Deposition Date
2025-02-01
Release Date
2026-01-28
Last Version Date
2026-02-04
Entry Detail
PDB ID:
9N43
Keywords:
Title:
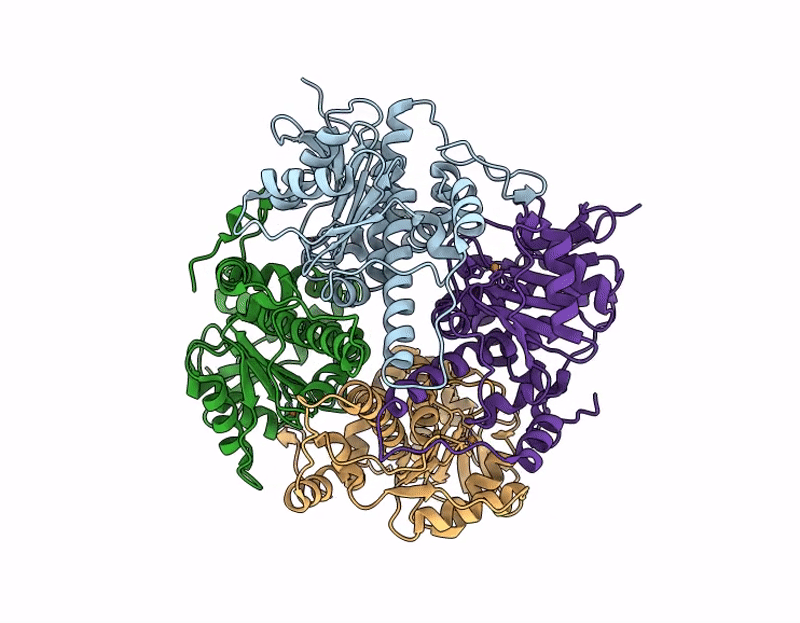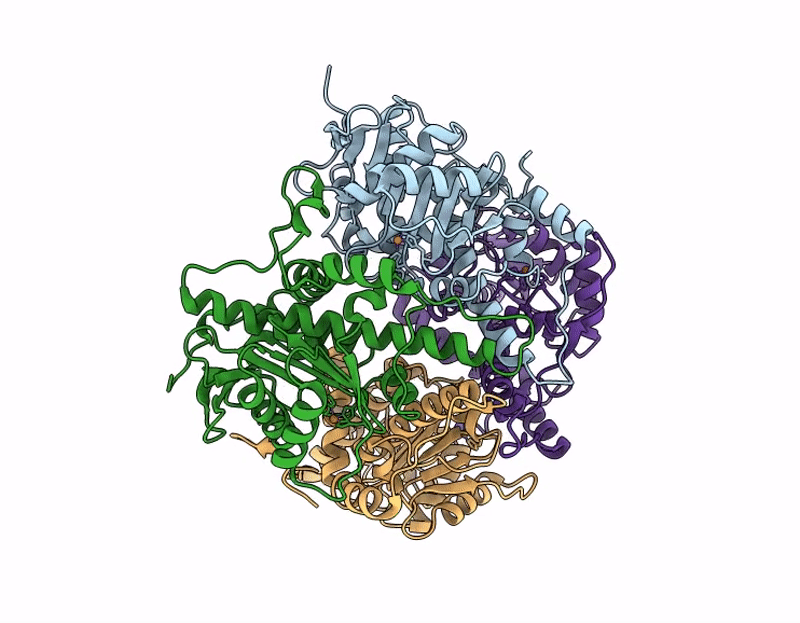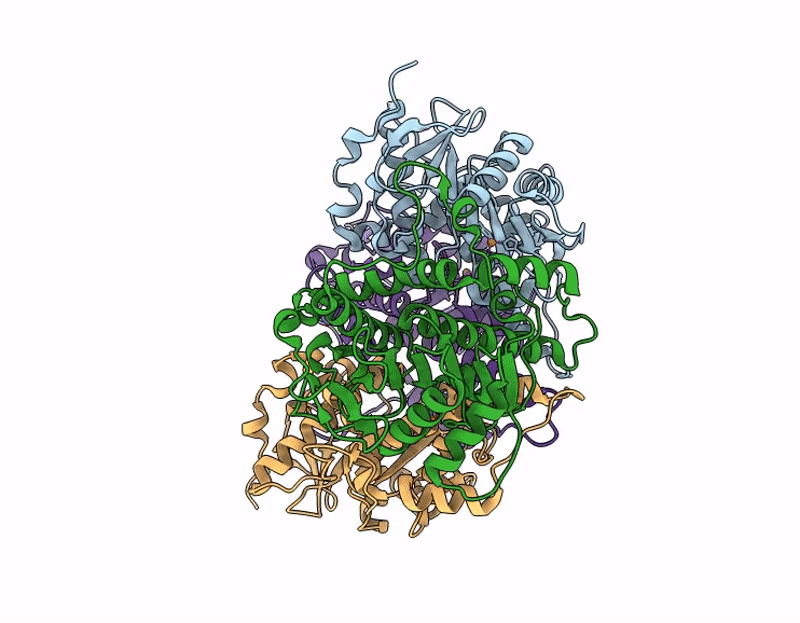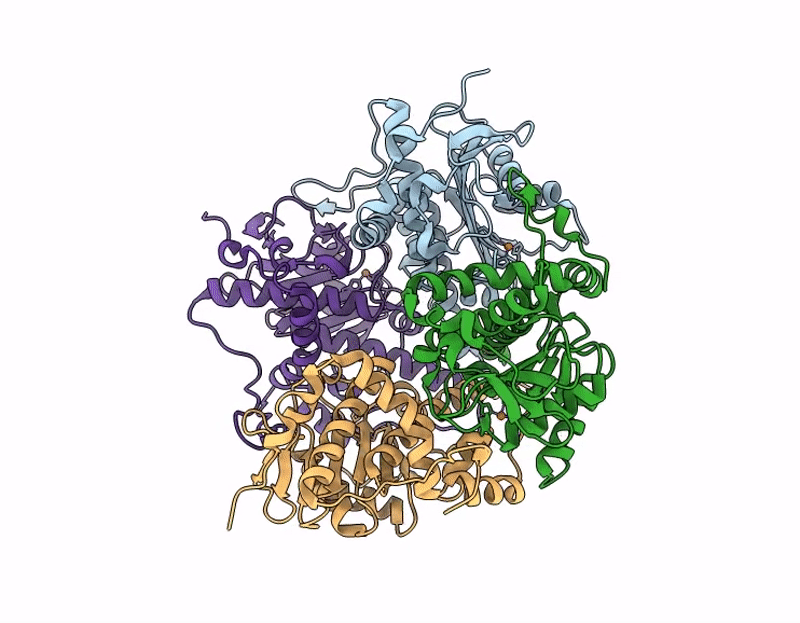
