Search Count: 54
All
Selected
 |
 |
 |
 |
 |
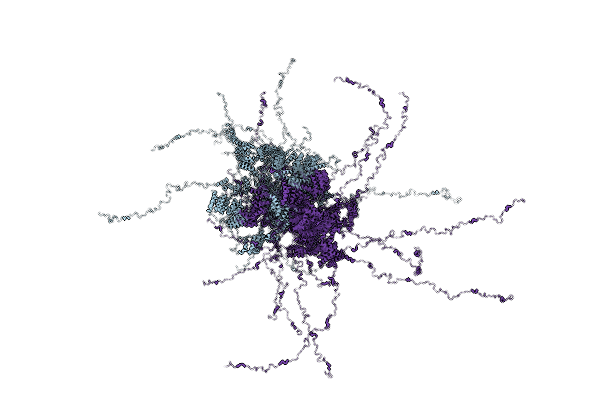 |
Organism: Saccharomyces cerevisiae s288c
Method: SOLUTION NMR Release Date: 2025-12-10 Classification: CHAPERONE |
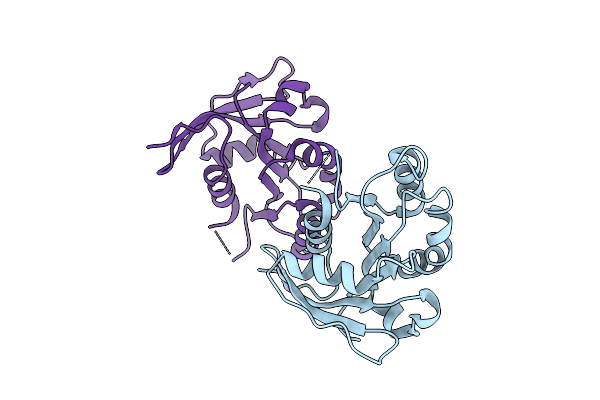 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2025-07-16 Classification: PROTEIN BINDING |
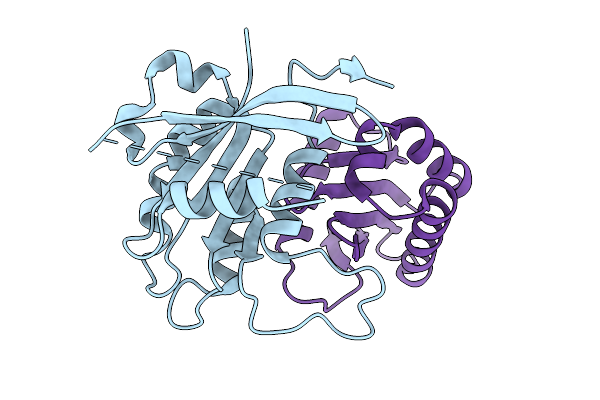 |
Aagab Pseudogtpase Domain In Complex With Ap-1 Clathrin Adaptor Complex Sigma 3 Subunit
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2025-07-16 Classification: PROTEIN BINDING |
 |
 |
 |
 |
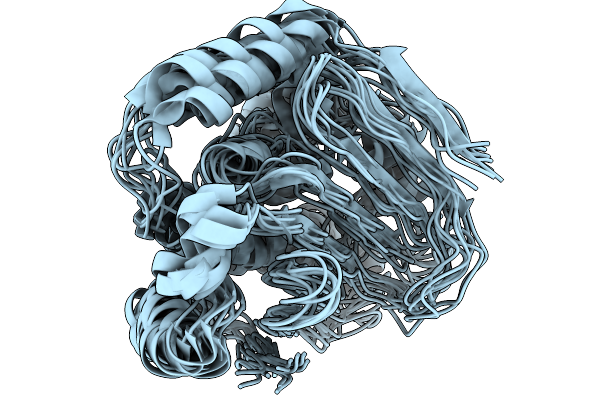
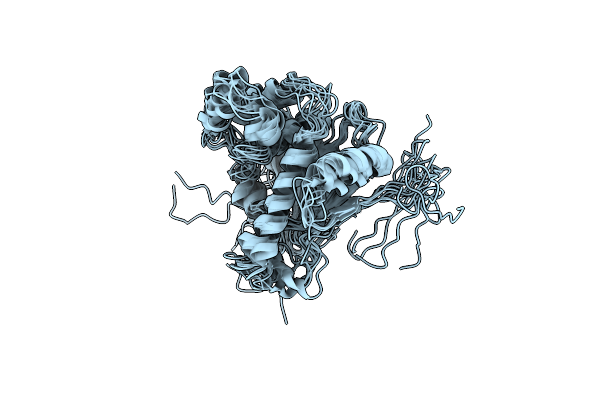
The Structure Of Htpg M Domain In Complex With Unstructured D131D Binding Site B
Organism: Escherichia coli, Staphylococcus aureus
Method: SOLUTION NMR Release Date: 2024-06-26 Classification: CHAPERONE |
 |
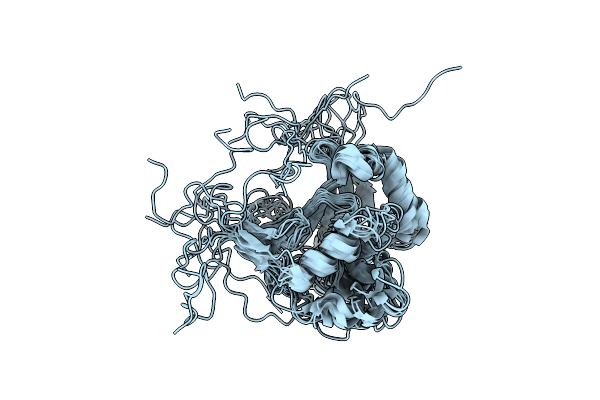
The Structure Of Htpg M Domain In Complex With Unstructured D131D Binding Site A
Organism: Escherichia coli, Staphylococcus aureus
Method: SOLUTION NMR Release Date: 2024-06-26 Classification: CHAPERONE |
 |
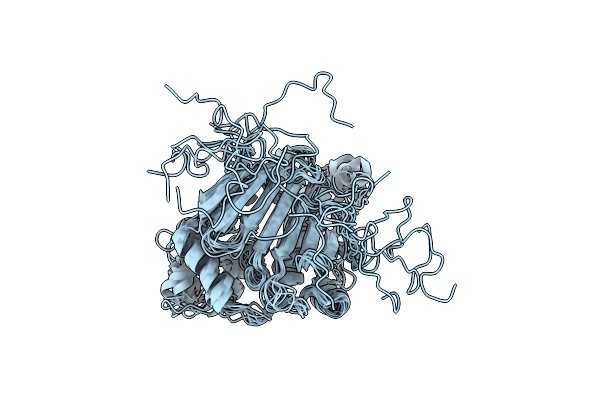
