Deposition Date
2026-03-18
Release Date
2026-04-01
Last Version Date
2026-04-01
Entry Detail
PDB ID:
11YO
Keywords:
Title:
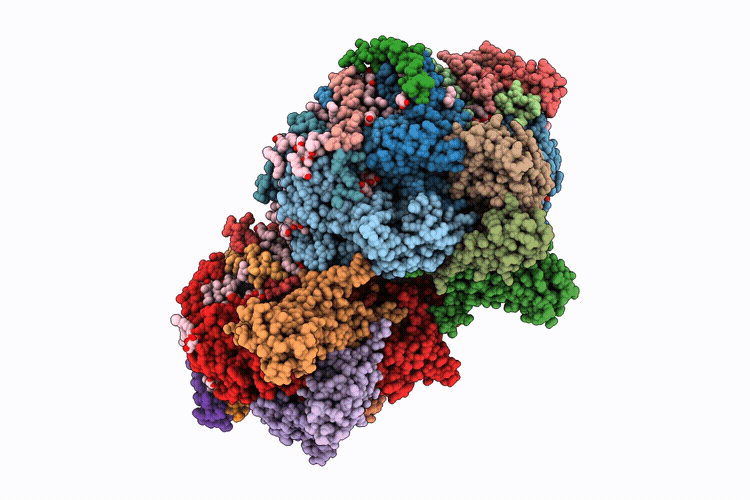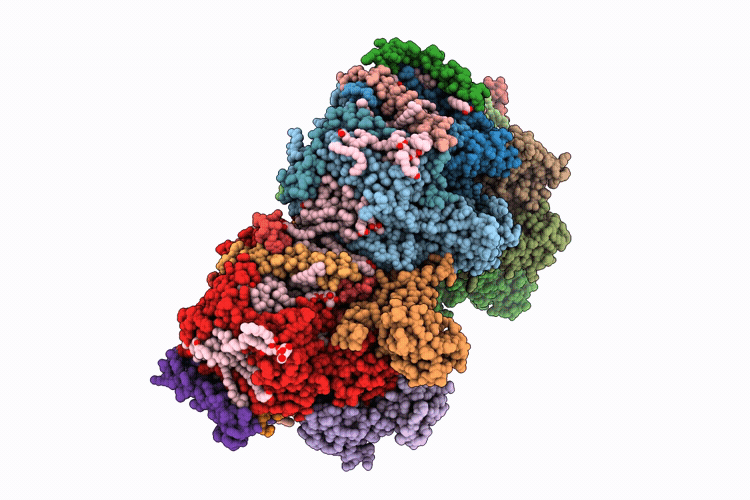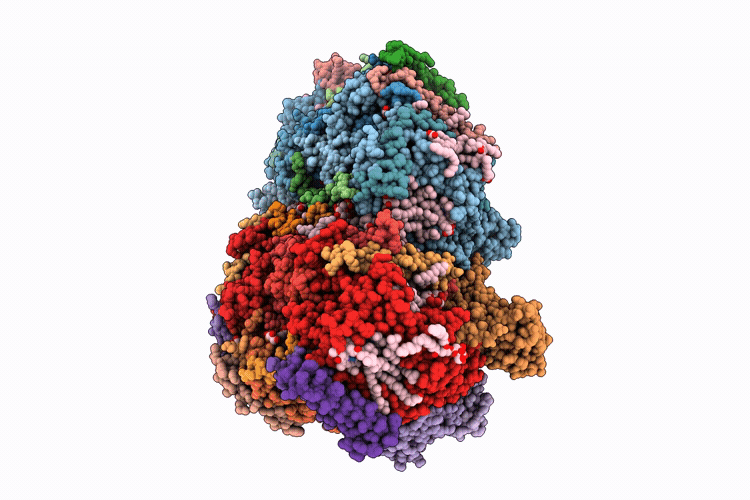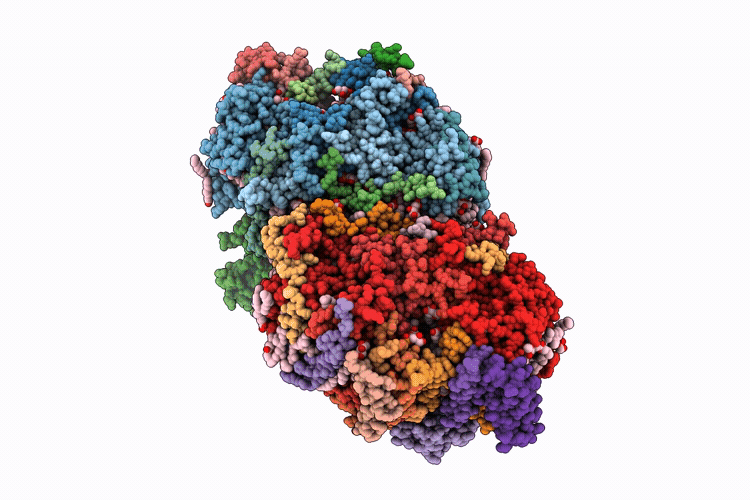