Crystal structure of none-heme iron enzyme (TqaM) from Trichoderma atroviride bound with iron
Biological Source:
Source Organism(s):
Trichoderma atroviride (Taxon ID: 63577)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.60 Å
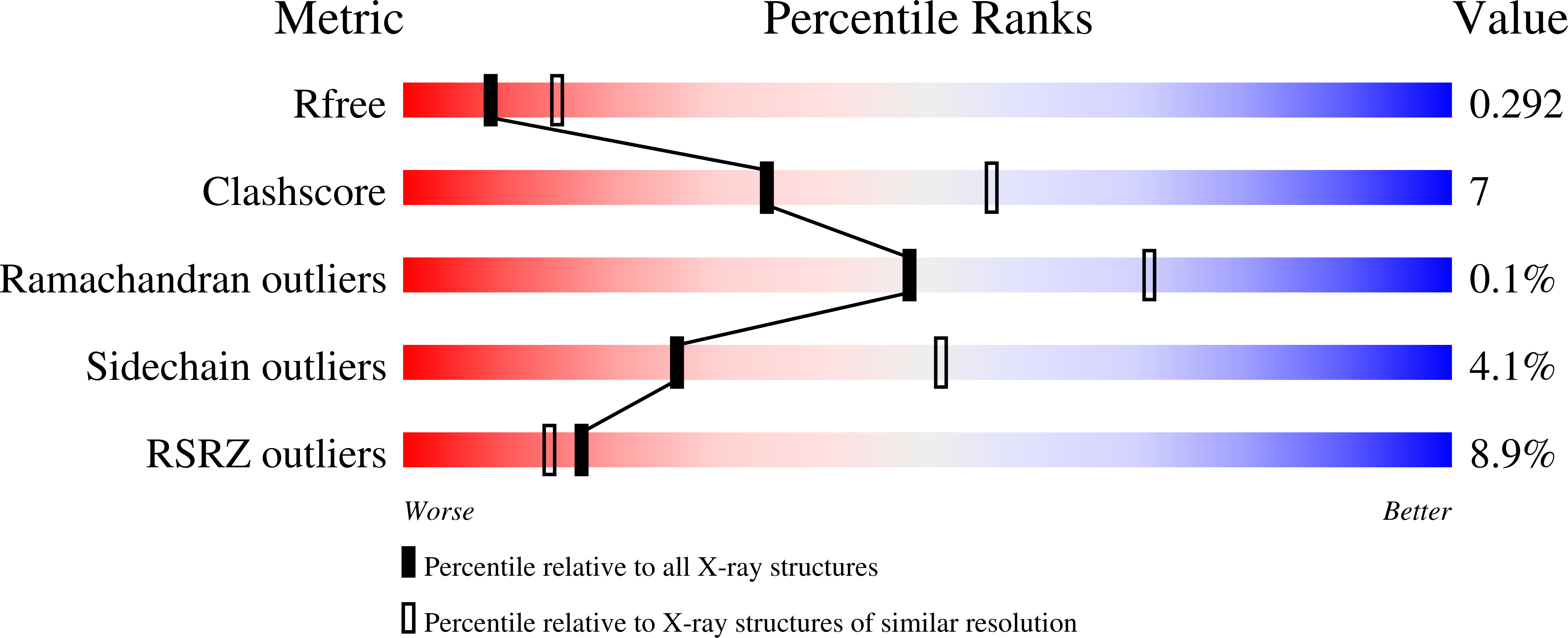
R-Value Free:
0.29
R-Value Work:
0.26
R-Value Observed:
0.26
Space Group:
C 2 2 21