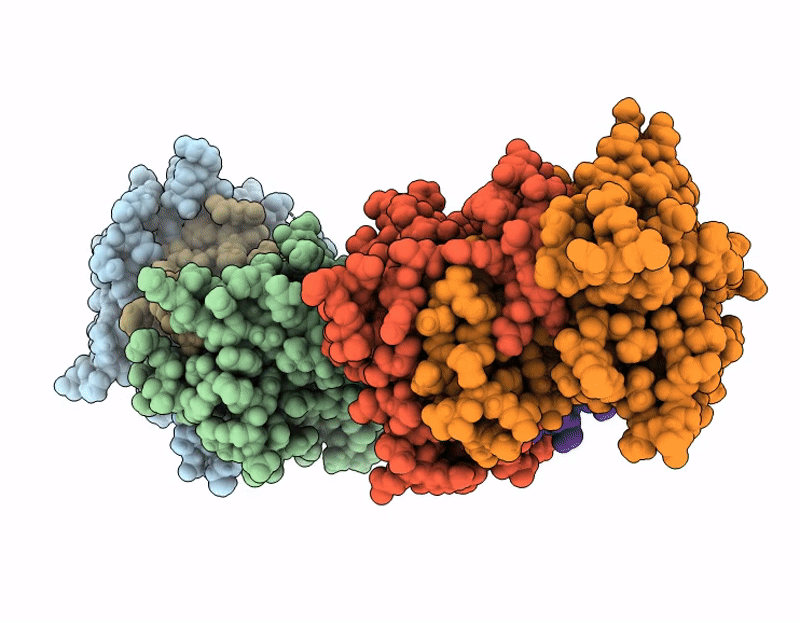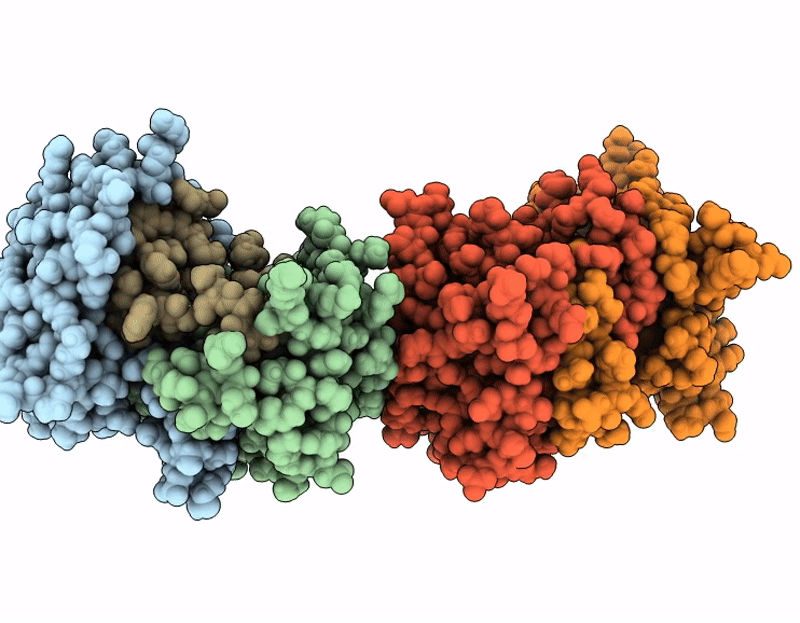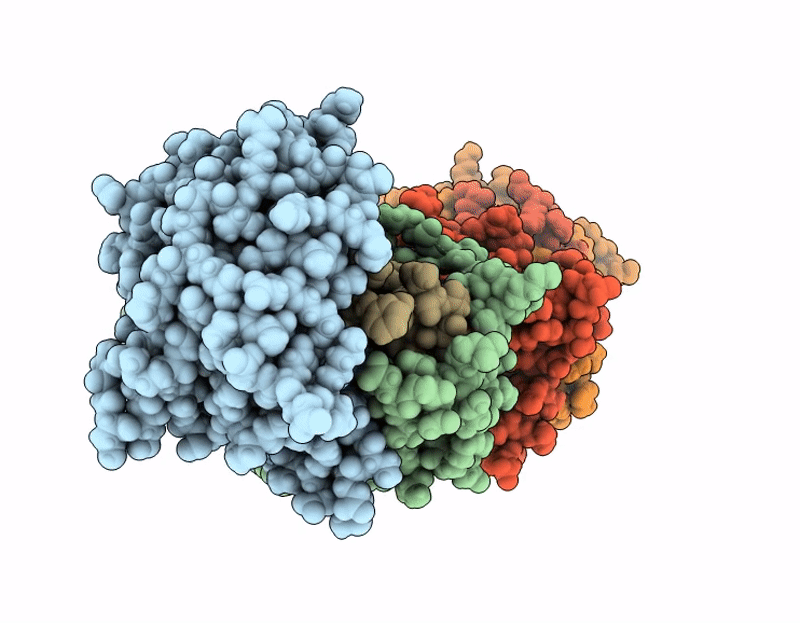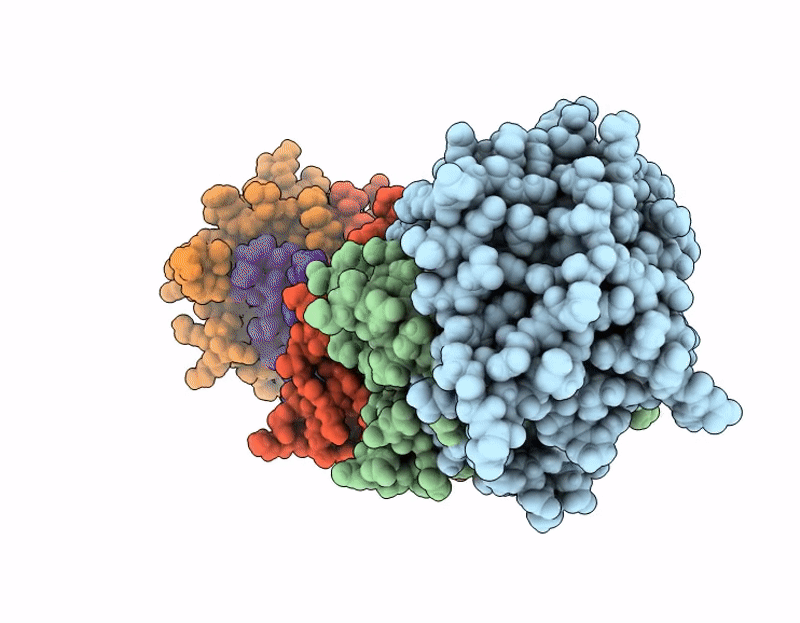
Deposition Date
2025-01-15
Release Date
2026-01-28
Last Version Date
2026-02-18
Entry Detail
PDB ID:
9MVG
Keywords:
Title:
Structure of SciW variant L66A bound to the Rhs1 transmembrane domain
Biological Source:
Source Organism(s):
Expression System(s):
Method Details:
Experimental Method:
Resolution:
3.05 Å
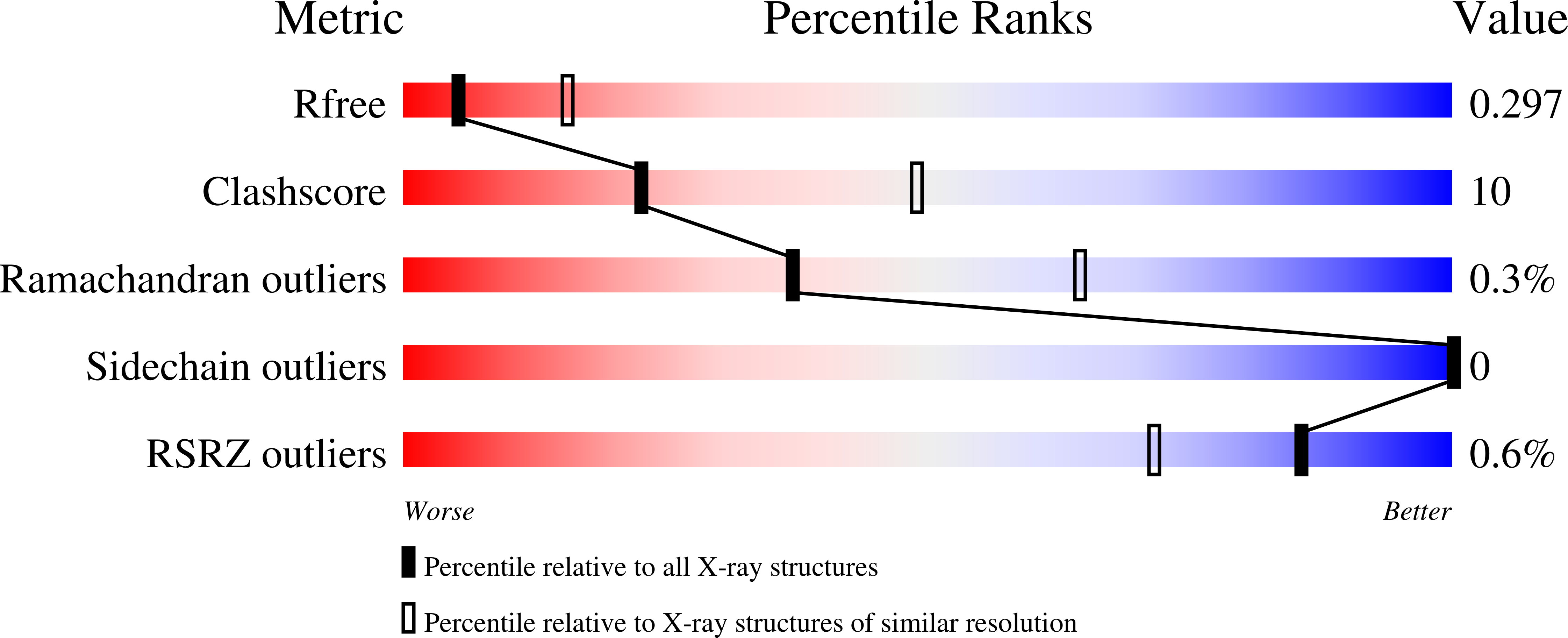
R-Value Free:
0.29
R-Value Work:
0.26
R-Value Observed:
0.26
Space Group:
C 1 2 1