Deposition Date
2025-03-19
Release Date
2026-03-25
Last Version Date
2026-04-08
Entry Detail
PDB ID:
9U4M
Keywords:
Title:
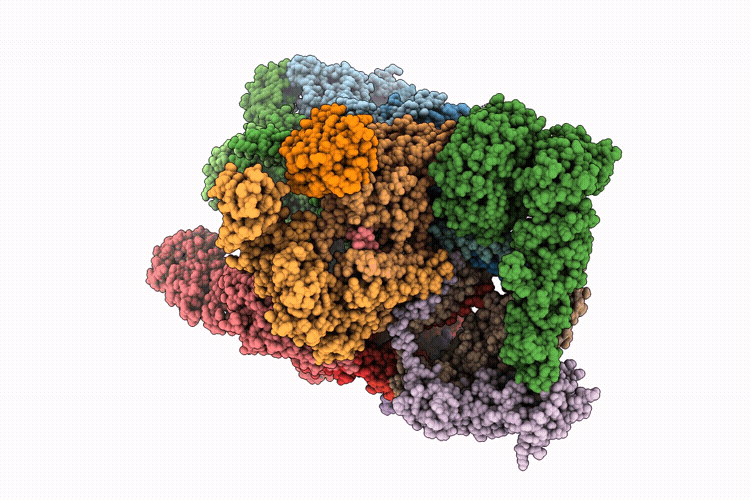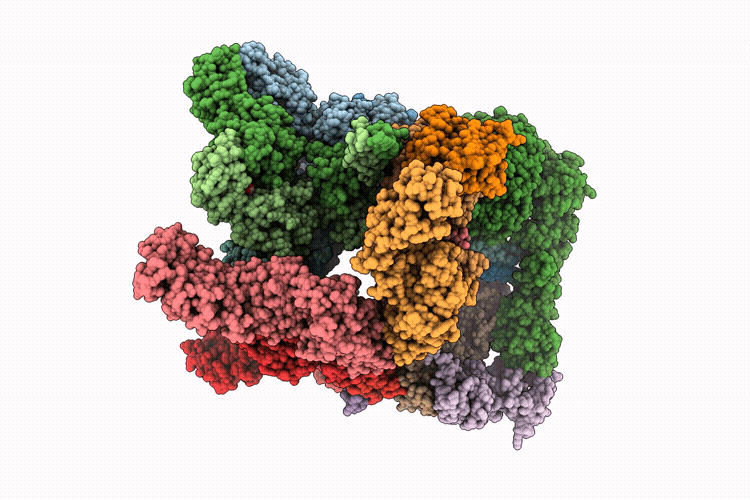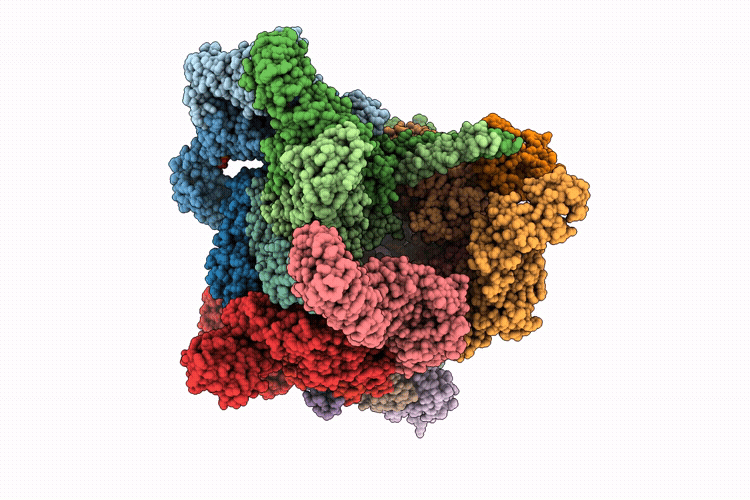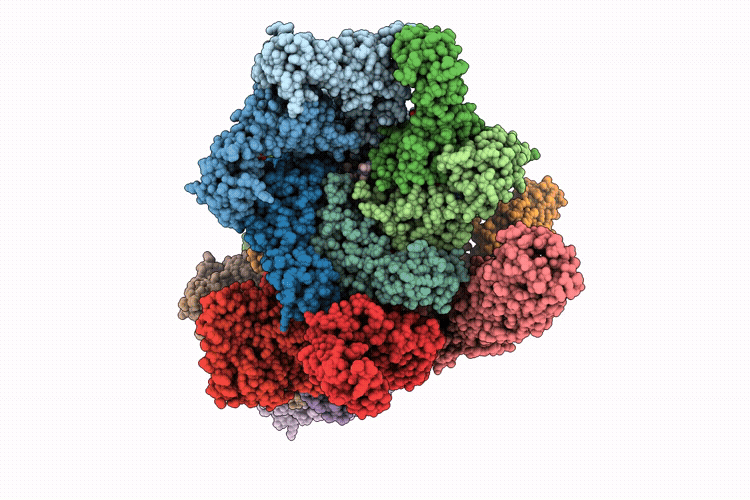
