Search Count: 2,649
All
Selected
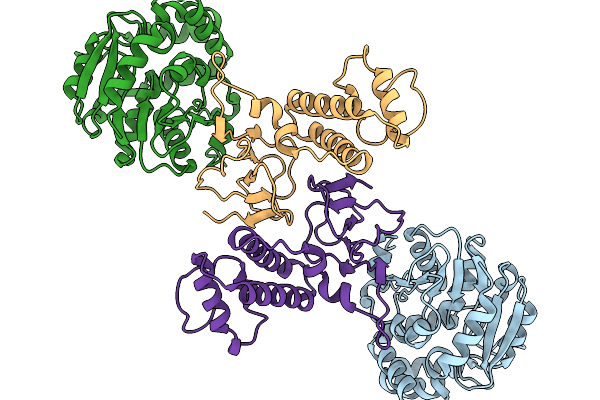 |
Organism: Oryza sativa japonica group, Tenuivirus oryzabrevis
Method: ELECTRON MICROSCOPY Resolution:3.67 Å Release Date: 2026-03-04 Classification: PLANT PROTEIN |
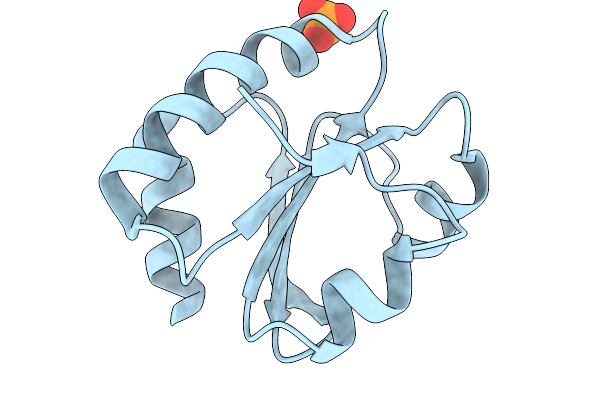 |
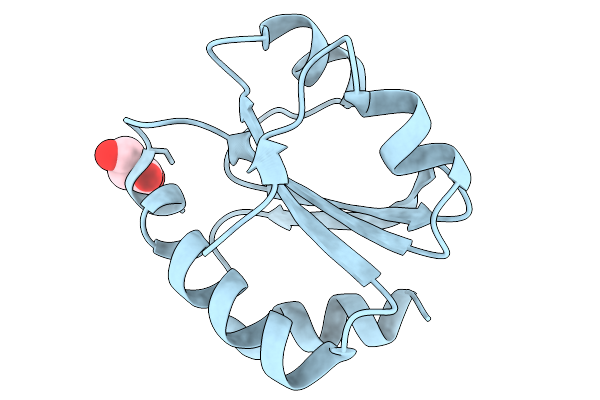
Organism: Treponema pallidum subsp. pallidum str. nichols
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2026-02-11 Classification: STRUCTURAL PROTEIN Ligands: PO4, CL |
 |
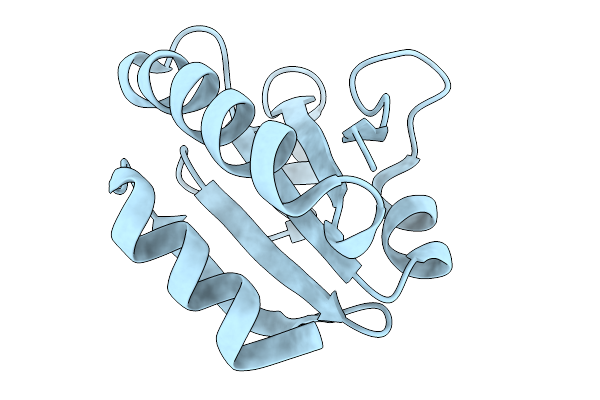
Organism: Treponema pallidum subsp. pallidum str. nichols
Method: X-RAY DIFFRACTION Resolution:2.64 Å Release Date: 2026-02-11 Classification: STRUCTURAL PROTEIN Ligands: 3OH |
 |
Organism: Treponema pallidum subsp. pallidum str. nichols
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-02-11 Classification: STRUCTURAL PROTEIN |
 |
Cryo-Em Structure Of Vsv-Indiana (Mudd-Summers Strain) Glycoprotein Under Its Acidic Conformation
Organism: Vesicular stomatitis virus
Method: ELECTRON MICROSCOPY Release Date: 2025-10-22 Classification: VIRAL PROTEIN |
 |
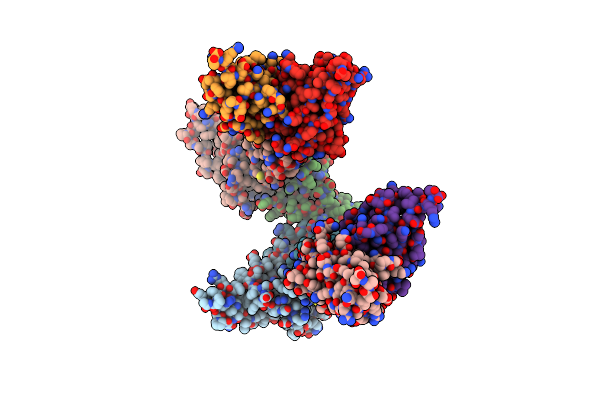
Organism: Hepatitis b virus genotype c, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-11-13 Classification: VIRUS LIKE PARTICLE |
 |
Organism: Hepatitis b virus genotype c, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-11-13 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
