Search Count: 9
All
Selected
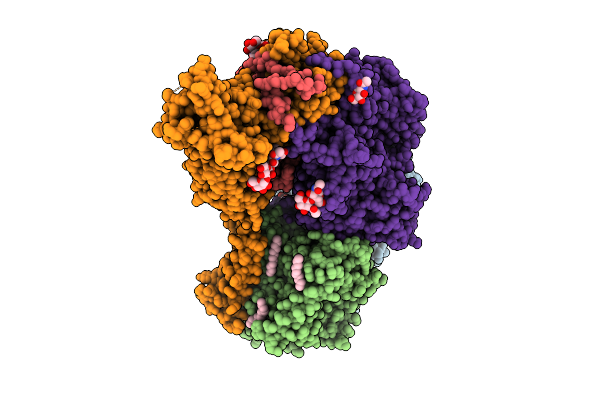 |
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN Ligands: Y01, XKP, C14, D12, D10, A1EOT, NAG |
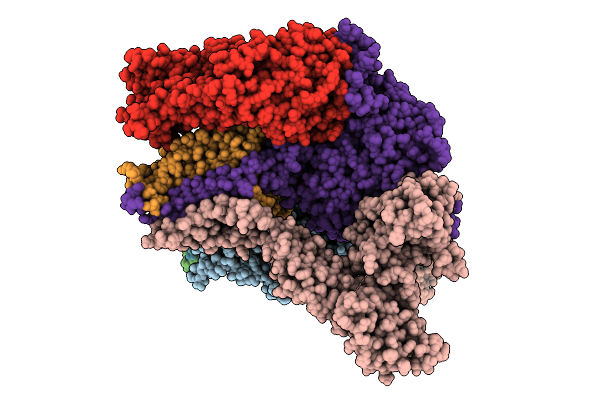 |
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c)
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN |
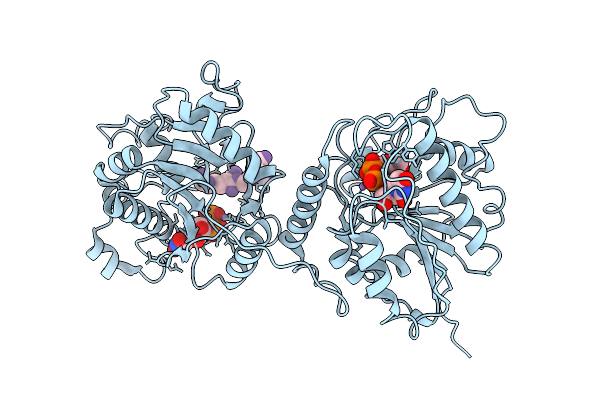 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.08 Å Release Date: 2026-04-01 Classification: TRANSFERASE Ligands: GDU, MN, UDP |
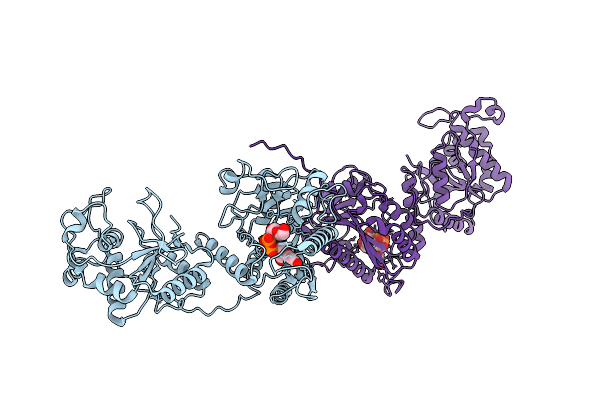 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2026-04-01 Classification: TRANSFERASE Ligands: GDU, MN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: TRANSFERASE Ligands: GDU, MN |
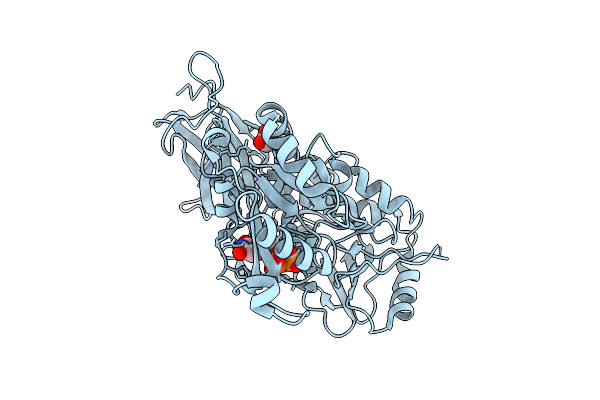 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-03-11 Classification: TRANSFERASE Ligands: UDP, GOL |
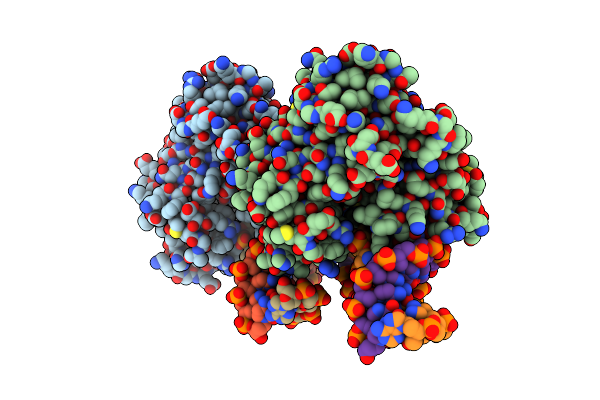 |
Organism: Thermoanaerobacter tengcongensis
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2013-03-20 Classification: HYDROLASE/DNA |
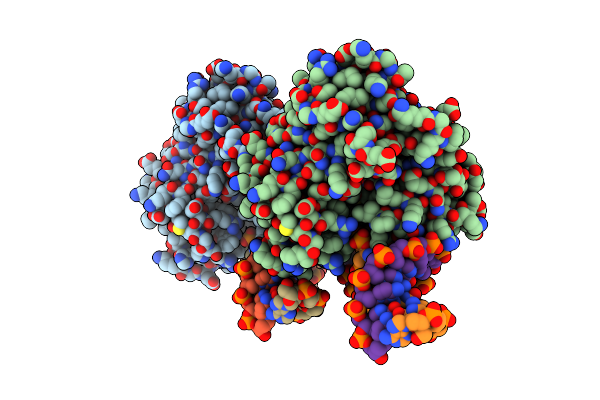 |
Organism: Thermoanaerobacter tengcongensis
Method: X-RAY DIFFRACTION Resolution:3.05 Å Release Date: 2013-03-20 Classification: HYDROLASE/DNA |
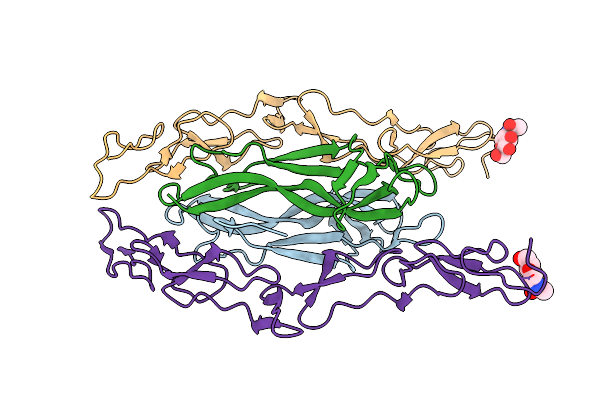 |
Organism: Homo sapiens, Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2008-07-15 Classification: SIGNALING PROTEIN Ligands: NAG |

