Deposition Date
2025-03-18
Release Date
2026-04-08
Last Version Date
2026-04-08
Entry Detail
PDB ID:
9QJ0
Keywords:
Title:
Crystal structure of S-adenosyl-L-homocysteine hydrolase from P. aeruginosa: Q65A mutant soaked with adenosine and probed with rubidium to confirm disruption of a potassium binding site.
Biological Source:
Source Organism(s):
Pseudomonas aeruginosa PAO1 (Taxon ID: 208964)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.40 Å
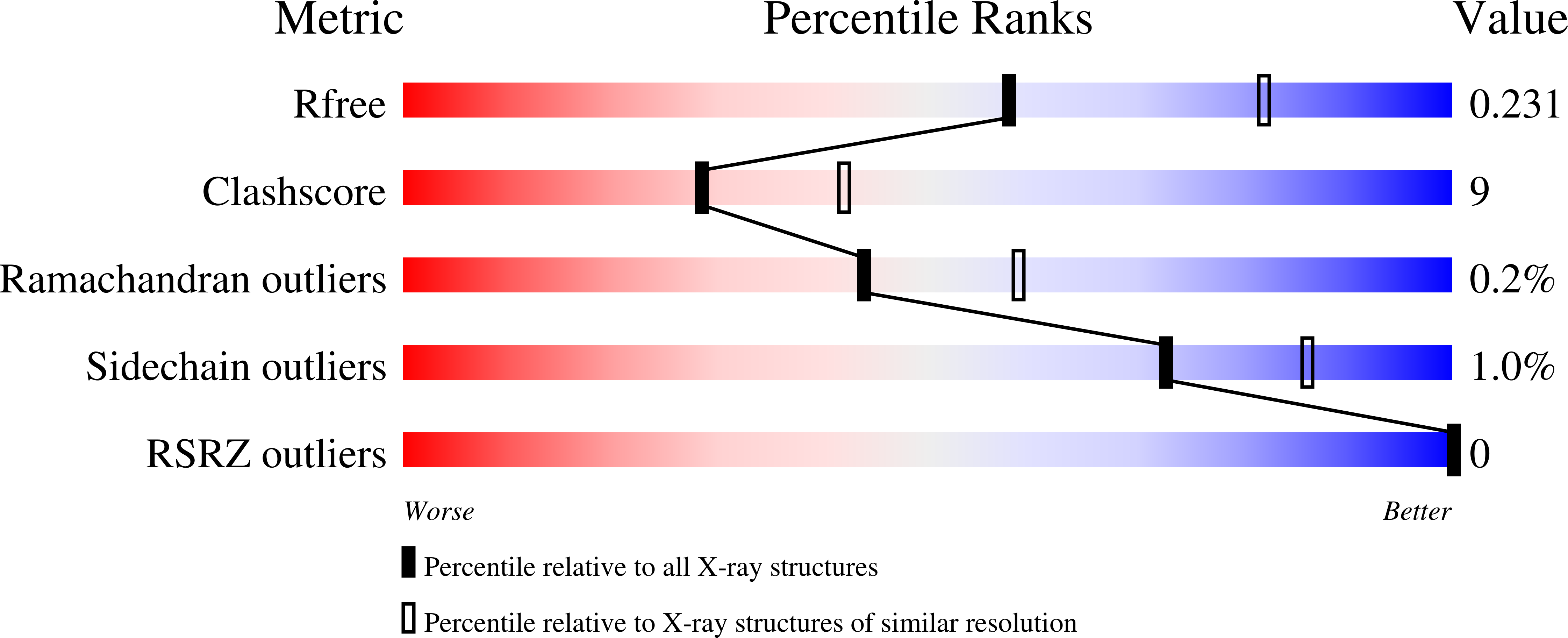
R-Value Free:
0.22
R-Value Work:
0.16
R-Value Observed:
0.17
Space Group:
P 1 21 1