Search Count: 4,748
All
Selected
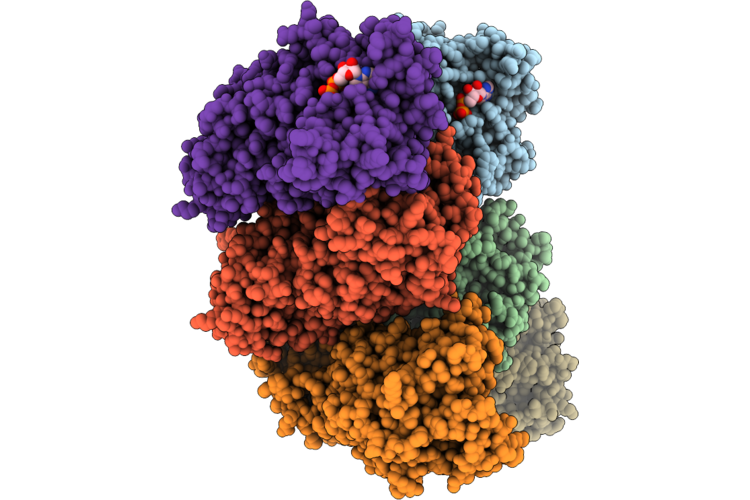 |
Crystal Structure Of Hera Like Helicae Yjgr With Ntpase Fold From Escherchia Coli
Organism: Escherichia coli str. k-12 substr. mg1655
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2026-05-06 Classification: GENE REGULATION Ligands: ATP, MG |
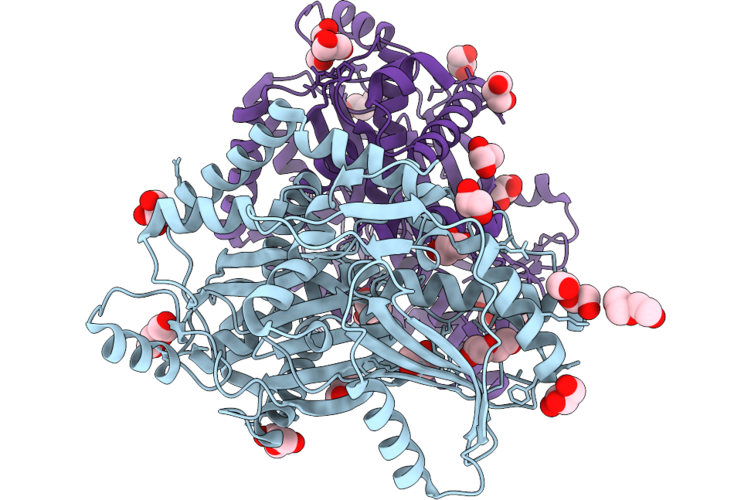 |
Organism: Coprococcus eutactus atcc 27759
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-04-29 Classification: BIOSYNTHETIC PROTEIN Ligands: GOL, PEG |
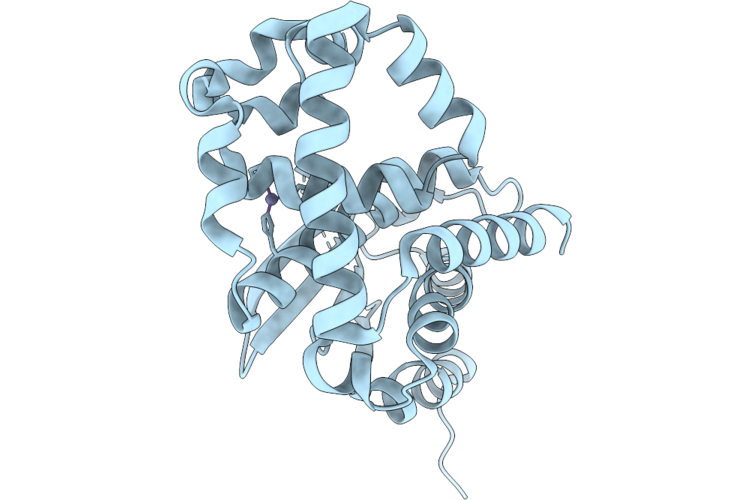 |
Crystal Structure Of Zinc-Binding Metallopeptidase (Uniprot: A0Aan3A8D2) From Bacteroides Ovatus
Organism: Bacteroides ovatus atcc 8483
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ZN |
 |
Crystal Structure Of Halide Methyltransferase From Dichomitus Squalens In Complex With Sah
Organism: Dichomitus squalens
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: SAH, ACT |
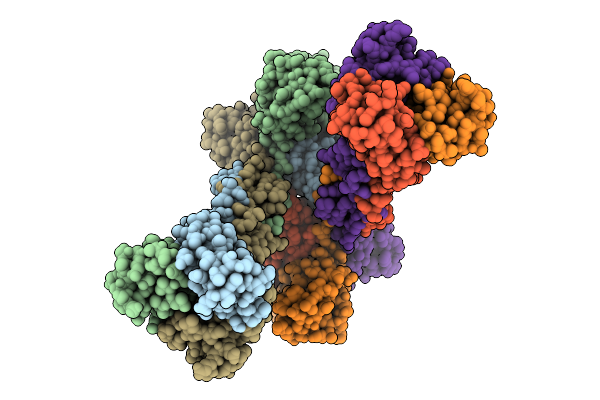 |
Crystal Structure Of Gp232 Tail Fiber Recognition Domain From Mycobacteriophage Bxz-1
Organism: Bixzunavirus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-15 Classification: VIRAL PROTEIN |
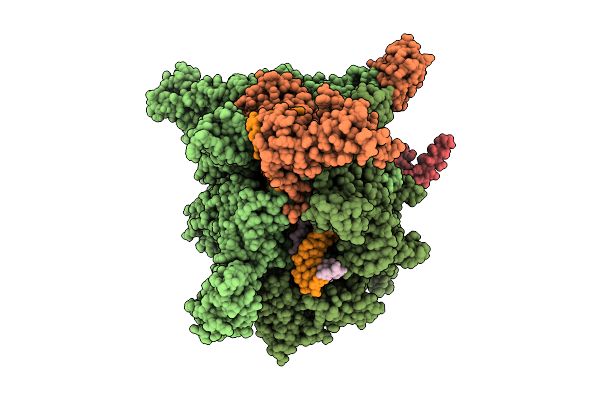 |
