Deposition Date
2024-02-19
Release Date
2025-03-19
Last Version Date
2026-04-01
Entry Detail
PDB ID:
8YD1
Keywords:
Title:
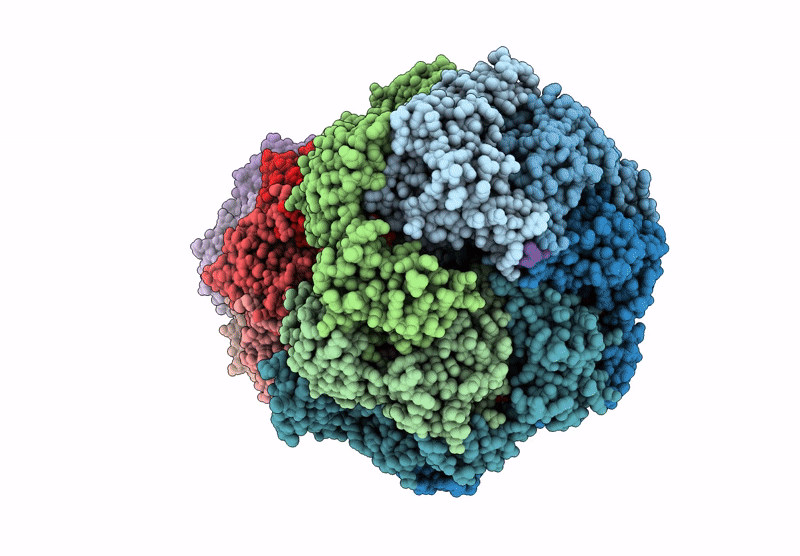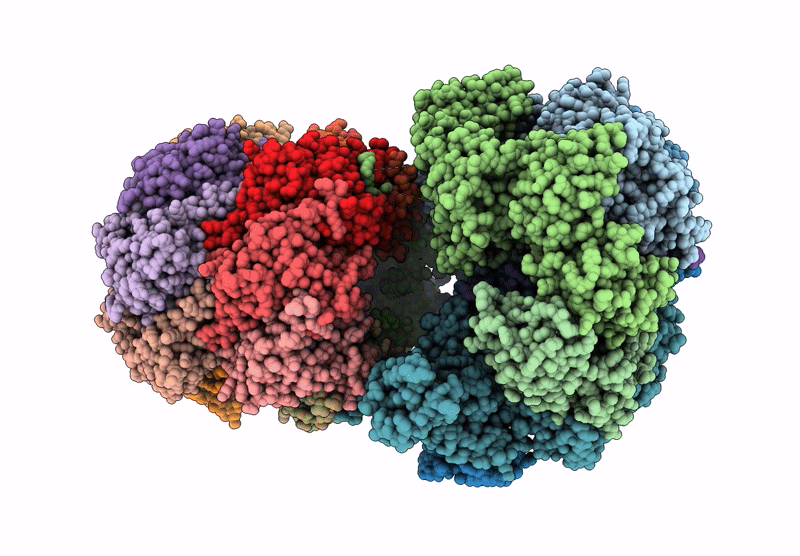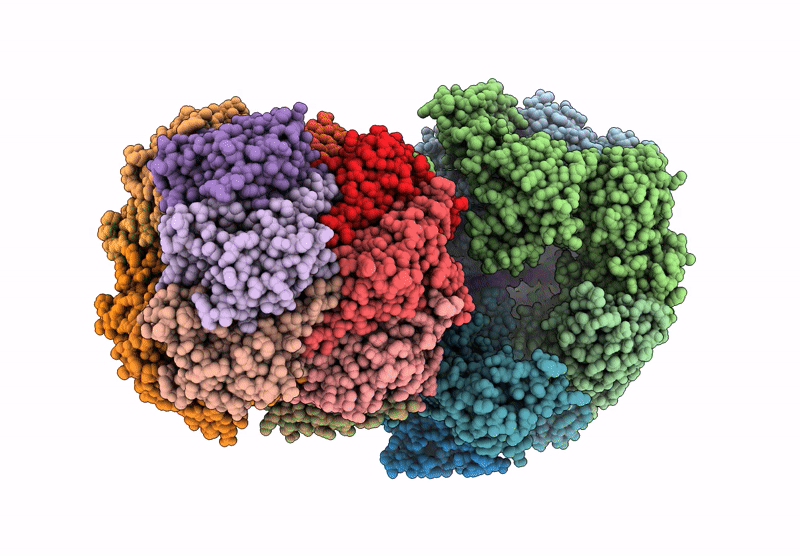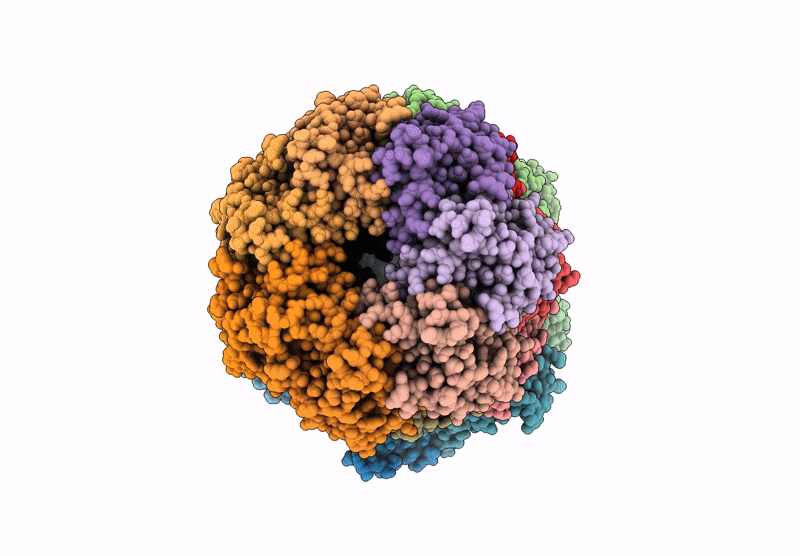
CryoEM structure of M. tuberculosis ClpC1P1P2 complex bound to bortezomib, conformation 1
Biological Source:
Source Organism(s):
Mycobacterium tuberculosis H37Rv (Taxon ID: 83332)
Bos grunniens (Taxon ID: 30521)
Bos grunniens (Taxon ID: 30521)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.81 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE