Search Count: 14
All
Selected
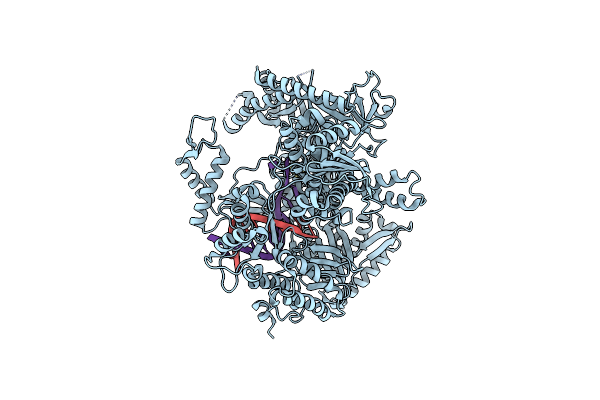 |
Organism: Saccharomyces cerevisiae, Dna molecule
Method: ELECTRON MICROSCOPY Release Date: 2024-10-09 Classification: DNA BINDING PROTEIN/DNA Ligands: SF4 |
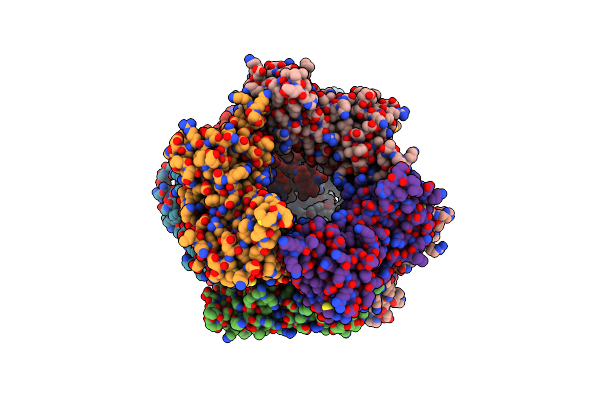 |
Cryo-Em Structure Of S. Cerevisiae Ctf18-Rfc-Pcna Complex In Apo State Conformation I
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2024-09-04 Classification: DNA BINDING PROTEIN/DNA Ligands: ADP, AGS, MG |
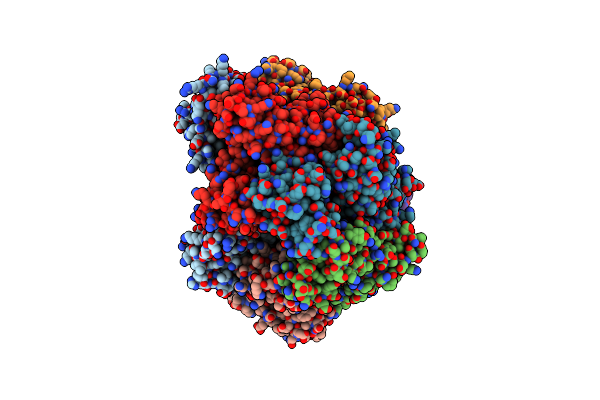 |
Cryo-Em Structure Of S. Cerevisiae Ctf18-Rfc-Pcna Complex In Apo State Conformation I
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2024-09-04 Classification: DNA BINDING PROTEIN/DNA Ligands: AGS, MG, ADP |
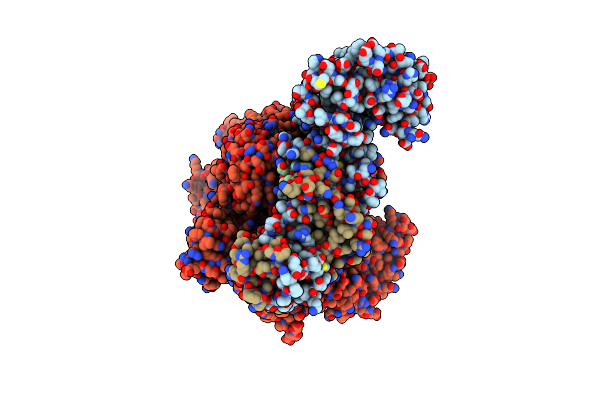 |
Organism: Saccharomyces cerevisiae, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2024-09-04 Classification: DNA BINDING PROTEIN/DNA Ligands: SF4 |
 |
Organism: Saccharomyces cerevisiae, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2024-09-04 Classification: DNA BINDING PROTEIN/DNA Ligands: SF4, AGS, MG, ADP |
 |
Organism: Saccharomyces cerevisiae, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2024-09-04 Classification: DNA BINDING PROTEIN/DNA Ligands: AGS, MG, ADP |
 |
Structure Of Eukaryotic Cmg Helicase At A Replication Fork And Implications
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c), Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Resolution:4.90 Å Release Date: 2017-02-08 Classification: REPLICATION Ligands: ANP |
