Deposition Date
2022-02-08
Release Date
2023-02-08
Last Version Date
2023-02-08
Entry Detail
PDB ID:
7R4D
Keywords:
Title:
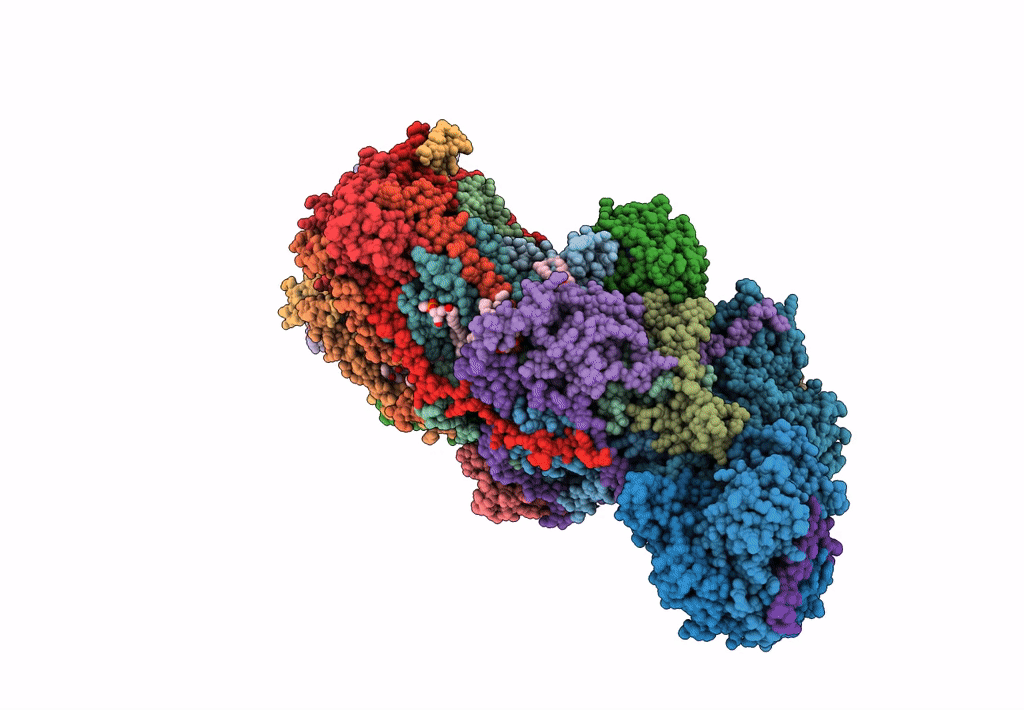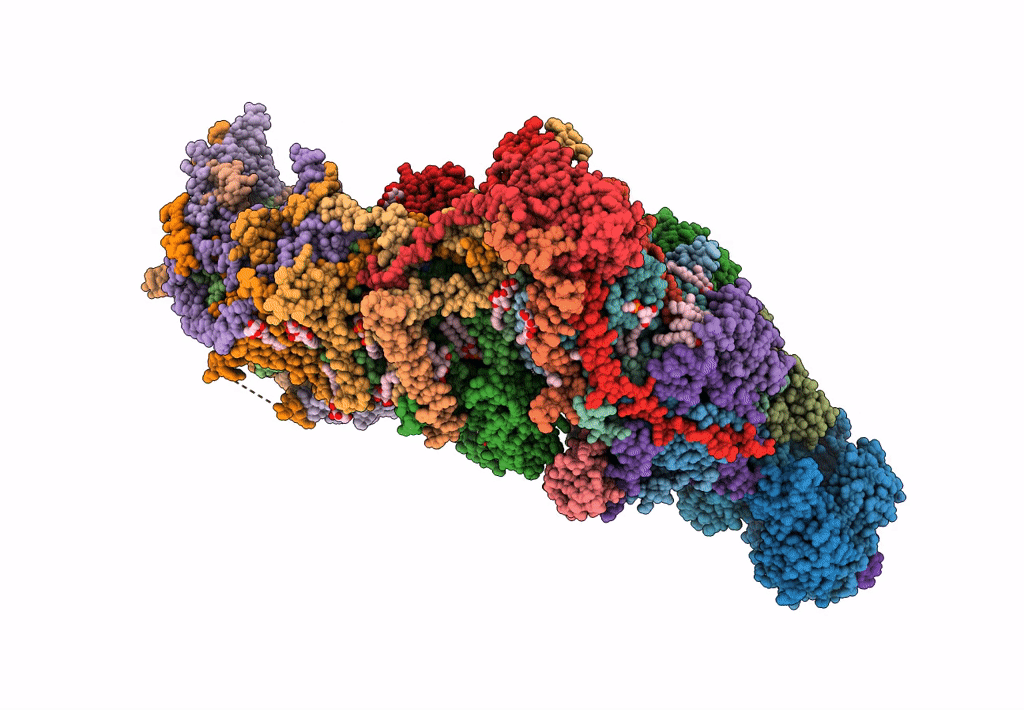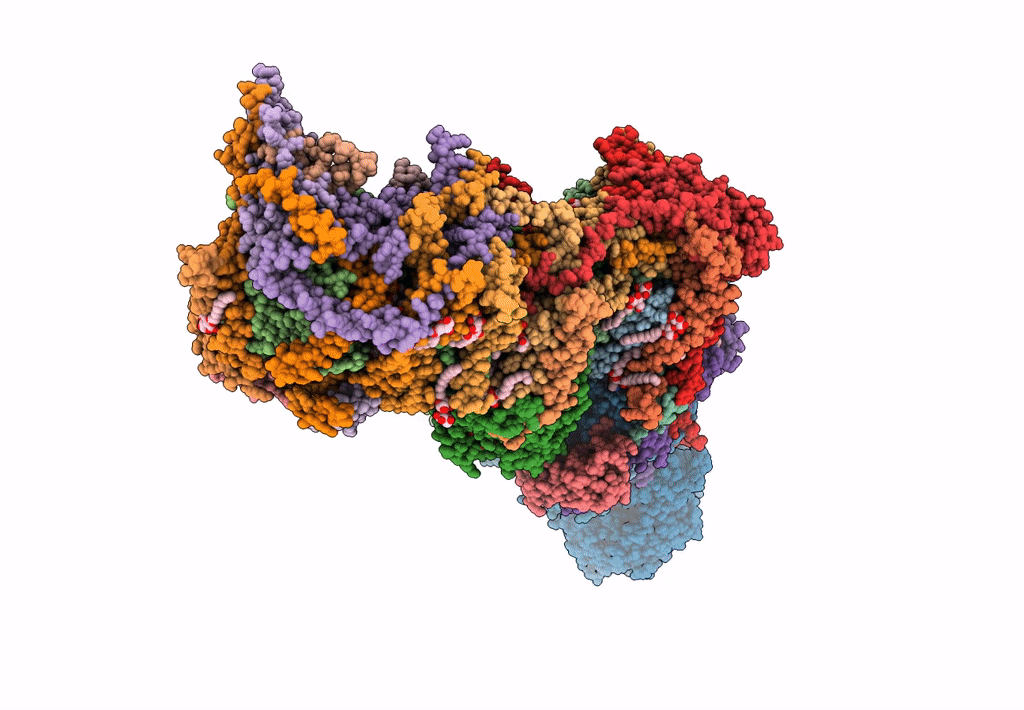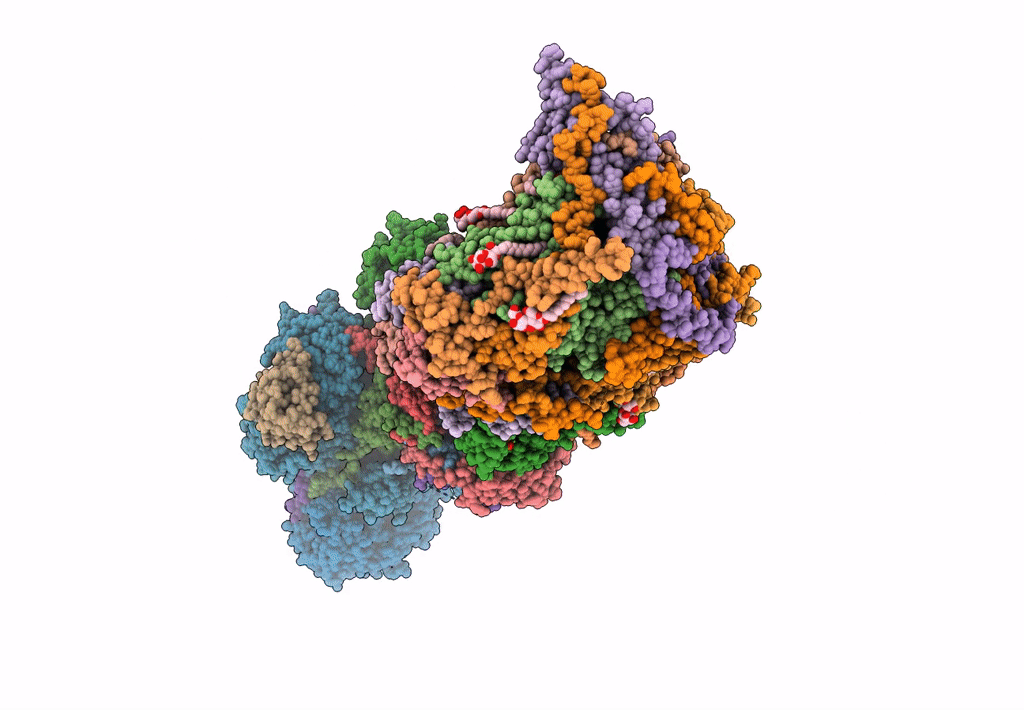
Bovine complex I in the presence of IM1761092, deactive class vi (Composite map)
Biological Source:
Source Organism(s):
Bos taurus (Taxon ID: 9913)
Method Details:
Experimental Method:
Resolution:
2.30 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE