Search Count: 264
All
Selected
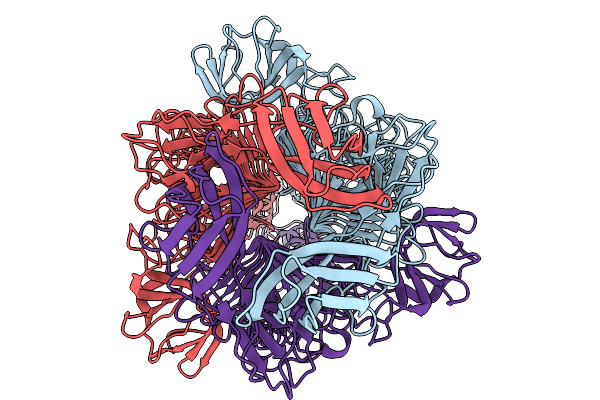 |
Organism: Klebsiella phage k64-1
Method: ELECTRON MICROSCOPY Resolution:2.24 Å Release Date: 2026-03-25 Classification: ANTIMICROBIAL PROTEIN |
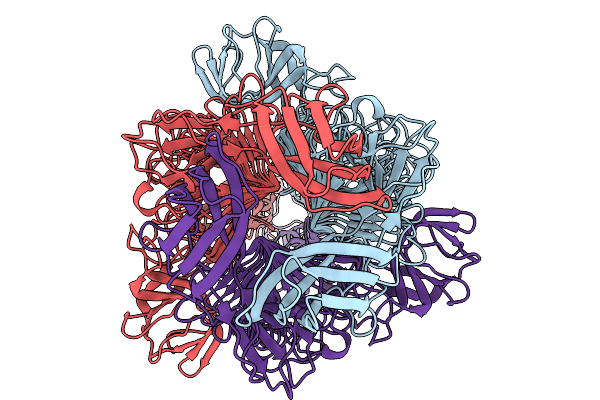 |
Organism: Klebsiella phage k64-1
Method: ELECTRON MICROSCOPY Resolution:2.18 Å Release Date: 2026-03-25 Classification: ANTIMICROBIAL PROTEIN |
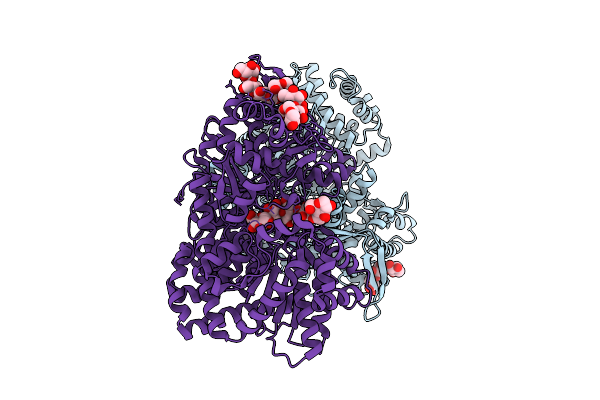 |
Carbohydrate-Bound Structure Of Alpha-Glucan Phosphorylase From Crocosphaera Subtropica Atcc 51142
Organism: Crocosphaera subtropica atcc 51142
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-03-18 Classification: TRANSFERASE |
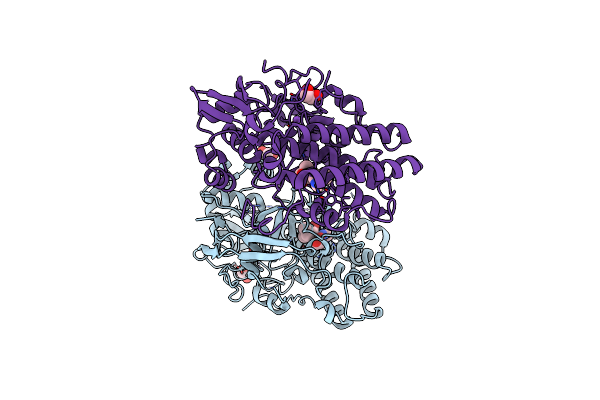 |
Egtb-Iv From Crocosphaera Subtropica, An Ergothioneine-Biosynthetic Type Iv Sulfoxide Synthase In Complex With Hercynine
Organism: Crocosphaera subtropica atcc 51142
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2024-08-07 Classification: OXIDOREDUCTASE Ligands: CL, NA, FE, AVJ, EDO |
 |
Egtb-Iv From Crocosphaera Subtropica, An Ergothioneine-Biosynthetic Type Iv Sulfoxide Synthase In Complex With Cysteine And Hercynine
Organism: Crocosphaera subtropica atcc 51142
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2024-08-07 Classification: OXIDOREDUCTASE |
 |
Egtb-Iv From Crocosphaera Subtropica, A Type Iv Sulfoxide Synthase Involved In Ergothioneine Biosynthesis
Organism: Crocosphaera subtropica atcc 51142
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2024-08-07 Classification: OXIDOREDUCTASE |
 |
Egtb-Iv From Crocosphaera Subtropica, An Ergothioneine-Biosynthetic Type Iv Sulfoxide Synthase In Complex With N,N-Dimethyl-Histidine
Organism: Crocosphaera subtropica atcc 51142
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2024-08-07 Classification: OXIDOREDUCTASE Ligands: CL, NA, FE, EDO, AVI |
 |
Ligand Free Structure Of Branching Enzyme Isoform 3 (Be3) From Crocosphaera Subtropica Atcc 51142
Organism: Crocosphaera subtropica atcc 51142
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2023-06-07 Classification: TRANSFERASE Ligands: GOL |
 |
Crystal Structure Of Branching Enzyme D434A Mutant From Cyanothece Sp. Atcc 51142
Organism: Crocosphaera subtropica atcc 51142
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2020-08-05 Classification: TRANSFERASE Ligands: MG, GOL |
 |
Crystal Structure Of Branching Enzyme From Cyanothece Sp. Atcc 51142 In Complex With Maltoheptaose
Organism: Cyanothece sp. (strain atcc 51142)