 |
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c), Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:6.10 Å Release Date: 2017-01-25 Classification: REPLICATION Ligands: ATP |
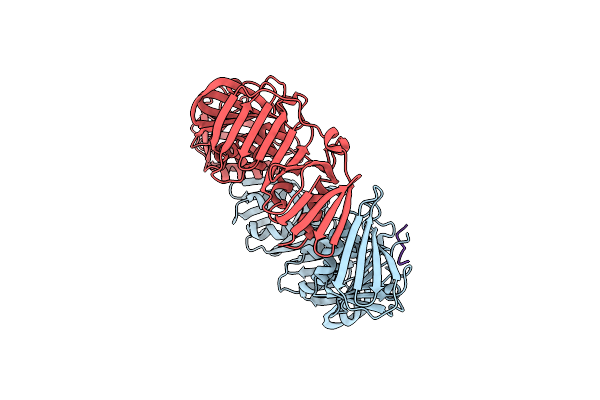 |
Crystal Structure Of E. Coli Sliding Clamp (Beta) Bound To A Polymerase Ii Peptide
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2008-07-29 Classification: TRANSFERASE, TRANSCRIPTION |
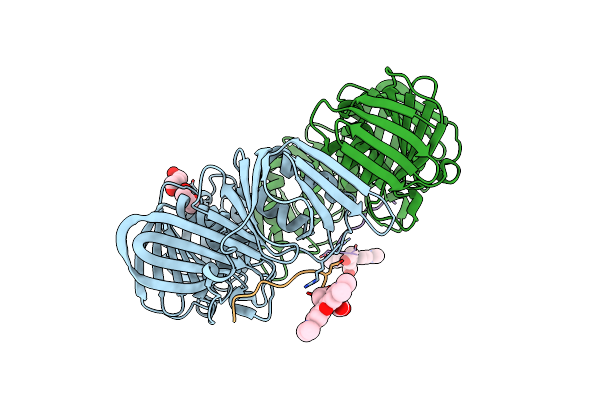 |
Crystal Structure Of E. Coli Sliding Clamp (Beta) Bound To A Polymerase Iii Peptide
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2008-07-29 Classification: TRANSFERASE, TRANSCRIPTION Ligands: PEG, 323 |
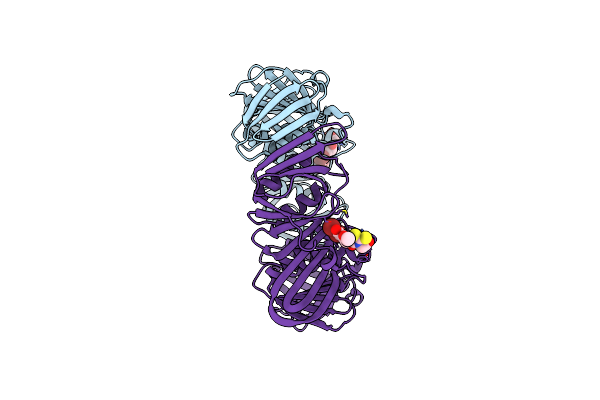 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2008-07-29 Classification: TRANSFERASE, TRANSCRIPTION Ligands: 322 |
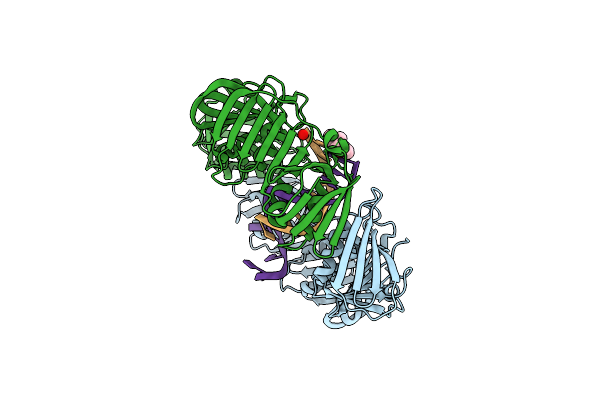 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.92 Å Release Date: 2008-01-29 Classification: TRANSFERASE, TRANSCRIPTION/DNA Ligands: 5CY |
 |
Mechanism Of Processivity Clamp Opening By The Delta Subunit Wrench Of The Clamp Loader Complex Of E. Coli Dna Polymerase Iii: Structure Of The Beta-Delta Complex
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2001-11-07 Classification: TRANSFERASE |
 |
Mechanism Of Processivity Clamp Opening By The Delta Subunit Wrench Of The Clamp Loader Complex Of E. Coli Dna Polymerase Iii: Structure Of Beta-Delta (1-140)
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2001-09-26 Classification: TRANSFERASE |