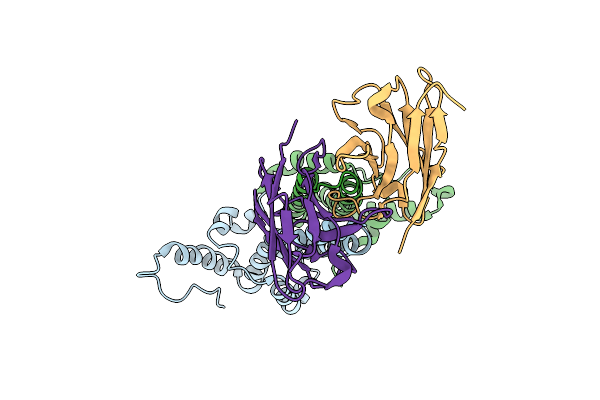 |
Organism: Homo sapiens, Hepatitis b virus genotype c
Method: ELECTRON MICROSCOPY Release Date: 2024-11-13 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
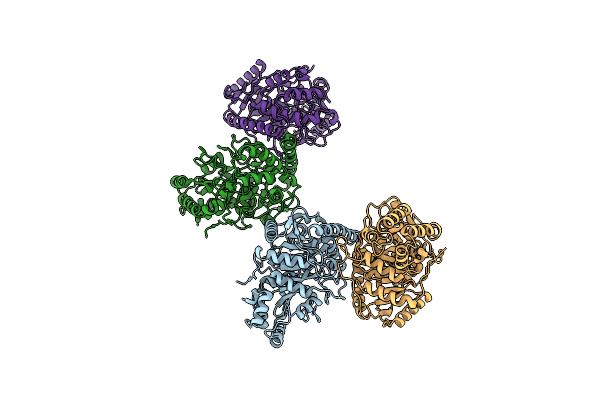 |
Organism: Pseudonocardia acaciae
Method: X-RAY DIFFRACTION Resolution:3.02 Å Release Date: 2024-10-09 Classification: HYDROLASE |
 |
Organism: Zymoseptoria tritici st99ch_1e4
Method: SOLUTION NMR Release Date: 2024-10-09 Classification: UNKNOWN FUNCTION |
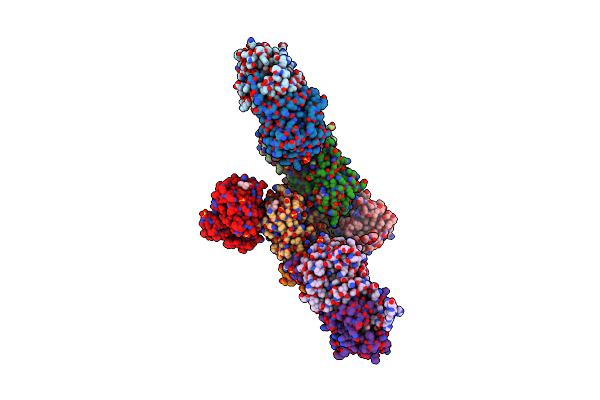 |
Dihydrofolate Reductase-Like Enzyme From Leptospira Interrogans (Selenomethionine Derivative)
Organism: Leptospira interrogans serovar pomona
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2024-06-26 Classification: OXIDOREDUCTASE Ligands: 1PE, NAP, SO4 |
 |
Organism: Leptospira interrogans serovar pomona
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2024-06-26 Classification: OXIDOREDUCTASE Ligands: 1PE, NAP, SO4, ACT |
 |
Dihydrofolate Reductase-Like Enzyme From Leptospira Interrogans With Additional Nadp+
Organism: Leptospira interrogans serovar pomona
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2024-06-26 Classification: OXIDOREDUCTASE Ligands: 1PE, NAP, SO4, ACT |
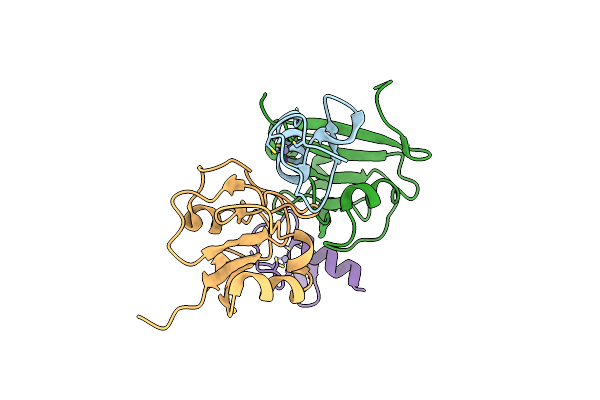 |
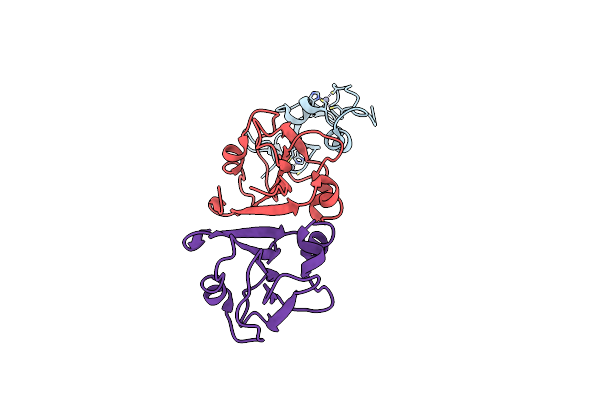
Organism: Arabidopsis thaliana, Onion yellows phytoplasma oy-m
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2024-02-28 Classification: PLANT PROTEIN Ligands: ZN |
 |
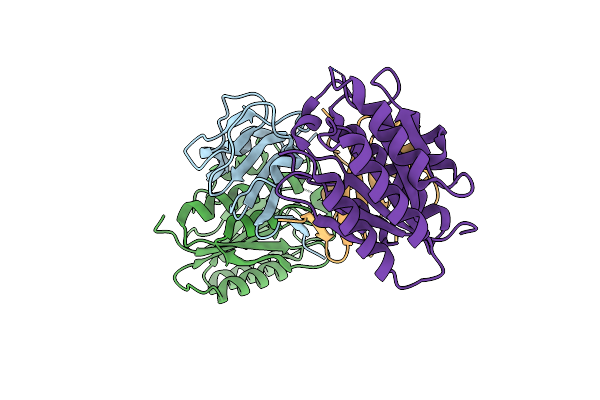
Organism: Arabidopsis thaliana, Onion yellows phytoplasma oy-m
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2024-02-28 Classification: PLANT PROTEIN Ligands: ZN |
 |
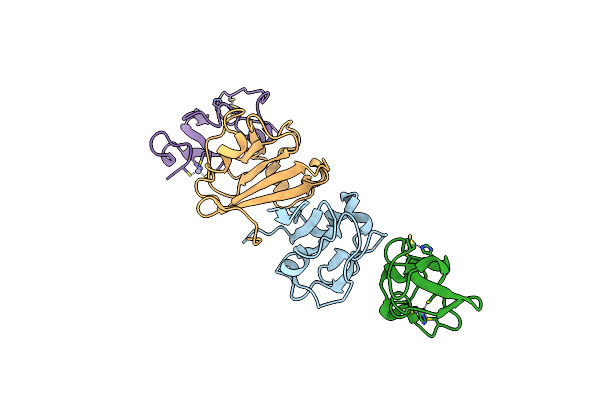
Organism: Onion yellows phytoplasma oy-m, Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2024-02-28 Classification: PLANT PROTEIN |
 |
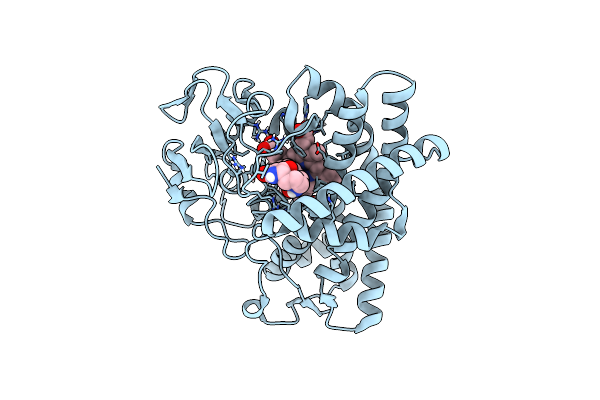
Organism: Onion yellows phytoplasma oy-m, Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2024-02-28 Classification: PLANT PROTEIN Ligands: ZN |
 |
Crystal Structure Of Cytochrome P450 Cfta From Streptomyces Torulosus Nrrl B-3889, In Complex With The Substrate Ikarugamycin
Organism: Streptomyces torulosus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-11-15 Classification: OXIDOREDUCTASE Ligands: EIA, HEM |
 |
Crystal Structure Of Cytochrome P450 Cfta From Streptomyces Torulosus Nrrl B-3889, In Complex With A Substrate Compound C
Organism: Streptomyces torulosus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-11-15 Classification: OXIDOREDUCTASE Ligands: EIU, HEM, NA |
 |
Structure Of M. Baixiangningiae Darr-Dna Complex Reveals Novel Dimer-Of-Dimers Dna Binding
Organism: Mycolicibacterium baixiangningiae
Method: X-RAY DIFFRACTION Resolution:3.49 Å Release Date: 2023-11-01 Classification: TRANSCRIPTION/DNA |

