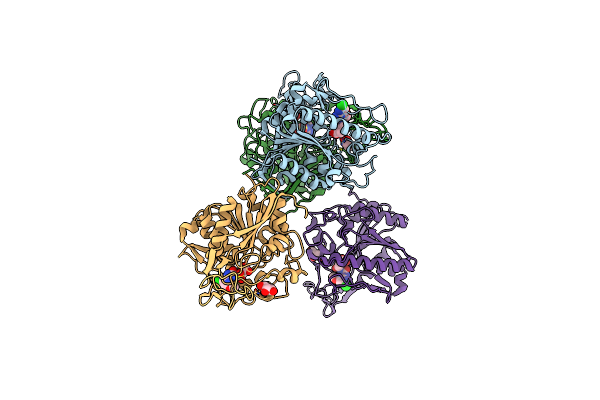
Organism: Hansschlegelia zhihuaiae
Method: X-RAY DIFFRACTION Resolution:1.29 Å Release Date: 2023-08-02 Classification: HYDROLASE Ligands: 1MM, GOL |
 |
Organism: Hansschlegelia zhihuaiae
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2023-08-02 Classification: HYDROLASE Ligands: IJC, GOL |
 |
Organism: Hansschlegelia zhihuaiae
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2023-08-02 Classification: HYDROLASE Ligands: GOL, JV6 |
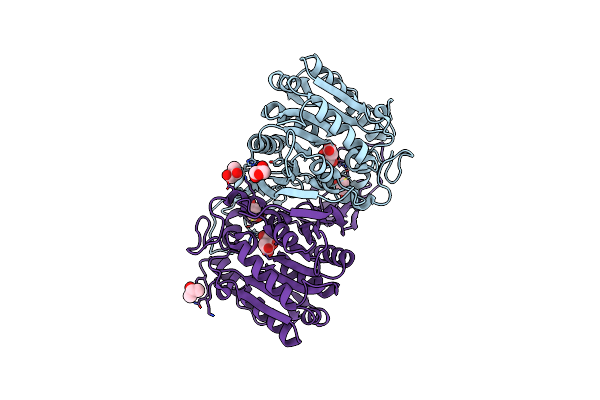 |
Organism: Hansschlegelia zhihuaiae
Method: X-RAY DIFFRACTION Resolution:1.46 Å Release Date: 2023-08-02 Classification: HYDROLASE Ligands: GOL, CIT |
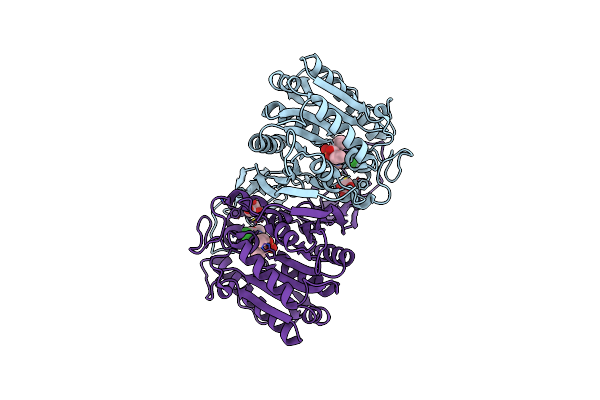 |
Organism: Hansschlegelia zhihuaiae
Method: X-RAY DIFFRACTION Resolution:1.44 Å Release Date: 2023-08-02 Classification: HYDROLASE Ligands: CIE, GOL |
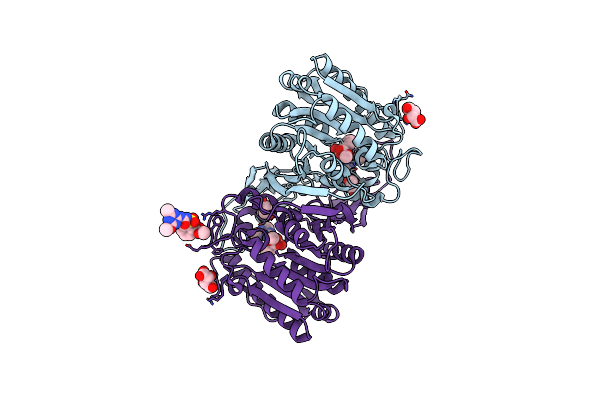 |
Organism: Hansschlegelia zhihuaiae
Method: X-RAY DIFFRACTION Resolution:1.32 Å Release Date: 2023-08-02 Classification: HYDROLASE Ligands: 1MM, GOL, TLA |
 |
Organism: Hansschlegelia zhihuaiae
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2023-08-02 Classification: HYDROLASE Ligands: 1MM, TLA, GOL |
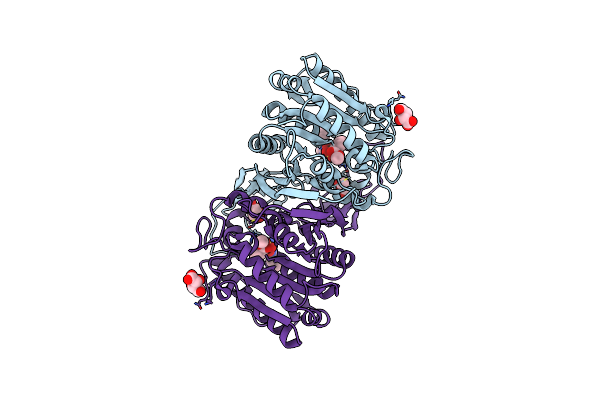 |
Organism: Hansschlegelia zhihuaiae
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2023-08-02 Classification: HYDROLASE Ligands: RXF, TLA, GOL |
 |
Organism: Hansschlegelia zhihuaiae
Method: X-RAY DIFFRACTION Resolution:1.42 Å Release Date: 2023-08-02 Classification: HYDROLASE Ligands: 1TB, TLA, GOL |
 |
Organism: Hansschlegelia zhihuaiae
Method: X-RAY DIFFRACTION Resolution:1.56 Å Release Date: 2023-08-02 Classification: HYDROLASE Ligands: CIE, TLA, GOL |