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2017-08-16 Classification: TRANSFERASE Ligands: GOL, MG |
 |
Crystal Structure Of Branching Enzyme Y500A Mutant From Cyanothece Sp. Atcc 51142
Organism: Cyanothece sp. (strain atcc 51142)
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2017-08-16 Classification: TRANSFERASE Ligands: GOL, MG |
 |
Crystal Structure Of Branching Enzyme D501A Mutant From Cyanothece Sp. Atcc 51142
Organism: Cyanothece sp. (strain atcc 51142)
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2017-08-16 Classification: TRANSFERASE Ligands: GOL, MG |
 |
Crystal Structure Of Branching Enzyme Y500A/D501A Mutant From Cyanothece Sp. Atcc 51142 In Complex With Maltoheptaose
Organism: Cyanothece sp. (strain atcc 51142)
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2017-08-16 Classification: TRANSFERASE Ligands: GOL, MG |
 |
Crystal Structure Of Branching Enzyme L541A Mutant From Cyanothece Sp. Atcc 51142
Organism: Cyanothece sp. (strain atcc 51142)
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2017-08-16 Classification: TRANSFERASE Ligands: GOL, MG |
 |
Crystal Structure Of Branching Enzyme L541A/W655A Mutant From Cyanothece Sp. Atcc 51142
Organism: Cyanothece sp. (strain atcc 51142)
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2017-08-16 Classification: TRANSFERASE Ligands: GOL, MG |
 |
Crystal Structure Of Branching Enzyme L541A Mutant From Cyanothece Sp. Atcc 51142 In Complex With Maltoheptaose
Organism: Cyanothece sp. (strain atcc 51142)
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2017-08-16 Classification: TRANSFERASE Ligands: GOL, MG |
 |
Crystal Structure Of Branching Enzyme W610A Mutant From Cyanothece Sp. Atcc 51142
Organism: Cyanothece sp. (strain atcc 51142)
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2017-08-16 Classification: TRANSFERASE Ligands: GOL, MG |
 |
Crystal Structure Of Branching Enzyme Y500A/D501A Double Mutant From Cyanothece Sp. Atcc 51142
Organism: Cyanothece sp. (strain atcc 51142)
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2017-08-16 Classification: TRANSFERASE Ligands: GOL, MG |
 |
Organism: Cyanothece sp. (strain atcc 51142)
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2017-02-22 Classification: TRANSFERASE Ligands: GOL, MG |
 |
Crystal Structure Of Branching Enzyme From Cyanothece Sp. Atcc 51142 In Complex With Maltohexaose
Organism: Cyanothece sp. (strain atcc 51142)
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2017-02-22 Classification: TRANSFERASE Ligands: GOL, MG |