Organism: Escherichia coli, Staphylococcus aureus
Method: SOLUTION NMR Release Date: 2024-06-26 Classification: CHAPERONE |
 |
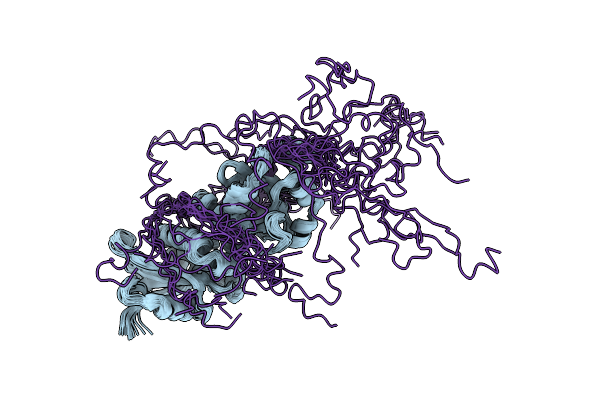
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2024-01-24 Classification: RNA |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.11 Å Release Date: 2023-01-18 Classification: CHAPERONE Ligands: PO4 |
 |
Organism: Yersinia pseudotuberculosis serotype i (strain ip32953)
Method: X-RAY DIFFRACTION Resolution:2.78 Å Release Date: 2022-12-21 Classification: TOXIN |
 |
Structure Of Candida Albicans Fructose-1,6-Bisphosphate Aldolase Complexed With G3P
Organism: Candida albicans sc5314
Method: X-RAY DIFFRACTION Resolution:2.98 Å Release Date: 2022-02-23 Classification: LYASE Ligands: ZN, G3H |
 |
Structure Of Candida Albicans Fructose-1,6-Bisphosphate Aldolase Mutation C157S With Cn39
Organism: Candida albicans sc5314
Method: X-RAY DIFFRACTION Resolution:2.34 Å Release Date: 2022-02-23 Classification: LYASE Ligands: ZN, EDO, 6Y3 |
 |
Organism: Candida albicans sc5314
Method: X-RAY DIFFRACTION Resolution:2.64 Å Release Date: 2020-12-30 Classification: LYASE Ligands: ZN, EDO |

