Search Count: 62
All
Selected
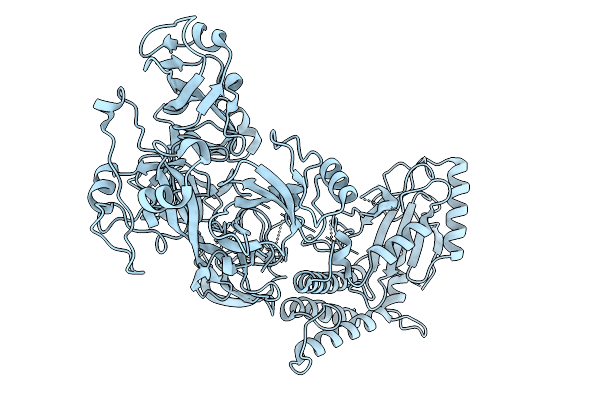 |
Organism: Pyrenophora tritici-repentis
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2026-01-21 Classification: LIGASE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-01-21 Classification: LIGASE Ligands: GOL, P4K |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Resolution:4.50 Å Release Date: 2025-12-10 Classification: LIGASE Ligands: ATP, MG, ZN |
 |
Cryo-Em Structure Of Mouse Rnf213:Ube2L3 Transthiolation Intermediate, Chemically Stabilized, And Atpgs
Organism: Mus musculus, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.80 Å Release Date: 2025-12-10 Classification: LIGASE Ligands: ATP, MG, ZN, ADP |
 |
Structure Of The 50S Ribosomal Subunit From The Antibiotic-Producing Bacterium Streptomyces Fradiae
Organism: Streptomyces fradiae atcc 10745 = dsm 40063
Method: ELECTRON MICROSCOPY Release Date: 2025-05-14 Classification: RIBOSOME |
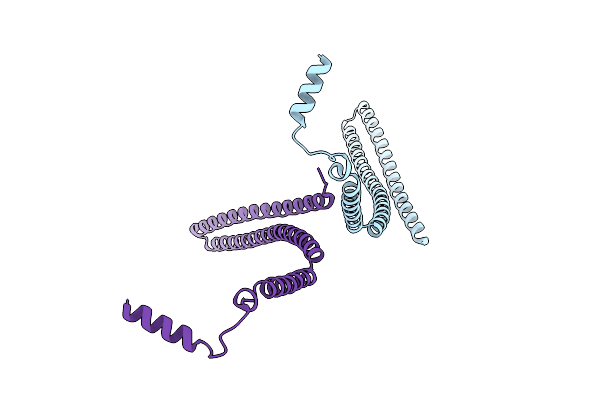 |
Organism: Candidatus heimdallarchaeota archaeon
Method: ELECTRON MICROSCOPY Release Date: 2025-04-16 Classification: MEMBRANE PROTEIN |
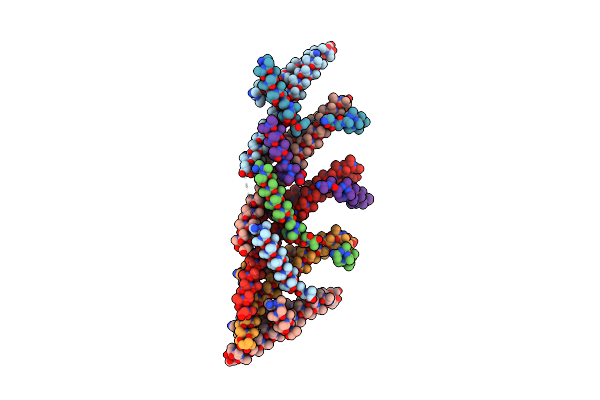 |
