Search Count: 15
All
Selected
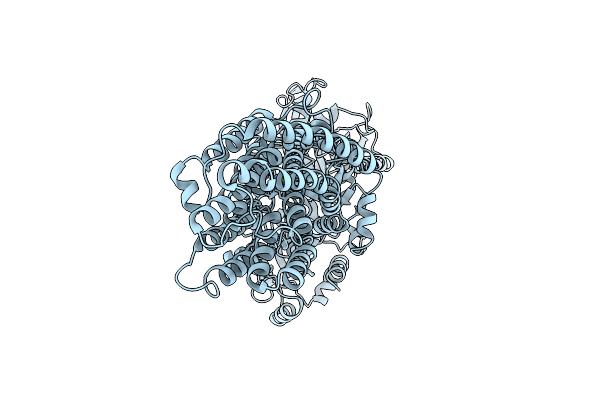 |
Cryo-Em Of Mgta In The E2-P State With Bound Mg2+ At 3.1 Angstrom Resolution
Organism: Lactococcus lactis subsp. lactis
Method: ELECTRON MICROSCOPY Resolution:3.07 Å Release Date: 2026-04-08 Classification: TRANSPORT PROTEIN Ligands: MG |
 |
Organism: Lactococcus lactis subsp. lactis
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: MG |
 |
Cryo-Em Structure Of The Magnesium Transporter Mgta In E1-Like And E2-P Conformations At 2.59 Angstroms
Organism: Lactococcus lactis subsp. lactis
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: TRANSPORT PROTEIN Ligands: MG, BEF |
 |
Cryo-Em Structure Of The Magnesium Transporter Mgta In The E2 Conformation With Bound Beryllium Fluoride (Bef3-) And Mg2+ At 2.65 A Resolution.
Organism: Lactococcus lactis subsp. lactis
Method: ELECTRON MICROSCOPY Resolution:2.59 Å Release Date: 2026-02-18 Classification: TRANSPORT PROTEIN Ligands: MG, BEF |
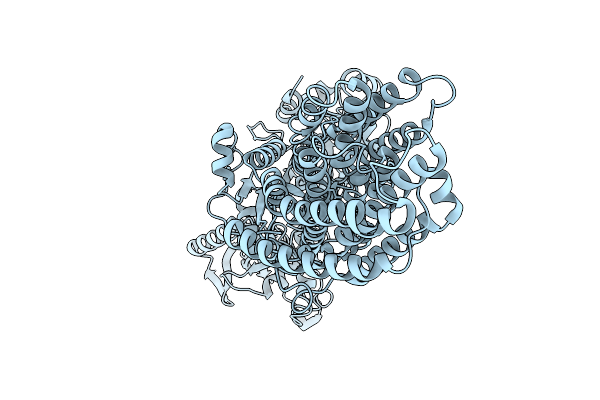 |
Cryo-Em Structure Of The Magnesium Transporter Mgta In The E2 Conformation Bound To Mg2+
Organism: Lactococcus lactis subsp. lactis
Method: ELECTRON MICROSCOPY Resolution:3.23 Å Release Date: 2026-01-21 Classification: TRANSPORT PROTEIN Ligands: MG |
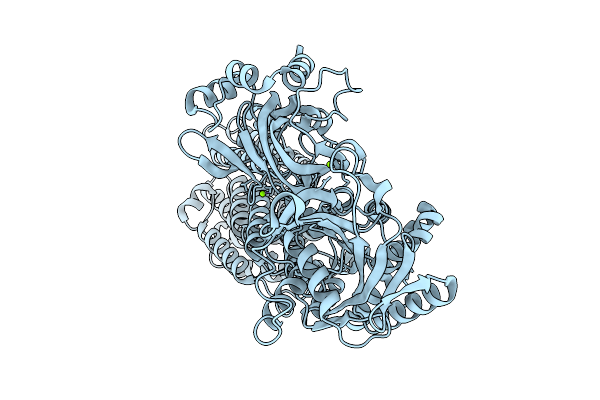 |
Cryo-Em Structure Of The Magnesium Transporter Mgta In An E1-Like Conformation With Bound Mg2+ Ions
