Search Count: 279
All
Selected
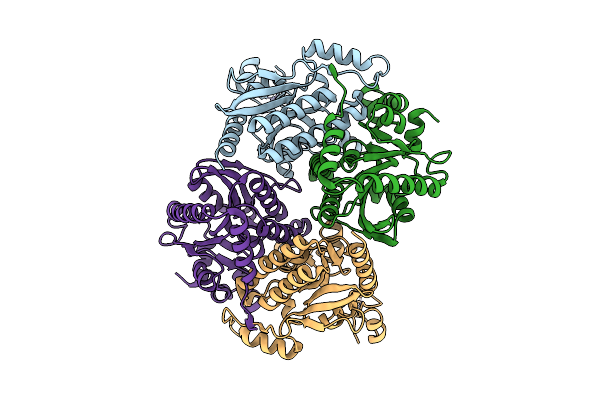 |
Crystal Structure Of The Carboxyltransferase Subunit Of Acetyl-Coa Carboxylase From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus j-10-fl
Method: X-RAY DIFFRACTION Resolution:2.78 Å Release Date: 2026-03-11 Classification: LIGASE Ligands: ZN |
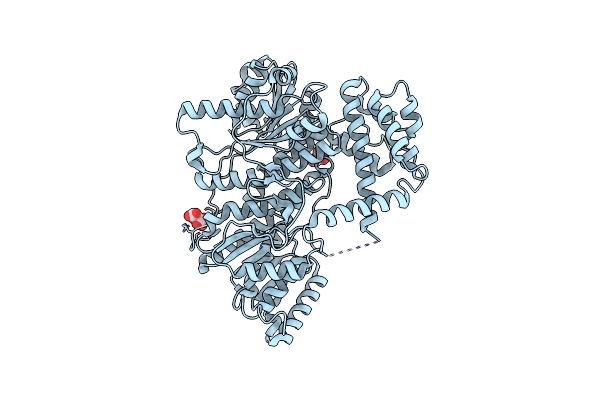 |
Structures Of The C-Terminal Subenzyme Of The Malonyl-Coa Reductase From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus (strain atcc 29366 / dsm 635 / j-10-fl)
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2024-04-17 Classification: OXIDOREDUCTASE Ligands: TLA, GOL |
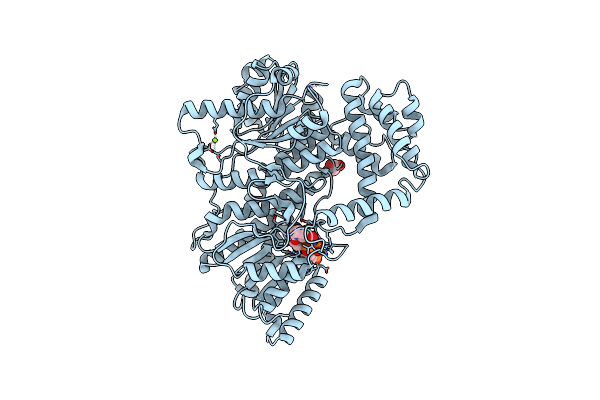 |
Structure Of The C-Terminal Subenzyme Of The Malonyl-Coa Reductase From Chloroflexus Aurantiacus, In Complex With Nadp+
Organism: Chloroflexus aurantiacus (strain atcc 29366 / dsm 635 / j-10-fl)
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2024-04-17 Classification: OXIDOREDUCTASE Ligands: NAP, GOL, MG |
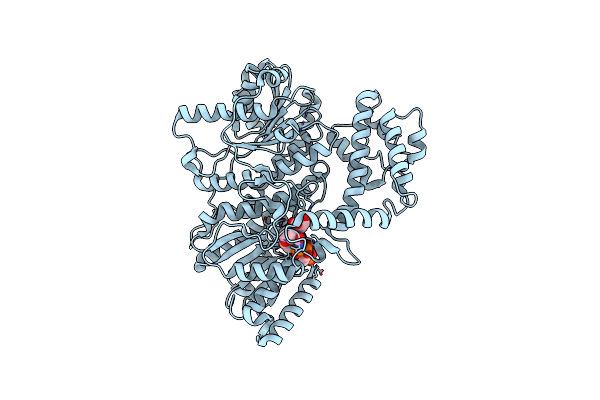 |
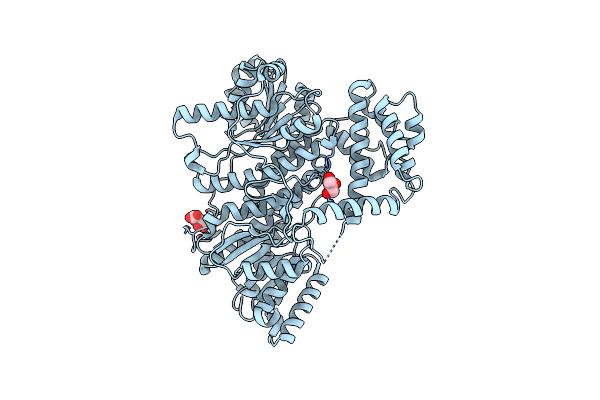
Structure Of The C-Terminal Subenzyme Of The Malonyl-Coa Reductase From Chloroflexus Aurantiacus, In Complex With Nadp+ And Malonate
Organism: Chloroflexus aurantiacus (strain atcc 29366 / dsm 635 / j-10-fl)
Method: X-RAY DIFFRACTION Resolution:2.13 Å Release Date: 2024-04-17 Classification: OXIDOREDUCTASE Ligands: NAP, MLA |
 |
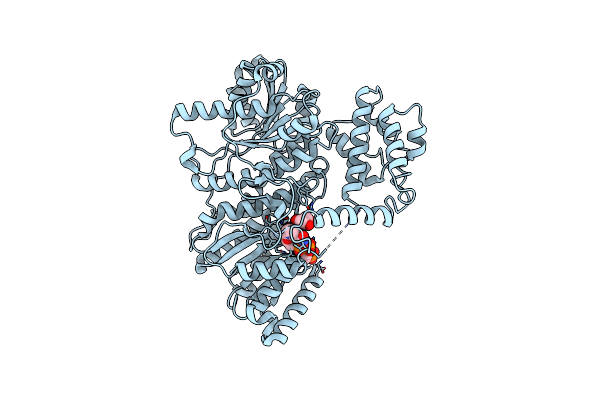
Structure Of The C-Terminal Subenzyme Of The Malonyl-Coa Reductase From Chloroflexus Aurantiacus, Mutant N940V/K1106W/S1114R
Organism: Chloroflexus aurantiacus (strain atcc 29366 / dsm 635 / j-10-fl)
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2024-04-17 Classification: OXIDOREDUCTASE Ligands: TLA, GOL |
 |