Cryo-Em Structure Of Escherichia Coli Transcription Initiation Complex With Gpa And Pseudouridimycin (Pum)
Organism: Escherichia coli str. k-12 substr. mg1655
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: TRANSCRIPTION, TRANSFERASE/DNA/RNA/INHIBITOR Ligands: ZN, MG, PUM |
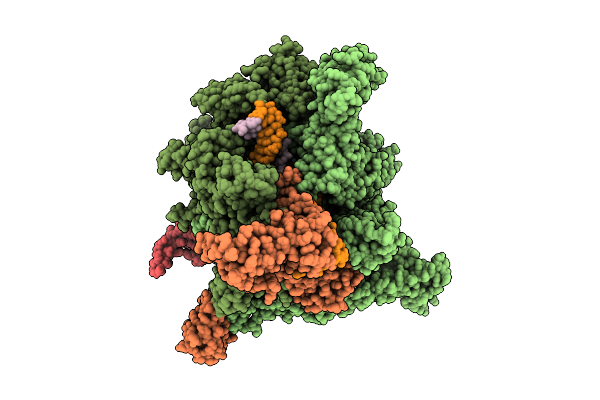 |
Cryo-Em Structure Of Escherichia Coli Transcription Initiation Complex With Gpa And Des-Hydroxy Pseudouridimycin (Des-Hydroxy Pum)
Organism: Escherichia coli str. k-12 substr. mg1655
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: TRANSFERASE/DNA/RNA/INHIBITOR Ligands: ZN, MG, A1CYH |
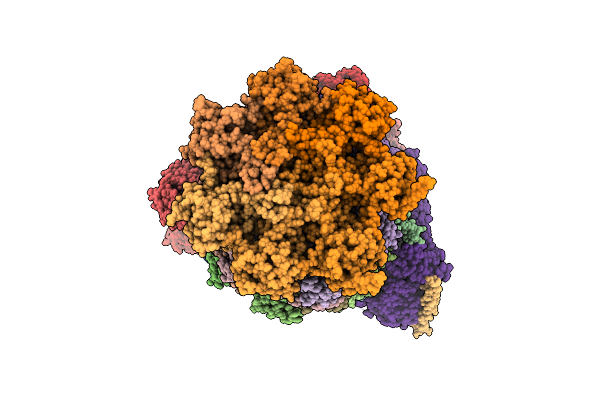 |
Organism: Homo sapiens, Escherichia coli str. k-12 substr. mg1655, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.42 Å Release Date: 2026-04-01 Classification: HYDROLASE Ligands: ZN, ATP, MG, ADP, LDZ |
 |
Crystal Structure Of Alcohol Dehydrogenase From Anoxybacillus Geothermalis D9
Organism: Anoxybacteroides rupiense
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: ZN |
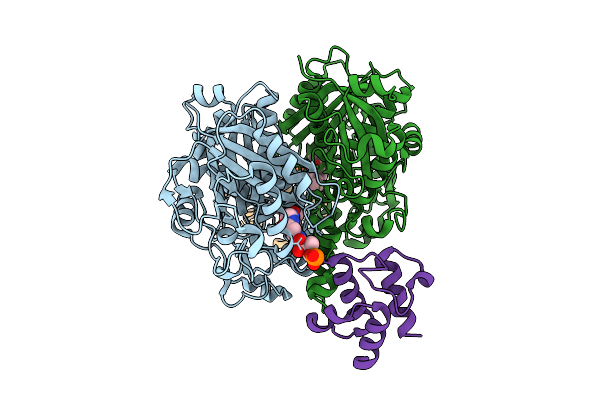 |
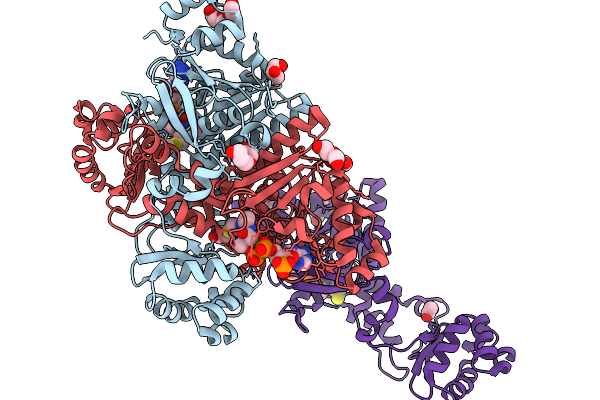
Crosslinked Complex Of Ketosynthase Fabb Mutant Fabbg107M And Acyl Carrier Protein Acpp From E.Coli With C8 Crosslinker
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2026-03-18 Classification: BIOSYNTHETIC PROTEIN Ligands: A1BMZ |
 |
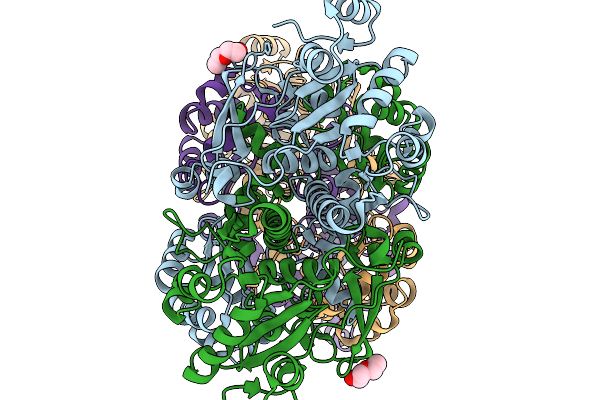
Crystal Structure Of Formyl-Coenzyme A Transferase From Brucella Melitensis In Complex With Coa
Organism: Brucella melitensis 16m1w
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: PGE, GOL, ACT, COA, PEG |
 |
Crystal Structure Of Formyl-Coenzyme A Transferase From Brucella Melitensis In Complex With Zinc
Organism: Brucella melitensis 16m1w
Method: X-RAY DIFFRACTION Resolution:2.34 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: PEG, ZN |
 |
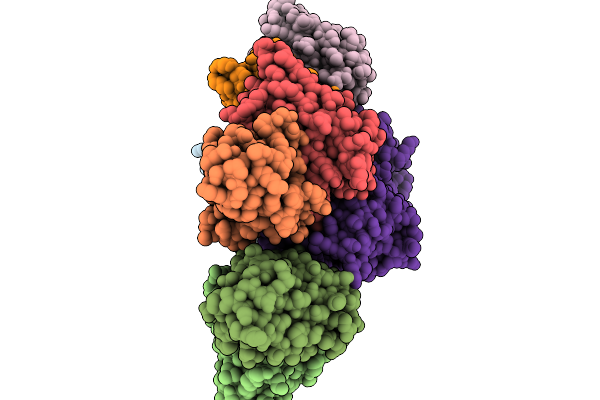
Organism: Dorea longicatena dsm 13814
Method: X-RAY DIFFRACTION Resolution:3.80 Å Release Date: 2026-02-18 Classification: STRUCTURAL PROTEIN |
 |
Stabilized Tandem Antigen Chimera Of Pfs230 And Pfs48/45 Bound By Potent Mabs
Organism: Homo sapiens, Rattus, Plasmodium falciparum
Method: ELECTRON MICROSCOPY Resolution:3.22 Å Release Date: 2026-02-04 Classification: IMMUNE SYSTEM |
 |
Cryo-Em Structure Of The Escherichia Phage Hk446 Rip1 In Complex With The Enterobacteria Phage T6 Small Terminase
Organism: Enterobacteria phage t6, Escherichia phage hk446
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: VIRAL PROTEIN |
 |
The Cryo-Em Structure Of Hera-Nura Complex With Amppnp From Thermococcus Kodakarensis
Organism: Thermococcus kodakarensis
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-02-04 Classification: TRANSLOCASE Ligands: ANP |
 |
The Cryo-Em Structure Of Hera-Nura Complex With Amppnp And Dsdna From Thermococcus Kodakarensis
Organism: Thermococcus kodakarensis, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.30 Å Release Date: 2026-02-04 Classification: DNA BINDING PROTEIN Ligands: ANP, MN |
 |
The Cryo-Em Structure Of Hera-Nura Complex With Atpgammas And Dsdna From Thermococcus Kodakarensis (State 1)
Organism: Thermococcus kodakarensis, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-02-04 Classification: DNA BINDING PROTEIN Ligands: AGS, MN |
 |
The Cryo-Em Structure Of Hera-Nura Complex With Atpgammas And Dsdna From Thermococcus Kodakarensis (State 2)
Organism: Thermococcus kodakarensis, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2026-02-04 Classification: DNA BINDING PROTEIN Ligands: AGS, MN, ADP |
 |
The Cryo-Em Structure Of Hera-Nura Complex With Atpgammas And Dsdna From Thermococcus Kodakarensis (State 3)
Organism: Thermococcus kodakarensis, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-02-04 Classification: DNA BINDING PROTEIN Ligands: AGS, MN, ADP |

