Search Count: 53
All
Selected
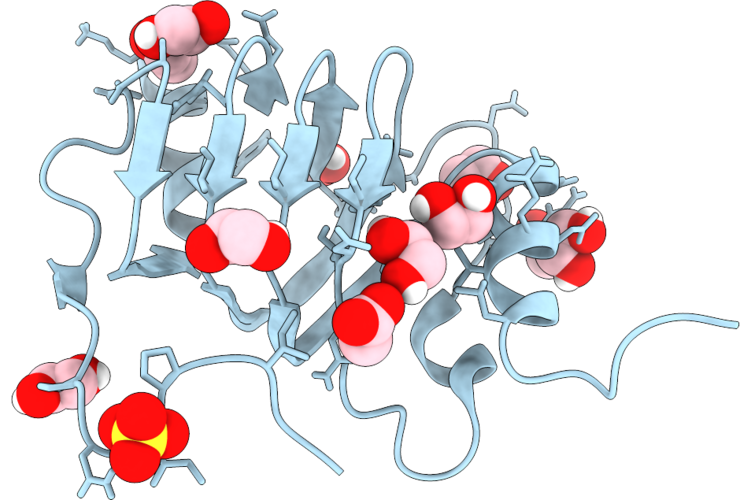 |
Organism: Brevundimonas sp. g8
Method: X-RAY DIFFRACTION Resolution:1.22 Å Release Date: 2026-04-22 Classification: UNKNOWN FUNCTION Ligands: SO4, ACT, EDO |
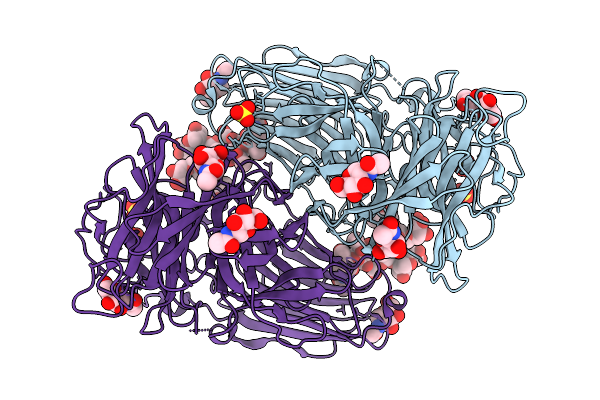 |
Organism: Komagataella phaffii cbs 7435
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: SO4, NAG |
 |
Structure Of D81A-Fructofuranosidase From Purpureocilum Lilacinum In Complex With Raffinose
Organism: Komagataella phaffii cbs 7435
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: FRU, TRS, NAG, GLC, GLA, SO4 |
 |
Structure Of D81A-Fructofuranosidase From Purpureocilum Lilacinum In Complex With Fructose
Organism: Komagataella phaffii cbs 7435
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: FRU, NAG, SO4 |
 |
Structure Of D81A-Fructofuranosidase From Purpureocilum Lilacinum In Complex With Sucrose
Organism: Komagataella phaffii cbs 7435
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: GLC, FRU, SO4, TRS, NAG, GOL |
 |
Organism: Enterococcus
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2024-11-27 Classification: HYDROLASE Ligands: ZN, Y7U |
 |
Organism: Enterococcus
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2024-11-27 Classification: HYDROLASE Ligands: ZN |
 |
Organism: Enterococcus
Method: X-RAY DIFFRACTION Resolution:1.56 Å Release Date: 2024-11-27 Classification: HYDROLASE Ligands: CAC, ZN |
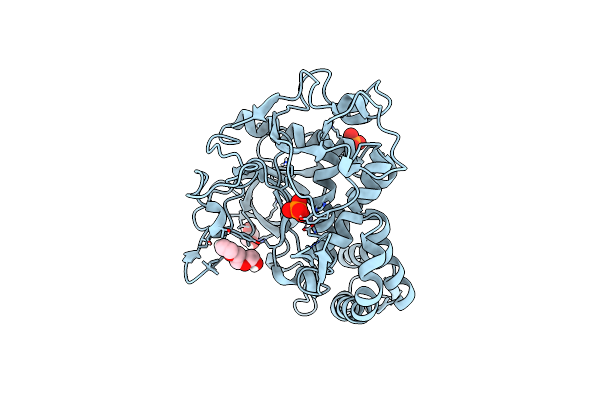 |
Crystal Structure Of The Catalytic Domain Of A Botulinum Neurotoxin Homologue From Enterococcus Faecium
Organism: Enterococcus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-08-30 Classification: TOXIN Ligands: ZN, PO4, PGE, EDO |
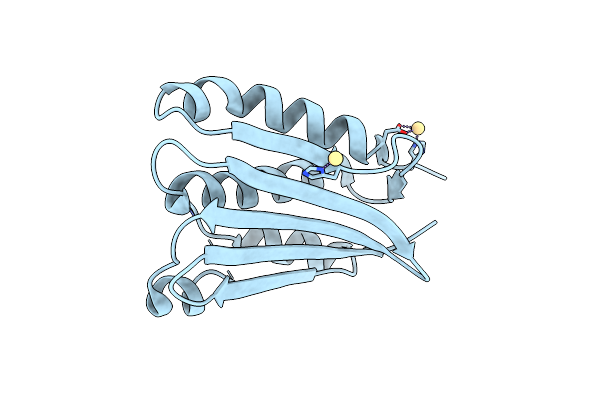 |
Organism: Enterococcus
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2023-03-22 Classification: SIGNALING PROTEIN Ligands: CD |
 |
Organism: Enterococcus
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2023-03-22 Classification: SIGNALING PROTEIN Ligands: CD |
 |
Organism: Enterococcus
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2023-03-22 Classification: SIGNALING PROTEIN Ligands: MG |
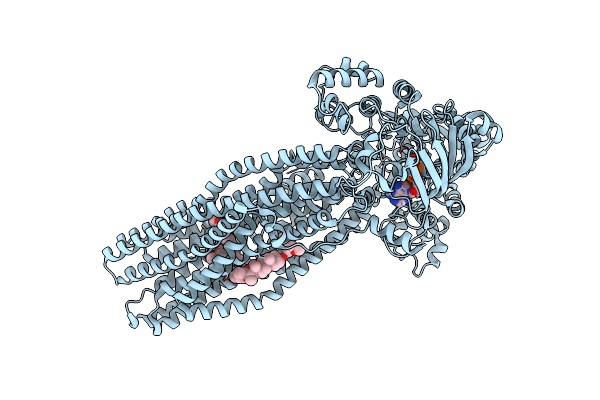 |
Cryo-Em Structure Of Abc Transporter Ste6-2P From Pichia Pastoris With Verapamil At 3.2 A Resolution
Organism: Komagataella phaffii cbs 7435
Method: ELECTRON MICROSCOPY Release Date: 2022-10-26 Classification: TRANSPORT PROTEIN Ligands: I6H, ANP, MG, 9Z9 |
 |
Cryo-Em Structure Of Abc Transporter Ste6-2P From Pichia Pastoris In Apo Conformation At 3.1 A Resolution
Organism: Komagataella phaffii cbs 7435
Method: ELECTRON MICROSCOPY Release Date: 2022-10-26 Classification: TRANSPORT PROTEIN Ligands: 9Z9 |
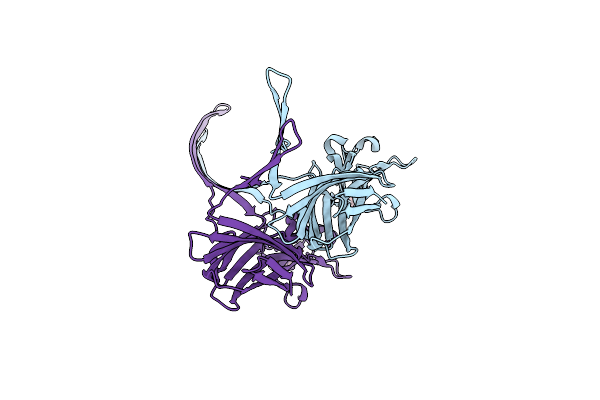 |
Organism: Enterococcus
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2022-03-16 Classification: TOXIN Ligands: MPD |
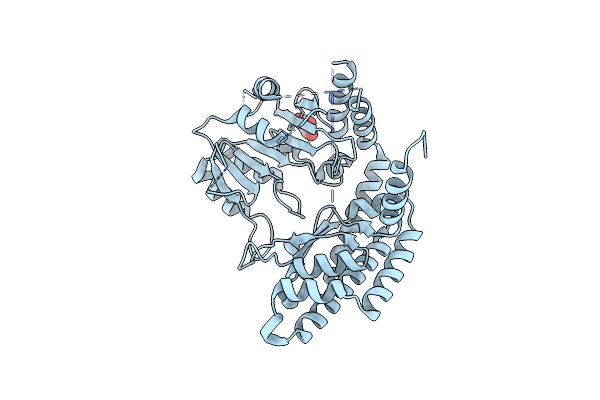 |
Crystallization And Structure Determination Of Cytoplasm Serine Hydroxymethyltransferase (Shmt) From Pichia Pastoris
Organism: Komagataella phaffii (strain atcc 76273 / cbs 7435 / cect 11047 / nrrl y-11430 / wegner 21-1)
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2018-03-14 Classification: TRANSFERASE Ligands: GOL |
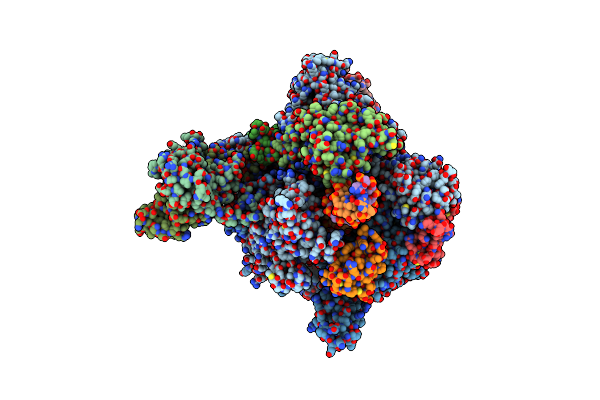 |
Organism: Komagataella pastoris, Komagataella phaffii, Synthetic construct, Komagataella phaffii (strain gs115 / atcc 20864), Komagataella phaffii (strain atcc 76273 / cbs 7435 / cect 11047 / nrrl y-11430 / wegner 21-1)
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2017-08-16 Classification: TRANSCRIPTION Ligands: ZN, MG, APC |
 |
Organism: Komagataella pastoris cbs 7435
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2016-03-09 Classification: OXIDOREDUCTASE Ligands: FAS, P6G, PGE, PO4, CA, GOL, CL |
 |
The Transient Complex Of Poplar Plastocyanin With Turnip Cytochrome F Determined With Paramagnetic Nmr
Organism: Populus nigra, Brassica rapa subsp. rapa
Method: SOLUTION NMR Release Date: 2005-05-24 Classification: PHOTOSYNTHESIS Ligands: CU, HEC |
 |

