Abstact
UNLABELLED
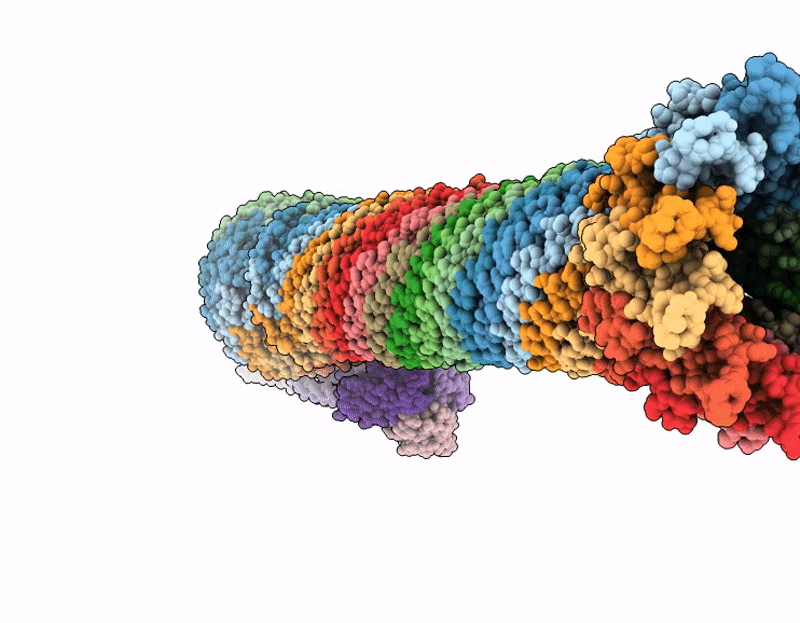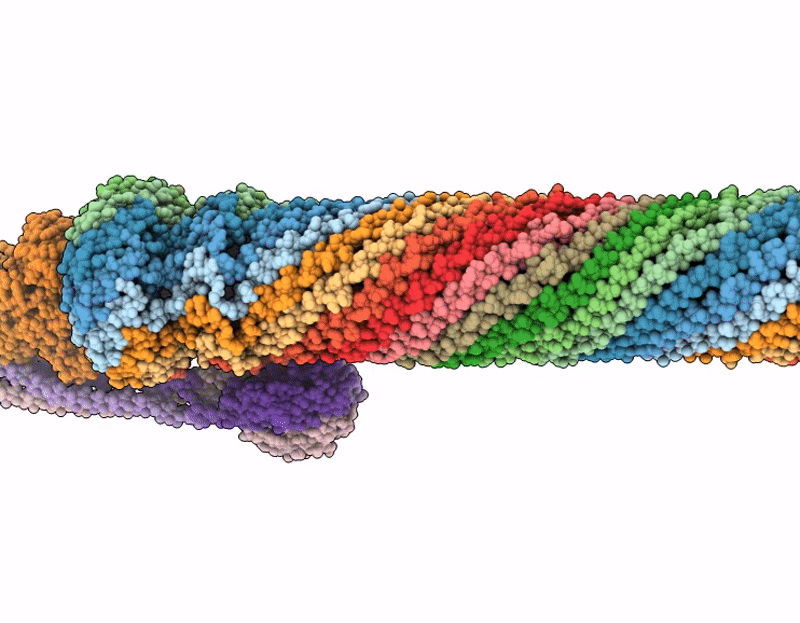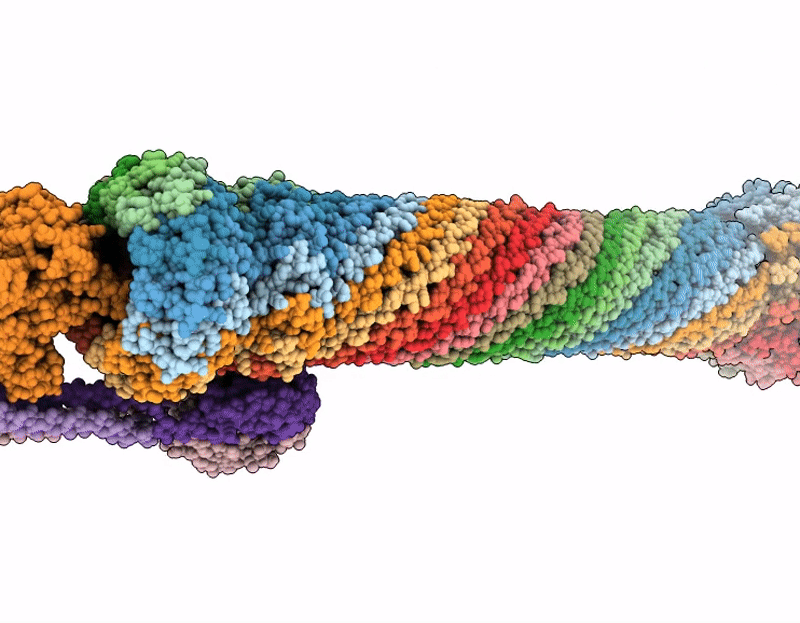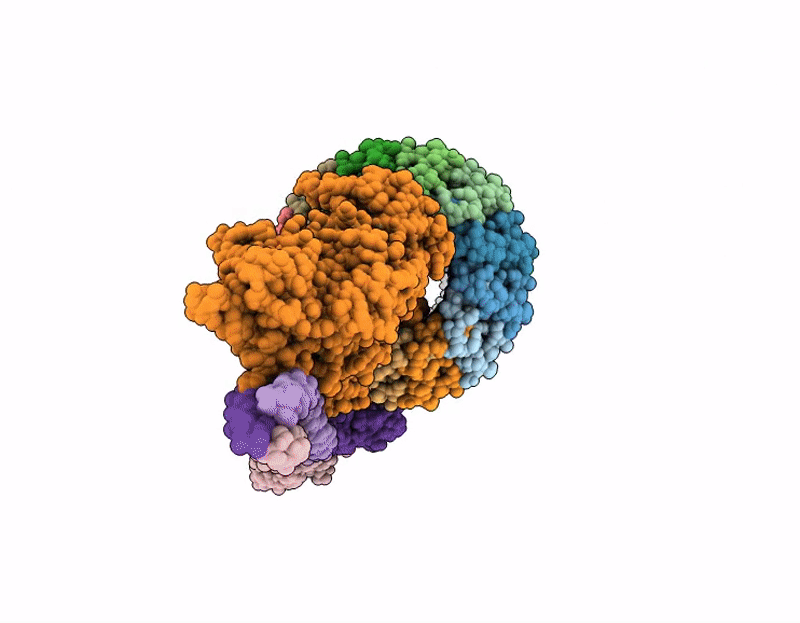
Pseudomonas aeruginosa bacteriophage (phage) DEV is a podovirus of the Schitoviridae family, related to the prototypical Escherichia coli phage N4. N4 uses the novel glycan receptor (NGR) surface glycan, presumably bound by the gp66 appendages, and the NGR transporter NfrA, recognized by the phage gp65 tail sheath, as receptors for adsorption. In contrast, DEV relies on the O-antigen moiety of lipopolysaccharide (LPS) as the primary receptor recognized by the gp53 long tail fibers. However, DEV can infect deep-rough strains that lack the O-antigen moiety by using another, still unknown receptor. Here, we provide evidence that the essential LPS transporter LptD serves as the DEV secondary receptor and that DEV gp54 is its cognate receptor-binding protein. gp54 is encoded within the essential gp56-gp55-gp54 operon, which also includes gp56, the short tail fiber gene. Using cryogenic electron microscopy, AlphaFold modeling, and genetic analysis, we show that DEV gp56, gp55, and gp54 assemble into a receptor-binding fiber (RBF) positioned laterally to a previously uncharacterized tail plug protein, gp74. The DEV RBF is functionally equivalent to the N4 sheath protein gp65, which associates with the tail plug gp53. Thus, DEV and N4 both use a glycan and its surface-exposing transporter as receptors for adsorption. To our knowledge, this is the first example of a P. aeruginosa phage using an essential outer membrane protein as a receptor, with implications for phage therapy.
IMPORTANCE
Pseudomonas aeruginosa phage DEV uses the O-antigen of lipopolysaccharide as its primary receptor. In this study, we found that LptD, an essential and highly conserved outer membrane protein, serves as the secondary receptor for DEV. This interaction is mediated by a specialized receptor-binding fiber composed of the DEV proteins gp54, gp55, and gp56. We posit that the gp56-gp55-gp54 genes form a functional module, possibly disseminated via horizontal gene transfer among distantly related phages, involved in tail sealing and the regulated unplugging of the tail upon interaction with the bacterial receptor. Given the high conservation of receptor-binding proteins among phages in the DEV Litunavirus genus, we anticipate that other members of this genus may also use LptD as their receptor. Since Litunaviruses are actively explored for phage therapy, insights into the interaction between DEV and its receptors could help develop more effective and targeted phage-based treatments.