Search Count: 124
All
Selected
 |
Crystal Structure Of Cytochrome P450 Mutant-T288G S289Q G290E T291I Of Cyp161H12 From Amycolatopsis Pretoriensis
Organism: Amycolatopsis pretoriensis
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-03-25 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM |
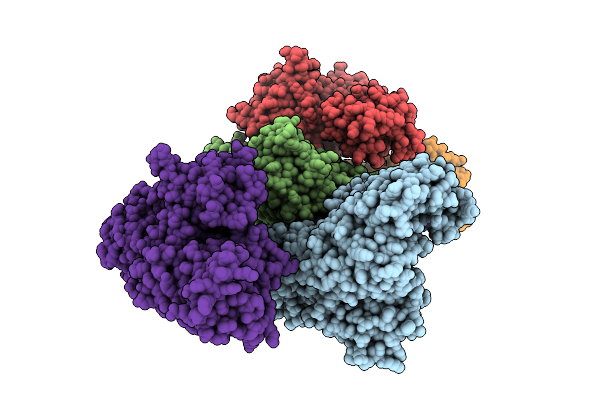 |
Crystal Structure Of A Wild-Type Tagose Isomerase (Tst4Ease Wt) From Thermotogota Bacterium
Organism: Thermotogota bacterium
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2025-12-24 Classification: ISOMERASE |
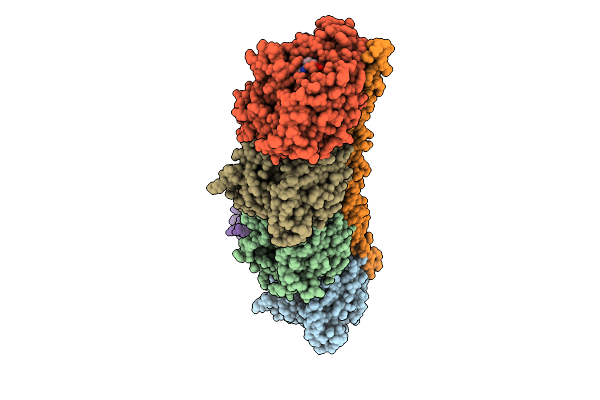 |
Organism: Sus scrofa, Bos taurus, Rattus norvegicus, Gallus gallus
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2025-11-05 Classification: STRUCTURAL PROTEIN Ligands: GTP, CA, MG, A1EE1, GDP, MES, ACP |
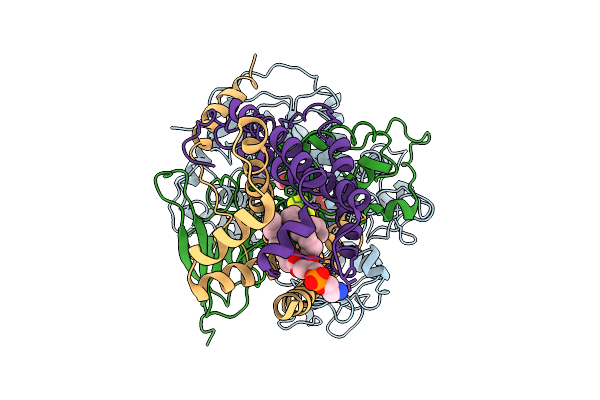 |
Cryo-Em Structure Of Saccharomyces Cerevisiae Mitochondrial Respiratory Complex Ii
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Resolution:3.36 Å Release Date: 2025-09-03 Classification: MEMBRANE PROTEIN Ligands: FAD, FES, SF4, F3S, PEE |
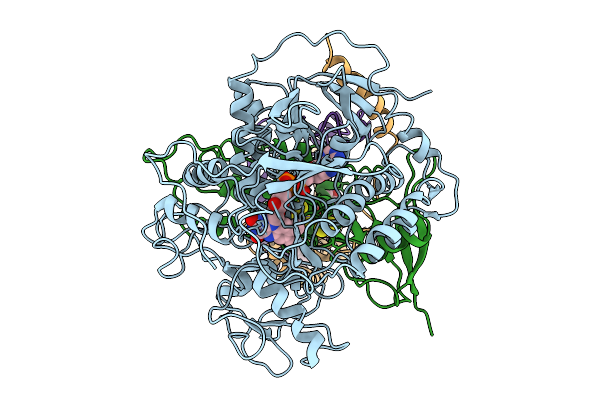 |
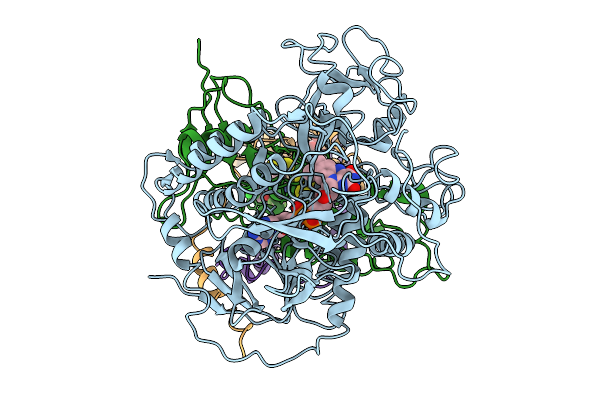
Cryo-Em Structure Of Saccharomyces Cerevisiae Mitochondrial Respiratory Complex Ii In Uq1-Bound State
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2025-09-03 Classification: MEMBRANE PROTEIN Ligands: FAD, FES, SF4, F3S, UQ1, PEE |
 |