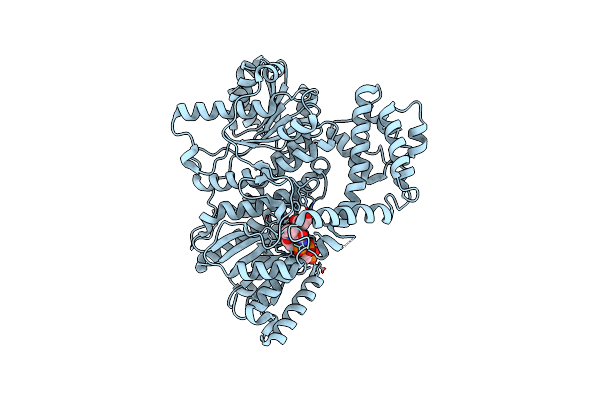
Structure Of The C-Terminal Subenzyme Of The Malonyl-Coa Reductase From Chloroflexus Aurantiacus, Mutant N940V/K1106W/S1114R In Complex With Nadp+
Organism: Chloroflexus aurantiacus (strain atcc 29366 / dsm 635 / j-10-fl)
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2024-04-17 Classification: OXIDOREDUCTASE Ligands: NAP, GOL |
 |
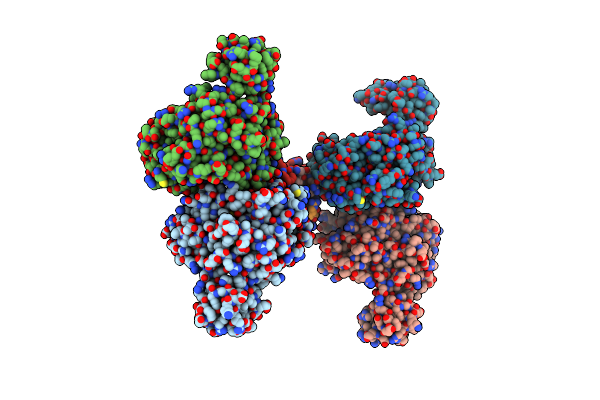
Structure Of The C-Terminal Subenzyme Of The Malonyl-Coa Reductase From Chloroflexus Aurantiacus, Mutant N940V/K1106W/S1114R In Complex With Nadp+ And Malonate
Organism: Chloroflexus aurantiacus (strain atcc 29366 / dsm 635 / j-10-fl)
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2024-04-17 Classification: OXIDOREDUCTASE Ligands: NAP, MLA |
 |
The Tetrameric Structure Of Biotin Carboxylase From Chloroflexus Aurantiacus In Complex With Bicarbonate
Organism: Chloroflexus aurantiacus (strain atcc 29366 / dsm 635 / j-10-fl)
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2024-01-10 Classification: LIGASE |
 |
The Homodimer Of A Biotin Carboxylase Isoform From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus (strain atcc 29366 / dsm 635 / j-10-fl)
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2024-01-10 Classification: LIGASE |
 |
Organism: Chloroflexus aurantiacus j-10-fl
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2023-04-05 Classification: OXIDOREDUCTASE Ligands: MG |
 |
Malonyl-Coa Reductase From Chloroflexus Aurantiacus - C-Terminal Nadp Bound
Organism: Chloroflexus aurantiacus j-10-fl
Method: X-RAY DIFFRACTION Resolution:1.54 Å Release Date: 2023-04-05 Classification: OXIDOREDUCTASE Ligands: NAP, MG |
 |
Malonyl-Coa Reductase From Chloroflexus Aurantiacus - C-Terminal Nadp And Malonate Bound
Organism: Chloroflexus aurantiacus j-10-fl
