Deposition Date
2003-05-19
Release Date
2003-10-09
Last Version Date
2024-10-23
Entry Detail
PDB ID:
1OGY
Keywords:
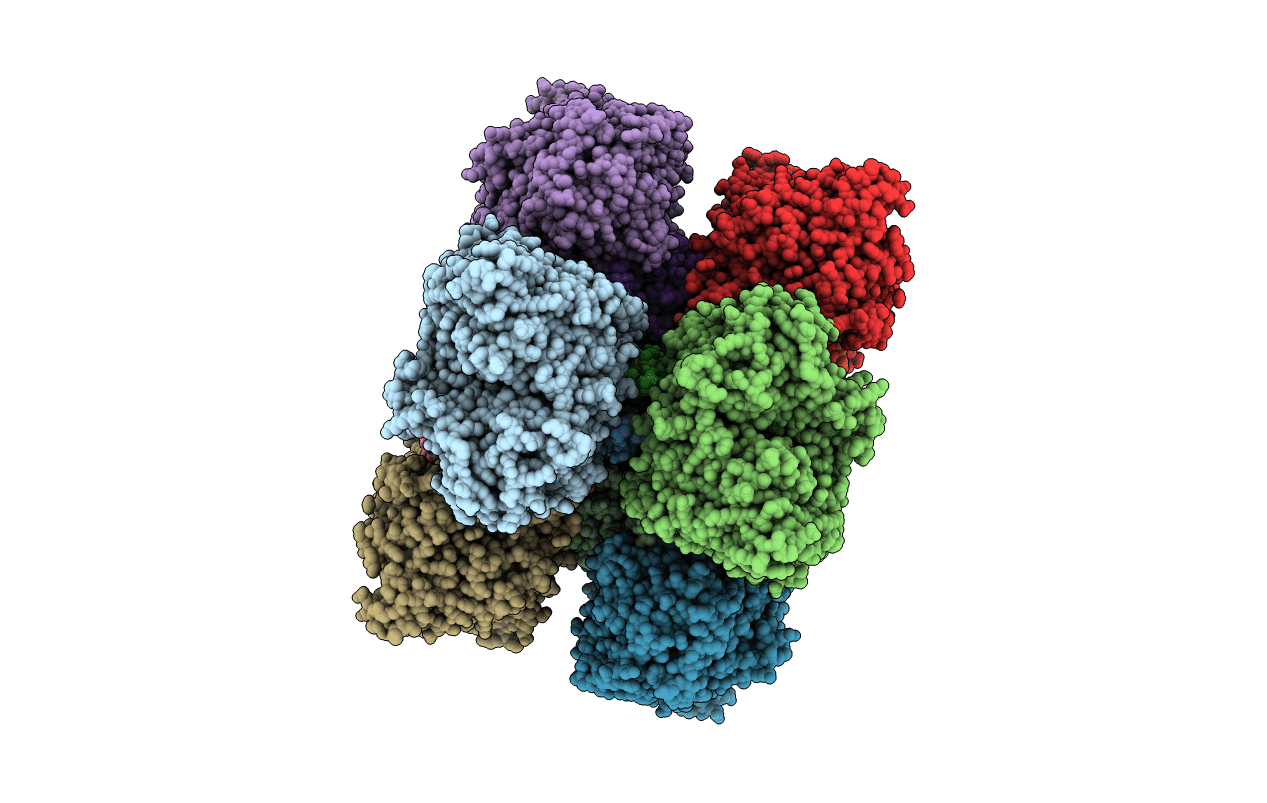
Title:
Crystal structure of the heterodimeric nitrate reductase from Rhodobacter sphaeroides
Biological Source:
Source Organism(s):
RHODOBACTER SPHAEROIDES (Taxon ID: 1063)
Expression System(s):
Method Details:
Experimental Method:
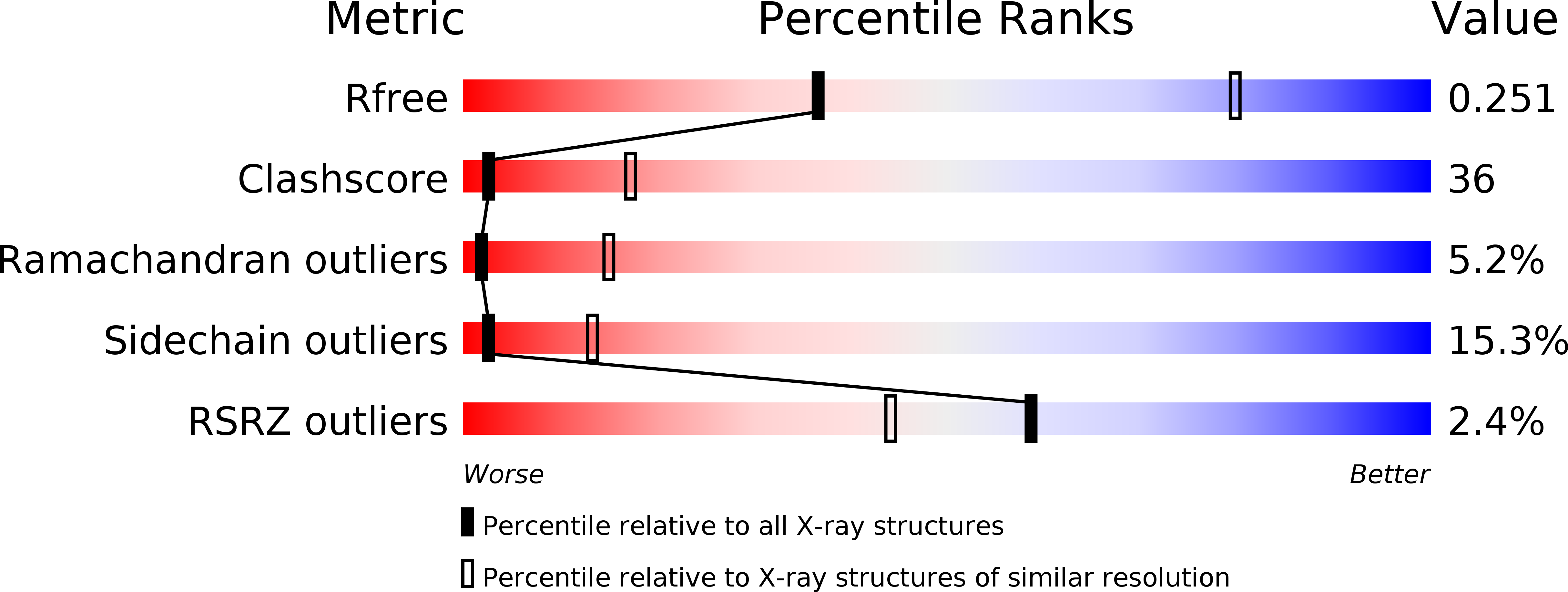
Resolution:
3.20 Å
R-Value Free:
0.26
R-Value Work:
0.25
R-Value Observed:
0.25
Space Group:
P 1 21 1