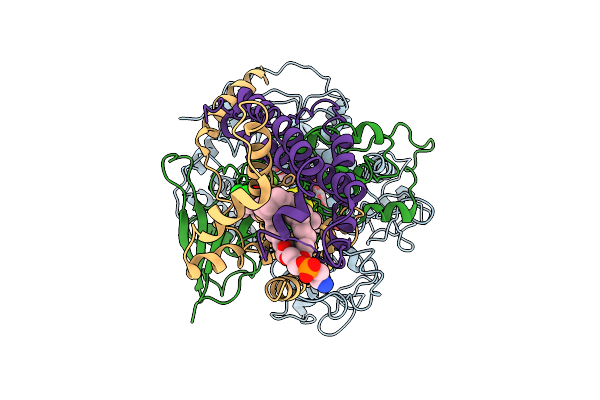
Cryo-Em Structure Of Saccharomyces Cerevisiae Mitochondrial Respiratory Complex Ii In Pydiflumetofen-Bound State
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2025-09-03 Classification: MEMBRANE PROTEIN Ligands: FAD, FES, SF4, F3S, A1EE4, PEE |
 |
Cryo-Em Structure Of Saccharomyces Cerevisiae Mitochondrial Respiratory Complex Ii In Y19315-Bound State
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2025-09-03 Classification: MEMBRANE PROTEIN Ligands: FAD, FES, SF4, F3S, A1EKV, PEE |
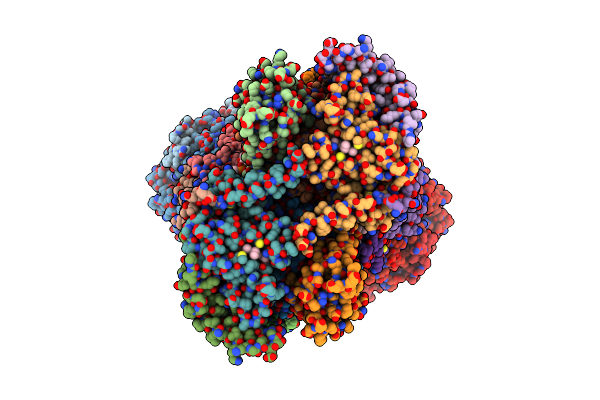 |
Organism: Arachis hypogaea
Method: ELECTRON MICROSCOPY Release Date: 2025-06-04 Classification: PLANT PROTEIN Ligands: ZN, CDL, A1D6K, HEM, 3PE, HEC, FES, PC1 |
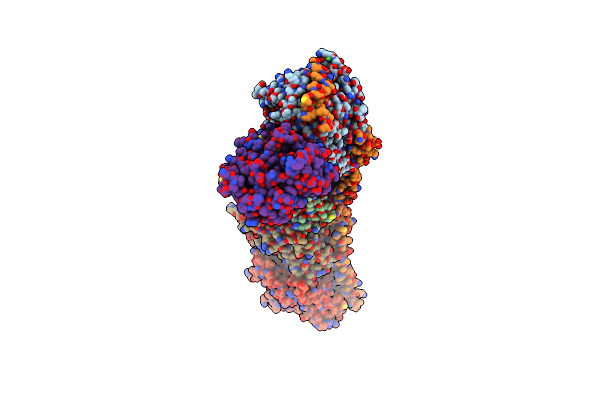 |
Organism: Sus scrofa, Bos taurus, Rattus norvegicus, Gallus gallus
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2025-05-07 Classification: STRUCTURAL PROTEIN Ligands: GTP, CA, MG, A1EPQ, GDP |
 |
Organism: Sus scrofa, Mus musculus, Gallus gallus
Method: X-RAY DIFFRACTION Resolution:2.67 Å Release Date: 2025-05-07 Classification: STRUCTURAL PROTEIN Ligands: GTP, MG, CA, A1EPR, GDP |
 |
Organism: Sus scrofa, Mus musculus, Gallus gallus
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2025-05-07 Classification: TRANSPORT PROTEIN Ligands: GTP, MG, CA, GDP, A1EPS |
 |
Organism: Sus scrofa, Mus musculus, Gallus gallus