Method: X-RAY DIFFRACTION Resolution:2.11 Å Release Date: 2023-04-05 Classification: OXIDOREDUCTASE Ligands: NAP, MLI |
 |
Malonyl-Coa Reductase From Chloroflexus Aurantiacus - C-Terminal R773A Variant
Organism: Chloroflexus aurantiacus j-10-fl
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2023-04-05 Classification: OXIDOREDUCTASE Ligands: SO4 |
 |
Malonyl-Coa Reductase From Chloroflexus Aurantiacus - C-Terminal R773Q Variant
Organism: Chloroflexus aurantiacus j-10-fl
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2023-04-05 Classification: OXIDOREDUCTASE |
 |
Malonyl-Coa Reductase From Chloroflexus Aurantiacus - C-Terminal Y731A Variant
Organism: Chloroflexus aurantiacus j-10-fl
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2023-04-05 Classification: OXIDOREDUCTASE Ligands: MG |
 |
Malonyl-Coa Reductase From Chloroflexus Aurantiacus - C-Terminal E779W Variant
Organism: Chloroflexus aurantiacus j-10-fl
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-04-05 Classification: OXIDOREDUCTASE Ligands: SO4 |
 |
Organism: Chloroflexus aurantiacus j-10-fl
Method: X-RAY DIFFRACTION Resolution:2.72 Å Release Date: 2023-04-05 Classification: OXIDOREDUCTASE |
 |
Organism: Chloroflexus aurantiacus j-10-fl
Method: X-RAY DIFFRACTION Resolution:2.49 Å Release Date: 2023-01-11 Classification: TRANSFERASE Ligands: OA9, COA, MEZ |
 |
Organism: Chloroflexus aurantiacus j-10-fl
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2022-12-28 Classification: TRANSFERASE Ligands: CL |
 |
Crystal Structure Of Isocitrate Lyase (Caur_3889) From Chloroflexus Aurantiacus In Complex With Manganese Ion
Organism: Chloroflexus aurantiacus (strain atcc 29366 / dsm 635 / j-10-fl)
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2021-07-28 Classification: LYASE Ligands: GOL, P3G |