Organism: Lactococcus lactis subsp. lactis
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2025-12-10 Classification: TRANSPORT PROTEIN Ligands: MG |
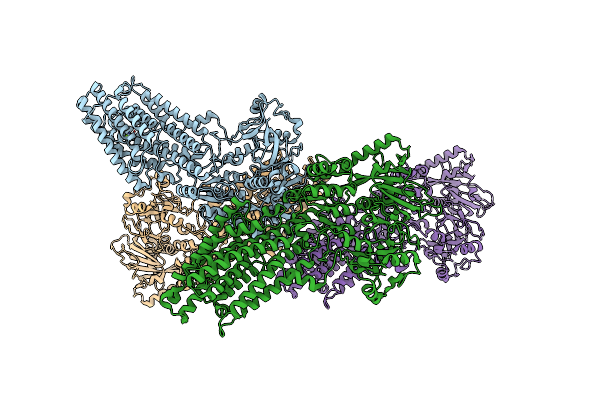 |
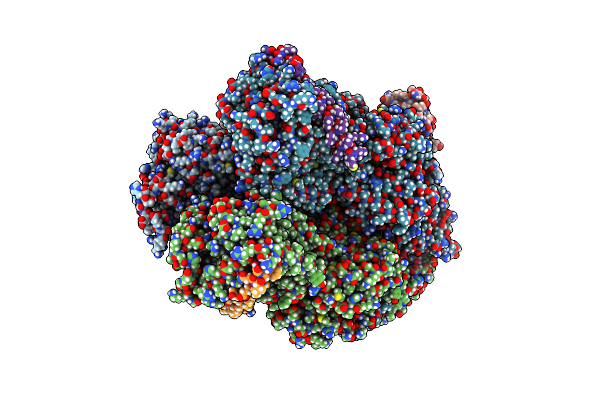
Crystal Structure Of Magnesium-Transporting Atpase Mgta In An E1-Like Magnesium-Bound State
Organism: Lactococcus lactis subsp. lactis cv56
Method: X-RAY DIFFRACTION Resolution:3.55 Å Release Date: 2025-02-26 Classification: METAL TRANSPORT Ligands: MG |
 |
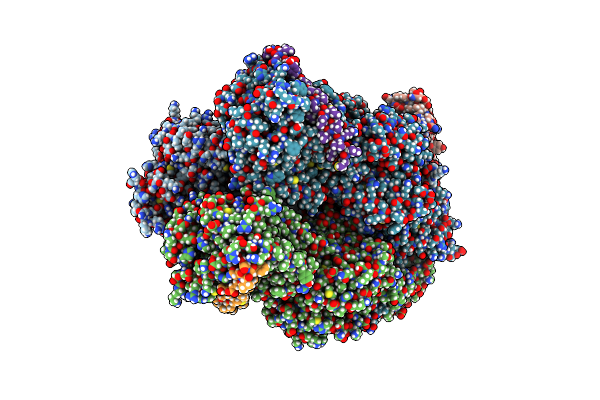
Cryo-Em Structure Of Escherichia Coli Acrbz And Darpin In Saposin A-Nanodisc With Cardiolipin
Organism: Escherichia coli k-12, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2020-05-13 Classification: MEMBRANE PROTEIN |
 |
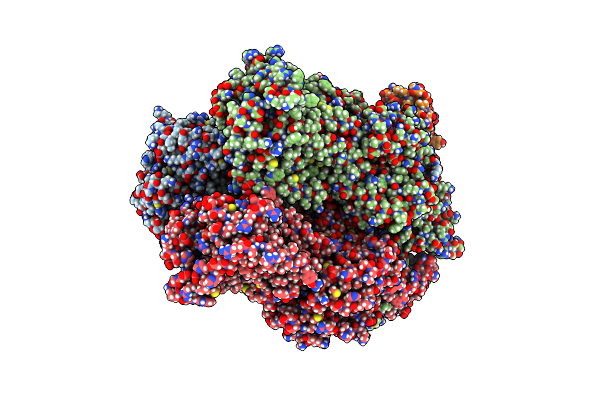
Cryo-Em Structure Of Escherichia Coli Acrbz And Darpin In Saposin A-Nanodisc
Organism: Escherichia coli k12, Synthetic construct, Shigella boydii 965-58
Method: ELECTRON MICROSCOPY Release Date: 2020-05-13 Classification: MEMBRANE PROTEIN |
 |
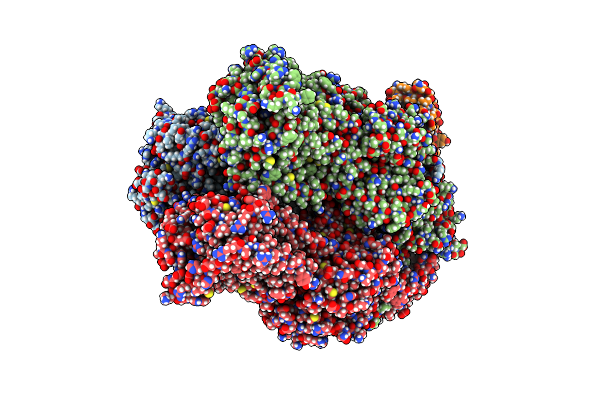
Cryo-Em Structure Of Escherichia Coli Acrb And Darpin In Saposin A-Nanodisc With Cardiolipin
Organism: Escherichia coli k12, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2020-05-13 Classification: MEMBRANE PROTEIN |
 |
Cryo-Em Structure Of Escherichia Coli Acrb And Darpin In Saposin A-Nanodisc