Organism: Candidatus heimdallarchaeota archaeon
Method: ELECTRON MICROSCOPY Release Date: 2025-04-16 Classification: MEMBRANE PROTEIN |
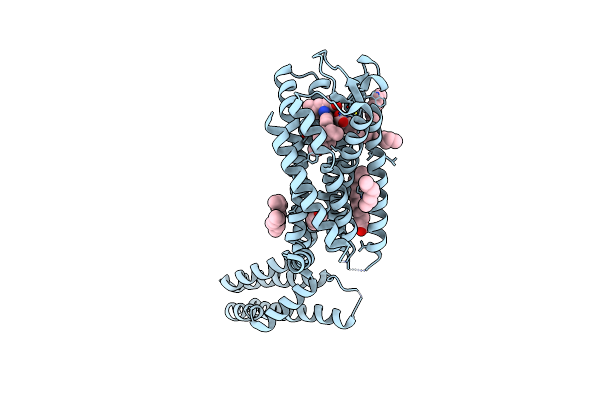 |
Crystal Structure Of A Peptidergic Gpcr In Complex With A Small Synthetic G Protein-Biased Agonist
Organism: Homo sapiens, Escherichia coli o139:h28 str. e24377a
Method: X-RAY DIFFRACTION Resolution:2.58 Å Release Date: 2024-12-04 Classification: MEMBRANE PROTEIN Ligands: A1D5N, OLA, 1PE |
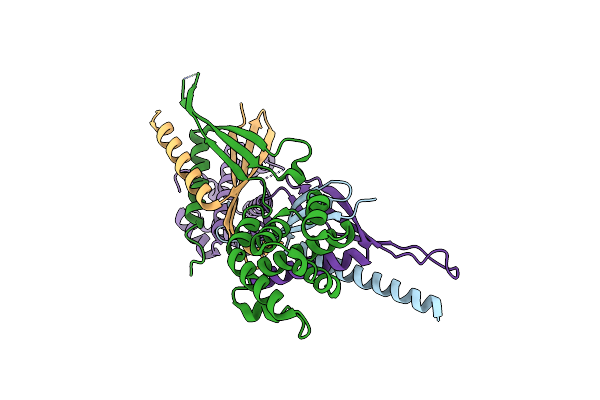 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.33 Å Release Date: 2023-05-24 Classification: ENDOCYTOSIS |
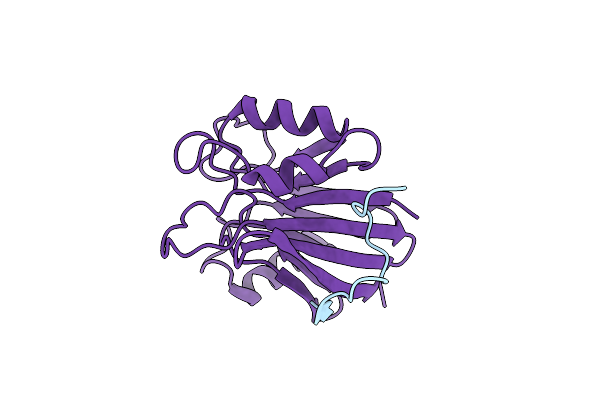 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2023-05-24 Classification: ENDOCYTOSIS |
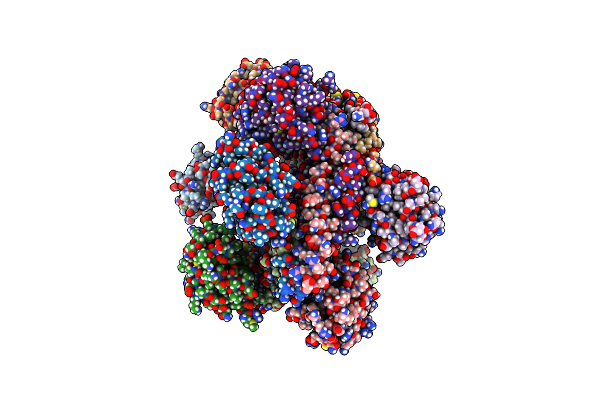 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-05-24 Classification: ENDOCYTOSIS |
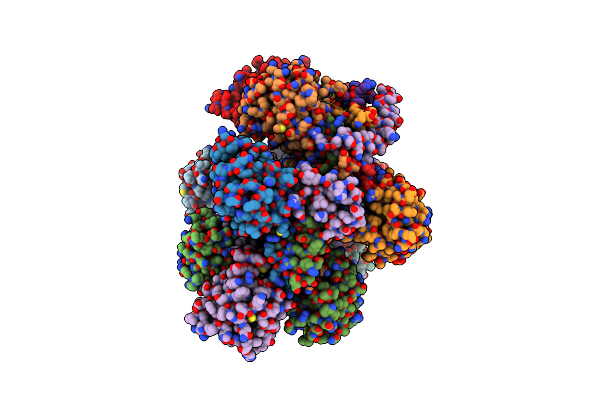 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-05-24 Classification: ENDOCYTOSIS |
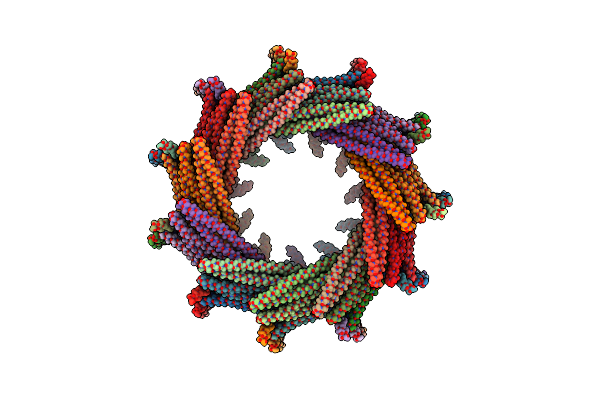 |
C11 Symmetry: Bacterial Vipp1 And Pspa Are Members Of The Ancient Escrt-Iii Membrane-Remodeling Superfamily.
