Search Count: 12
All
Selected
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-11-05 Classification: RIBOSOME Ligands: ZN, MG |
 |
Organism: Rhodococcus rhodochrous
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2025-11-05 Classification: BIOSYNTHETIC PROTEIN Ligands: CL, FMN, PUJ, GOL |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.18 Å Release Date: 2019-06-05 Classification: TRANSCRIPTION Ligands: 9FX |
 |
Organism: Porphyromonas gingivalis, Lama glama
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2018-02-07 Classification: PROTEIN TRANSPORT |
 |
Organism: Flavobacterium johnsoniae
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2018-02-07 Classification: PROTEIN TRANSPORT Ligands: EDO |
 |
Organism: Porphyromonas gingivalis
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2018-02-07 Classification: PROTEIN TRANSPORT Ligands: ZN |
 |
Organism: Porphyromonas gingivalis, Lama glama
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2018-02-07 Classification: PROTEIN TRANSPORT |
 |
Organism: Pectobacterium carotovorum subsp. carotovorum
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2008-12-30 Classification: LIPID BINDING PROTEIN Ligands: PLM |
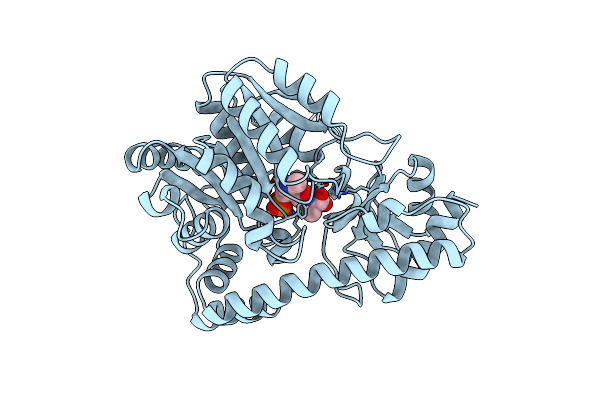 |
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 1993-10-31 Classification: TRANSFERASE(AMINOTRANSFERASE) Ligands: PLA |
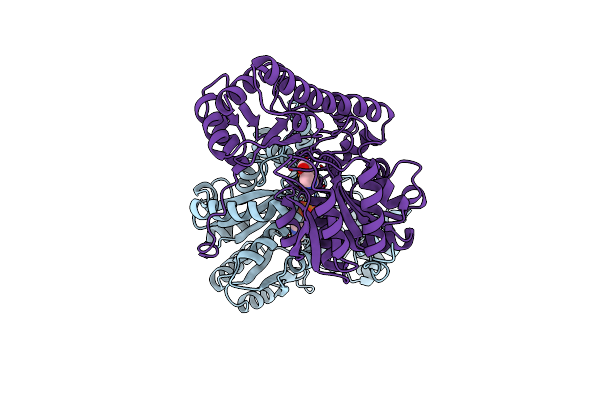 |
X-Ray Structure Refinement And Comparison Of Three Forms Of Mitochondrial Aspartate Aminotransferase
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 1993-10-31 Classification: TRANSFERASE(AMINOTRANSFERASE) Ligands: PLP |
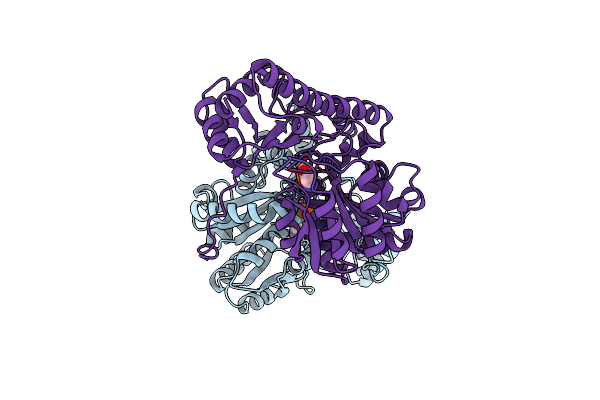 |
X-Ray Structure Refinement And Comparison Of Three Forms Of Mitochondrial Aspartate Aminotransferase
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 1993-10-31 Classification: TRANSFERASE(AMINOTRANSFERASE) Ligands: PLP |
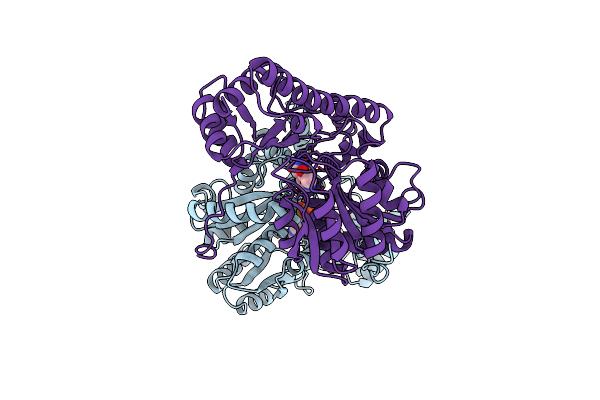 |
X-Ray Structure Refinement And Comparison Of Three Forms Of Mitochondrial Aspartate Aminotransferase
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 1993-10-31 Classification: TRANSFERASE(AMINOTRANSFERASE) Ligands: PMP |