Method: X-RAY DIFFRACTION Resolution:2.59 Å Release Date: 2025-05-07 Classification: TRANSPORT PROTEIN Ligands: GTP, MG, CA, A1EPT, GDP |
 |
Organism: Sus scrofa, Rattus norvegicus, Gallus gallus
Method: X-RAY DIFFRACTION Resolution:2.74 Å Release Date: 2025-02-26 Classification: STRUCTURAL PROTEIN Ligands: GTP, CA, MG, GDP, MES, A1D6D, ACP |
 |
Organism: Sus scrofa, Rattus norvegicus, Gallus gallus
Method: X-RAY DIFFRACTION Resolution:2.71 Å Release Date: 2025-02-26 Classification: STRUCTURAL PROTEIN Ligands: GTP, CA, MG, A1D6E, GDP, MES, ACP |
 |
Organism: Sus scrofa, Rattus norvegicus, Gallus gallus
Method: X-RAY DIFFRACTION Resolution:3.05 Å Release Date: 2025-02-26 Classification: STRUCTURAL PROTEIN Ligands: GTP, CA, MG, A1D6L, GDP, MES, ACP |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.06 Å Release Date: 2025-02-19 Classification: RNA Ligands: GTP, TAC, MG |
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-02-19 Classification: RNA Ligands: TAC, MG |
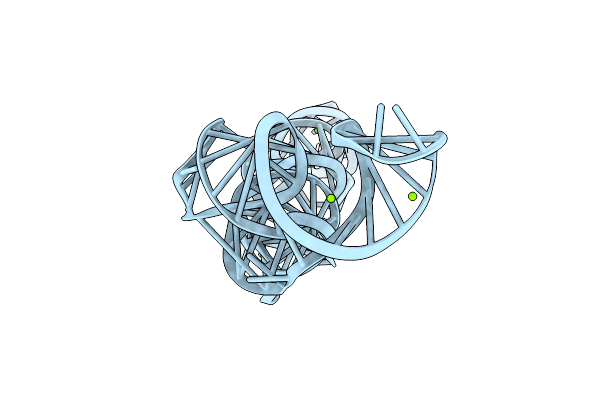 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-02-19 Classification: RNA Ligands: MG |
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-02-19 Classification: RNA Ligands: GTP, MG |
 |
Cryo-Em Structure Of Saccharomyces Cerevisiae Bc1 Complex In Pyraclostrobin-Bound State
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2024-12-25 Classification: MEMBRANE PROTEIN Ligands: 6PH, 9PE, HEM, 8PE, CN5, UQ6, A1D6K, 7PH, CN3 |