Organism: Nostoc punctiforme
Method: ELECTRON MICROSCOPY Release Date: 2021-08-04 Classification: LIPID BINDING PROTEIN |
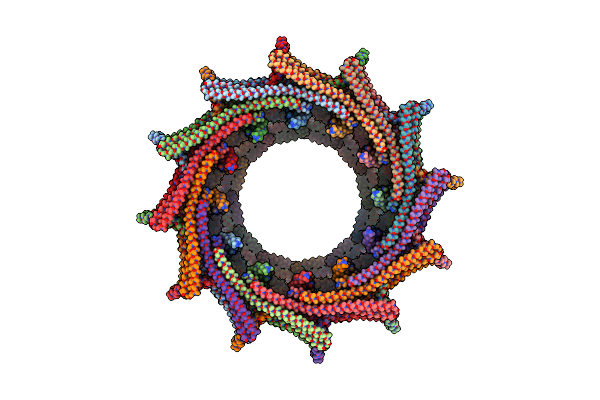 |
C12 Symmetry: Bacterial Vipp1 And Pspa Are Members Of The Ancient Escrt-Iii Membrane-Remodeling Superfamily.
Organism: Nostoc punctiforme
Method: ELECTRON MICROSCOPY Release Date: 2021-08-04 Classification: LIPID BINDING PROTEIN |
 |
C13 Symmetry: Bacterial Vipp1 And Pspa Are Members Of The Ancient Escrt-Iii Membrane-Remodeling Superfamily.
Organism: Nostoc punctiforme
Method: ELECTRON MICROSCOPY Release Date: 2021-08-04 Classification: LIPID BINDING PROTEIN |
 |
C14 Symmetry: Bacterial Vipp1 And Pspa Are Members Of The Ancient Escrt-Iii Membrane-Remodeling Superfamily.
Organism: Nostoc punctiforme
Method: ELECTRON MICROSCOPY Release Date: 2021-08-04 Classification: LIPID BINDING PROTEIN |
 |
C15 Symmetry: Bacterial Vipp1 And Pspa Are Members Of The Ancient Escrt-Iii Membrane-Remodeling Superfamily.
Organism: Nostoc punctiforme
Method: ELECTRON MICROSCOPY Release Date: 2021-08-04 Classification: LIPID BINDING PROTEIN |
 |
C16 Symmetry: Bacterial Vipp1 And Pspa Are Members Of The Ancient Escrt-Iii Membrane-Remodeling Superfamily.
Organism: Nostoc punctiforme
Method: ELECTRON MICROSCOPY Release Date: 2021-08-04 Classification: LIPID BINDING PROTEIN |
 |
C17 Symmetry: Bacterial Vipp1 And Pspa Are Members Of The Ancient Escrt-Iii Membrane-Remodeling Superfamily.
Organism: Nostoc punctiforme
Method: ELECTRON MICROSCOPY Release Date: 2021-08-04 Classification: LIPID BINDING PROTEIN |
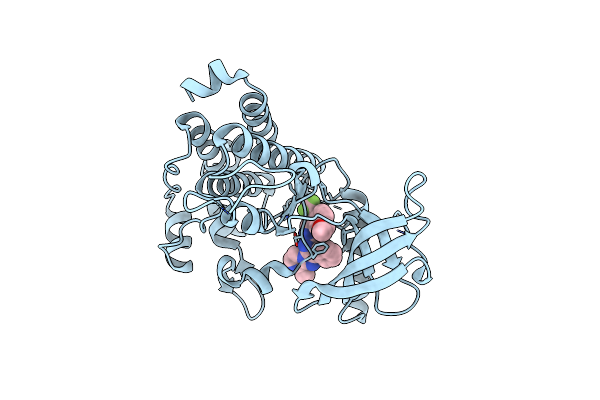 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.06 Å Release Date: 2020-07-01 Classification: TRANSFERASE Ligands: OO7 |

