Deposition Date
2026-01-16
Release Date
2026-04-01
Last Version Date
2026-04-08
Entry Detail
PDB ID:
22MM
Keywords:
Title:
Structure of human 26S proteasome complexed with midnolin(1-111+337-468)
Biological Source:
Source Organism(s):
Homo sapiens (Taxon ID: 9606)
Escherichia coli str. K-12 substr. MG1655 (Taxon ID: 511145)
synthetic construct (Taxon ID: 32630)
Escherichia coli str. K-12 substr. MG1655 (Taxon ID: 511145)
synthetic construct (Taxon ID: 32630)
Expression System(s):
Method Details:
Experimental Method:
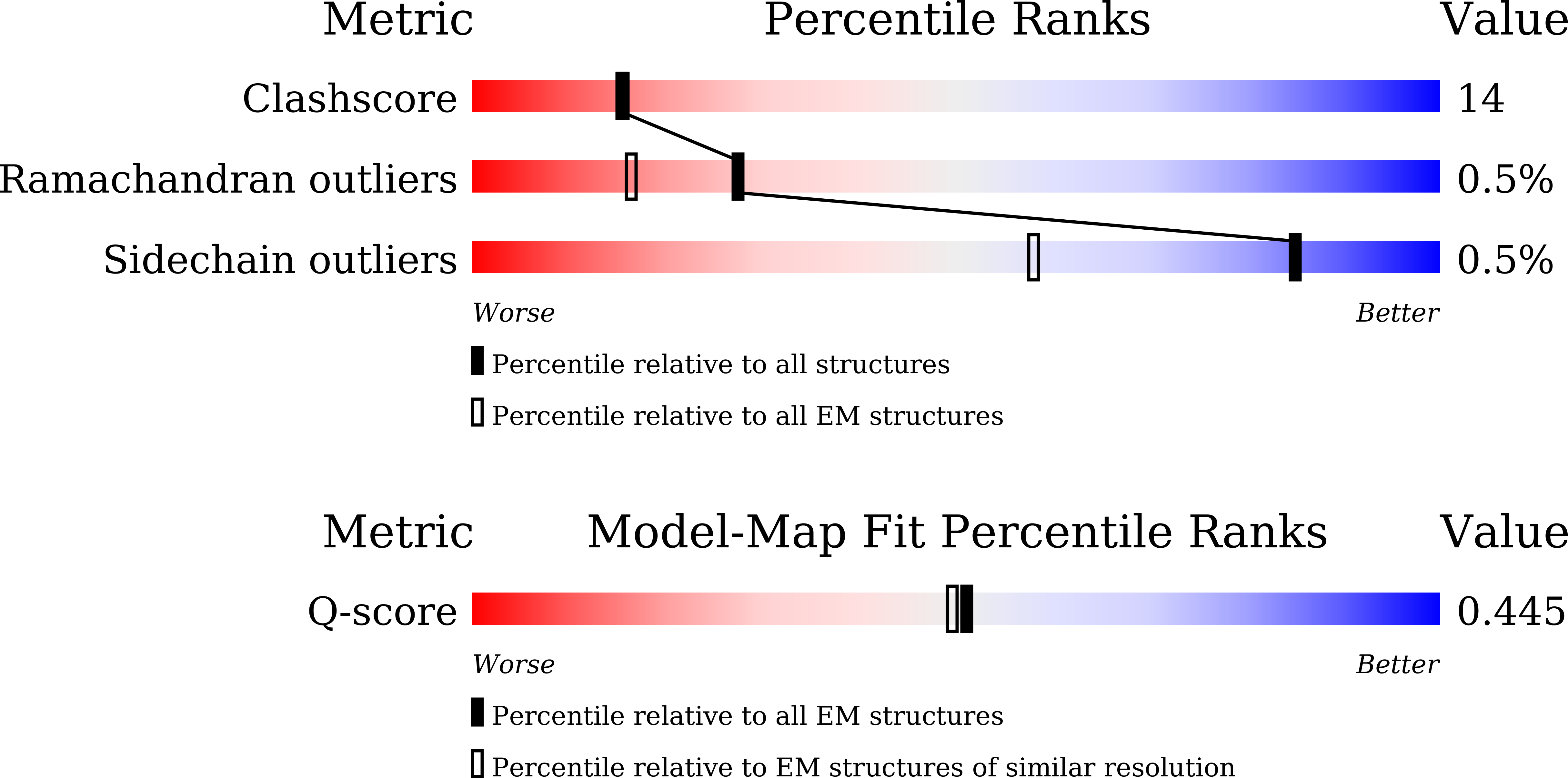
Resolution:
3.42 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE