Deposition Date
2025-02-27
Release Date
2026-01-14
Last Version Date
2026-02-18
Entry Detail
PDB ID:
9M2D
Keywords:
Title:
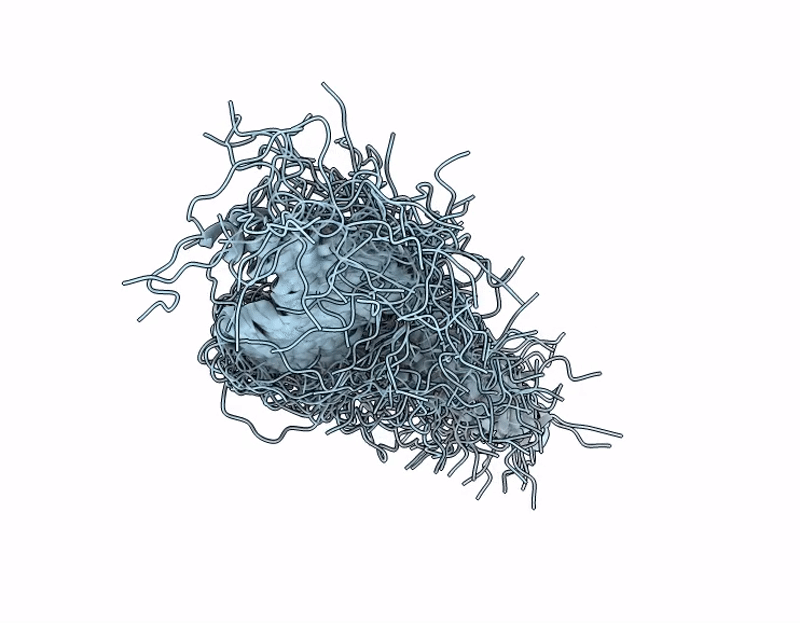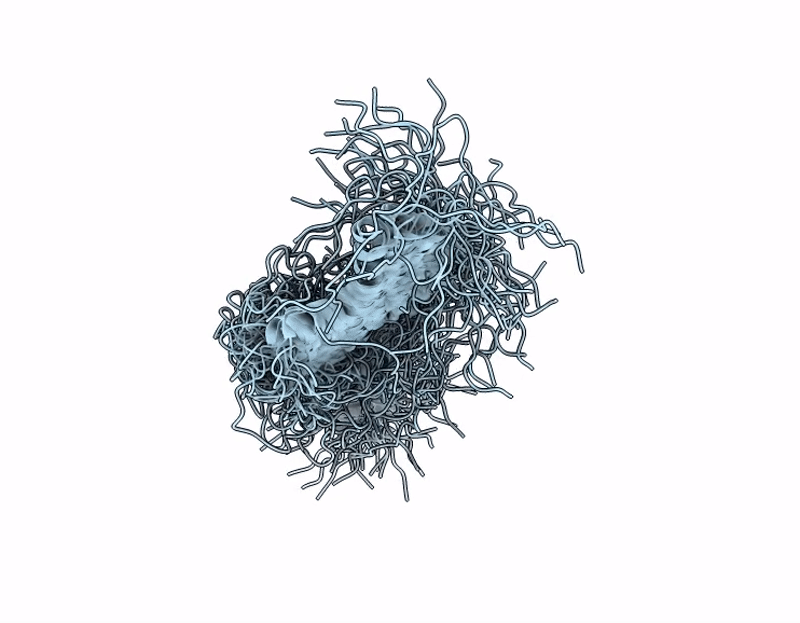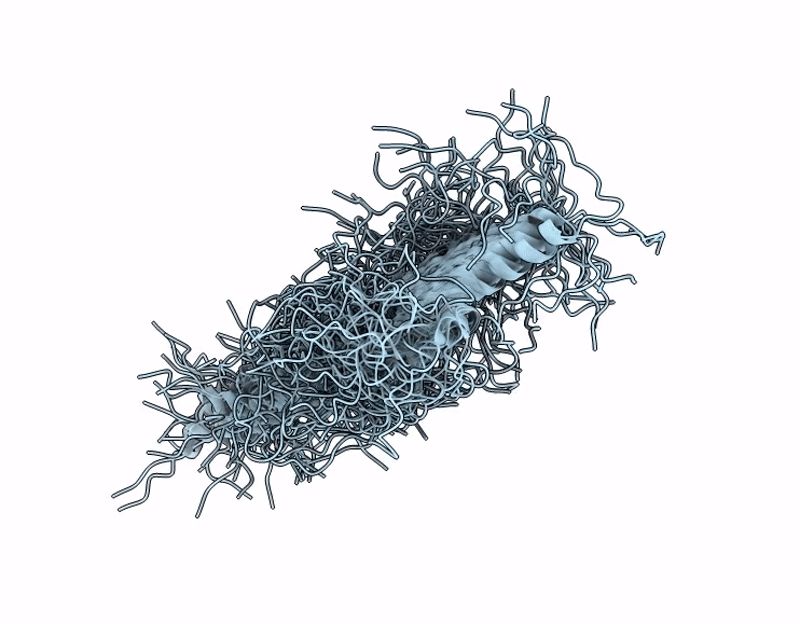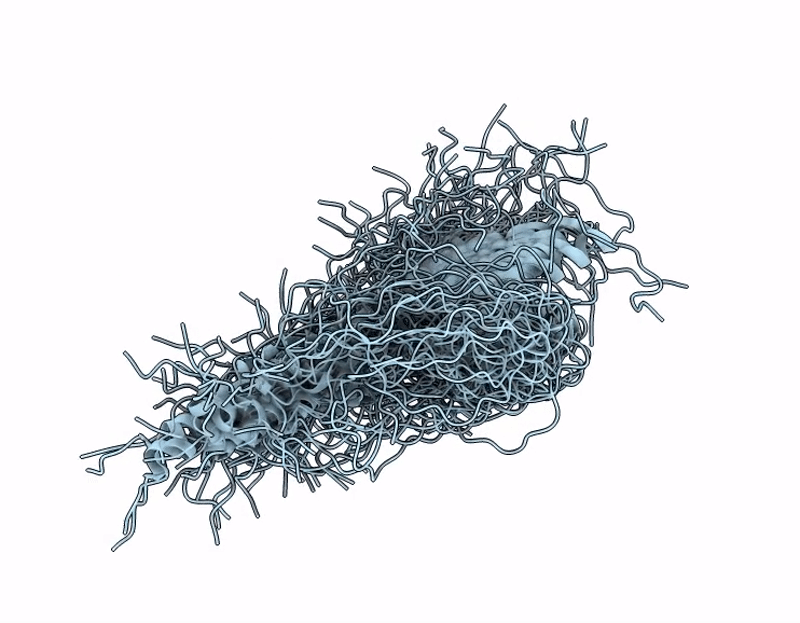
SERF1_HUMAN short isoform of Small EDRK-rich factor 1, serf1a at pH 6.8.
Biological Source:
Source Organism(s):
Homo sapiens (Taxon ID: 9606)
Expression System(s):
Method Details:
Experimental Method:
Conformers Calculated:
2000
Conformers Submitted:
100
Selection Criteria:
structures with the lowest energy