Organism: Escherichia coli k12, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2020-05-13 Classification: MEMBRANE PROTEIN |
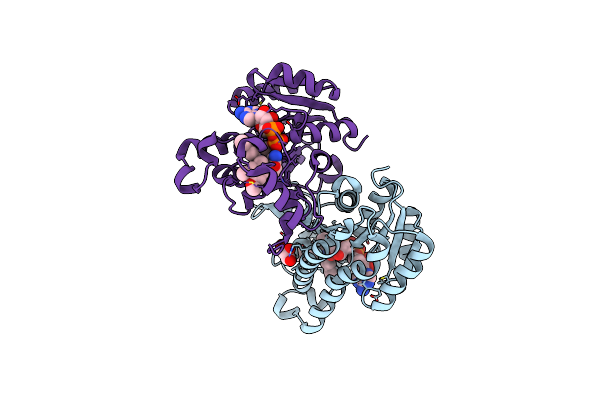 |
Crystal Structures Of Fabi From F. Tularensis In Complex With Novel Inhibitors Based On The Benzimidazole Scaffold
Organism: Francisella tularensis subsp. tularensis
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2014-07-23 Classification: OXIDOREDUCTASE/OXIDOREDUCTASE INHIBITOR Ligands: NAD, 1JN, GOL |
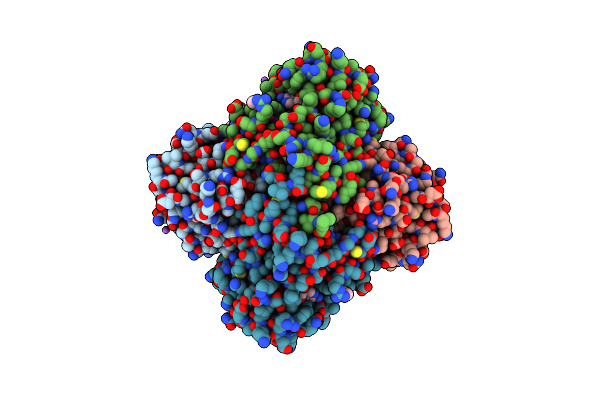 |
Crystal Structure Of Fabi From F. Tularensis In Complex With Novel Inhibitors Based On The Benzimidazole Scaffold.
Organism: Francisella tularensis subsp. tularensis
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2014-07-23 Classification: OXIDOREDUCTASE/OXIDOREDUCTASE INHIBITOR Ligands: 1JT, NAD, GOL, ACT, NA |
 |
Crystal Structure Of Fabi From F. Tularensis In Complex With Novel Inhibitors Based On The Benzimidazole Scaffold
Organism: Francisella tularensis subsp. tularensis
Method: X-RAY DIFFRACTION Resolution:2.34 Å Release Date: 2014-07-23 Classification: OXIDOREDUCTASE/OXIDOREDUCTASE INHIBITOR Ligands: NAD, 1JU, GOL, NA, ACT |
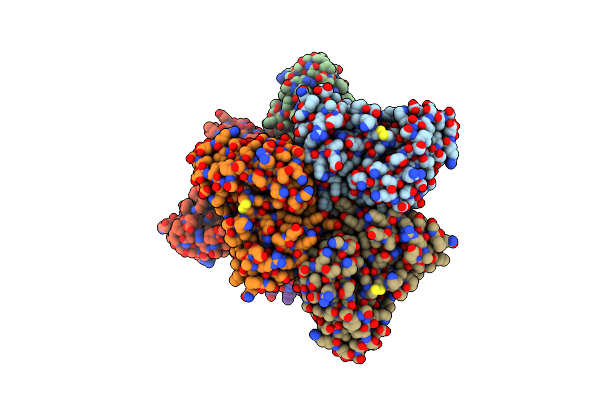 |
Predicting Amino Acid Preferences In The Complementarity Determining Regions Of An Antibody-Antigen Recognition Interface
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2012-02-22 Classification: IMMUNE SYSTEM |