Single-conformation model re-refinement of 2F/S3-rich PSII intermediate structure at 2.09 Angstrom resolution
Biological Source:
Source Organism(s):
Thermosynechococcus vestitus BP-1 (Taxon ID: 197221)
Method Details:
Experimental Method:
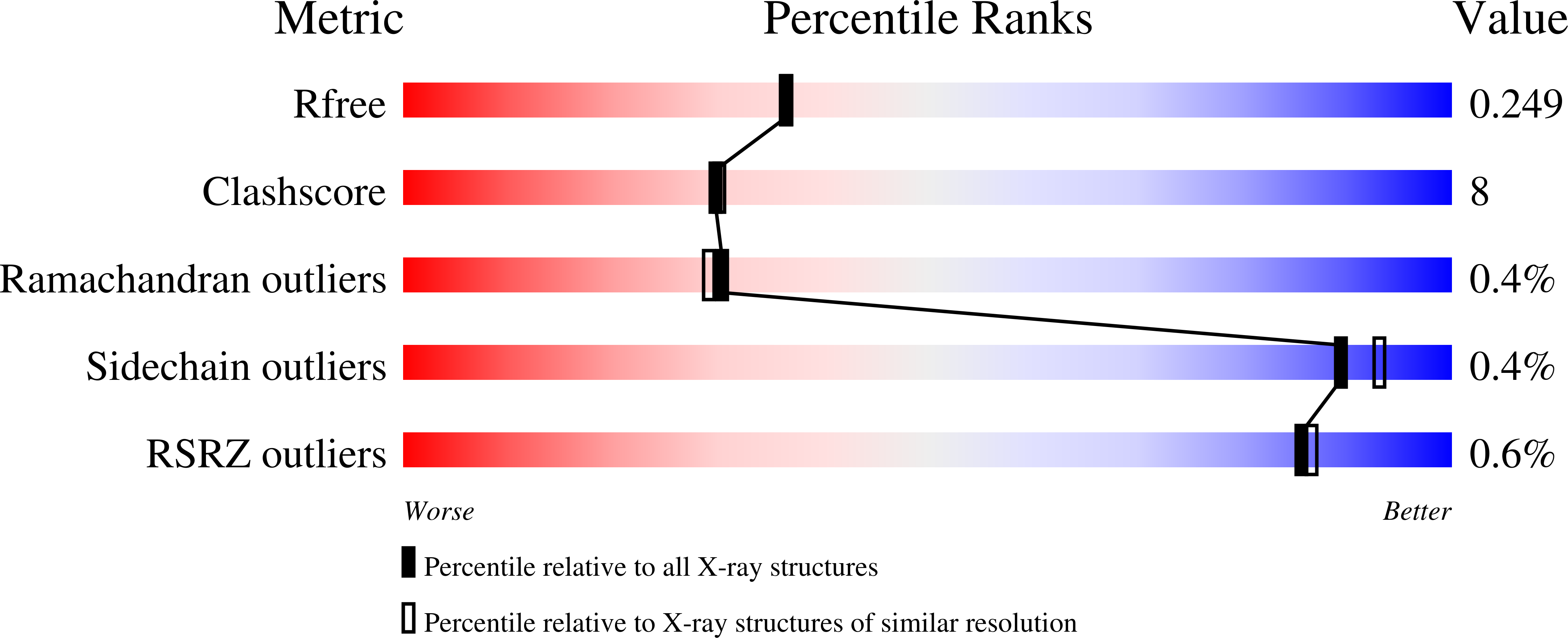
Resolution:
2.09 Å
R-Value Free:
0.24
R-Value Work:
0.18
Space Group:
P 21 21 21