Focused refinement of 19S in the substrate-engaged human 26S proteasome bound to midnolin with RPT6 at top of spiral staircase
Biological Source:
Source Organism(s):
Homo sapiens (Taxon ID: 9606)
Method Details:
Experimental Method:
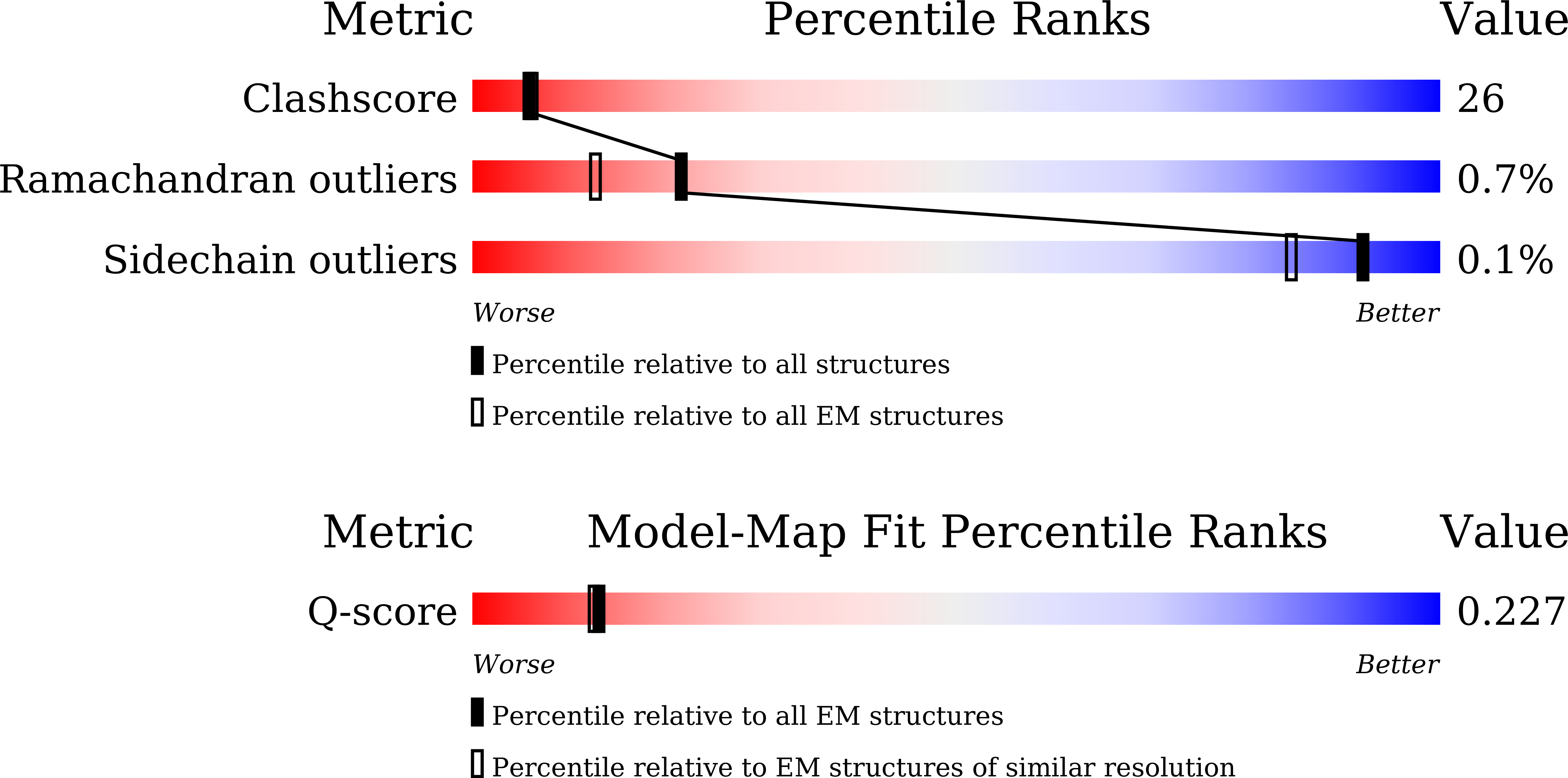
Resolution:
4.14 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE