Search Count: 77
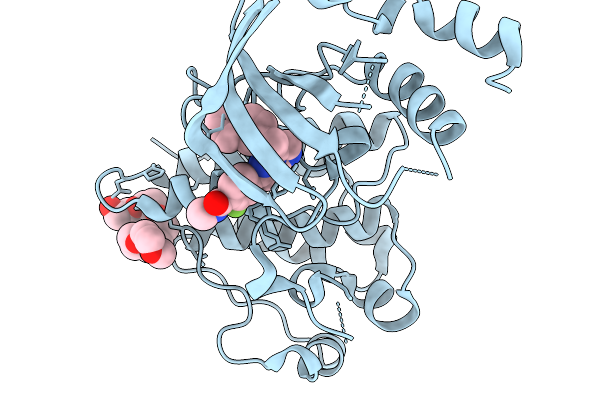 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.32 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: A1JRF, 12P |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.29 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: A1JRF |
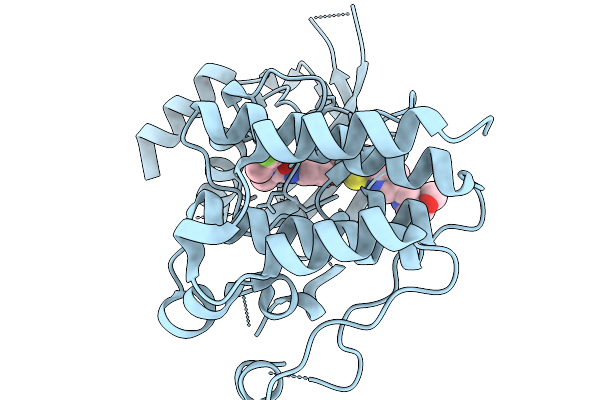 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.46 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: A1JSO |
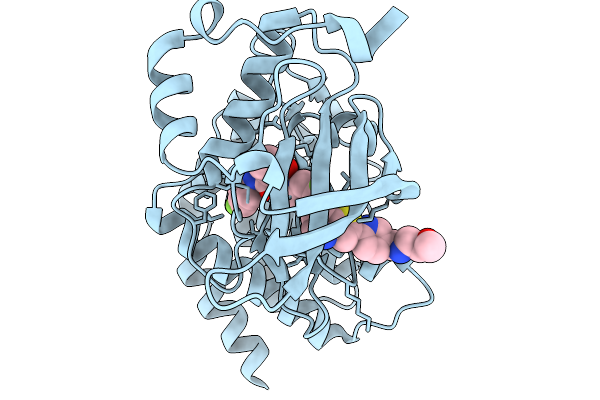 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.54 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: A1JSO |
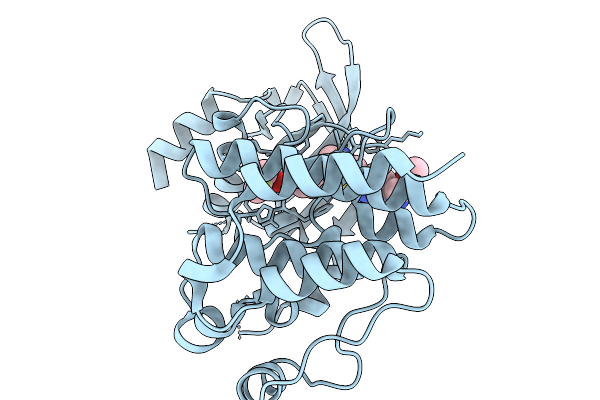 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: A1JSR |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: A1JSR |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: A1JS8 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: A1JS8 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.13 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: A1JT6, 15P, PEG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2025-12-24 Classification: HYDROLASE Ligands: A1JH5 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.51 Å Release Date: 2025-12-24 Classification: HYDROLASE Ligands: A1JKT, EDO |
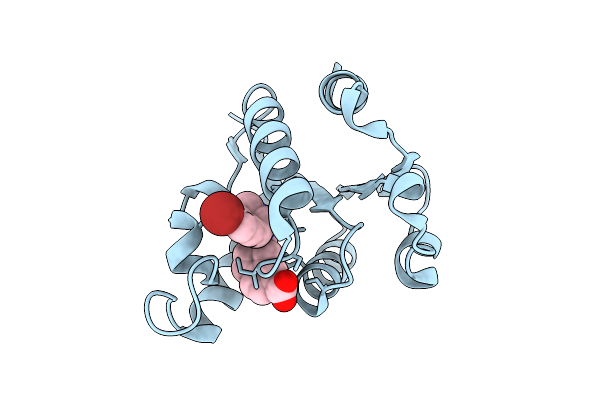 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-12-24 Classification: HYDROLASE Ligands: A1JKZ |
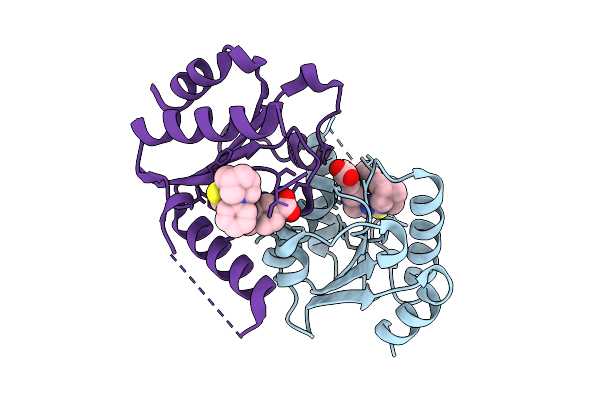 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2025-12-24 Classification: HYDROLASE Ligands: CL, A1JK6 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: A1IZR |
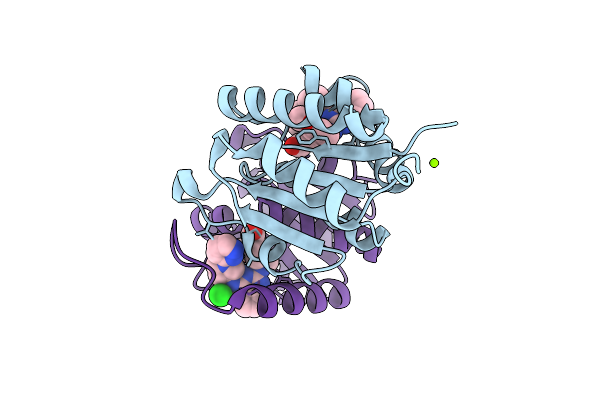 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.22 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: MG, A1IZW, CL |
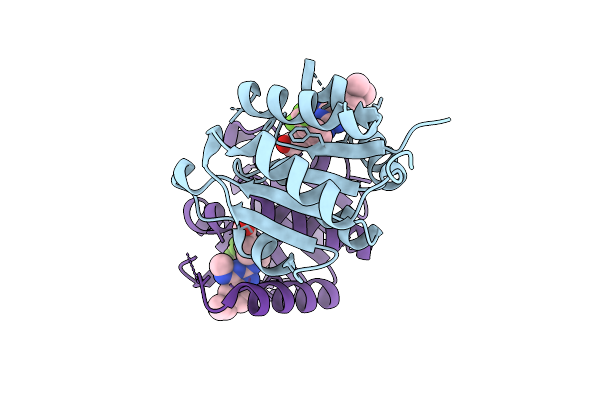 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.21 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: A1IZX |
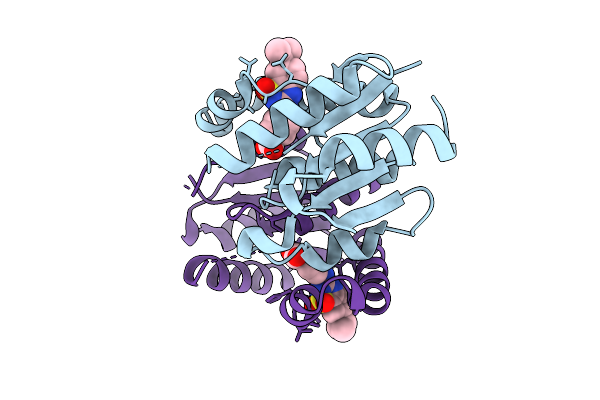 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: A1I0K |
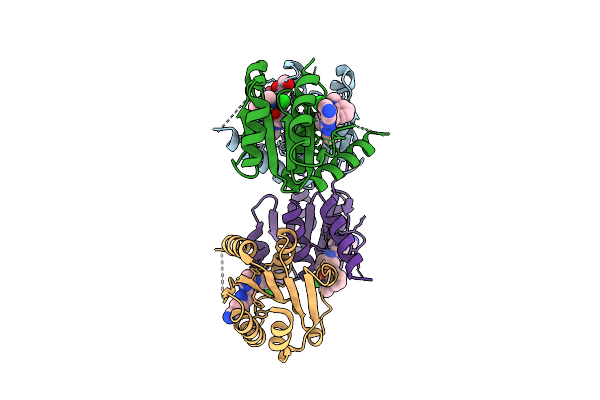 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: EOH, EDO, A1I1M, CL |
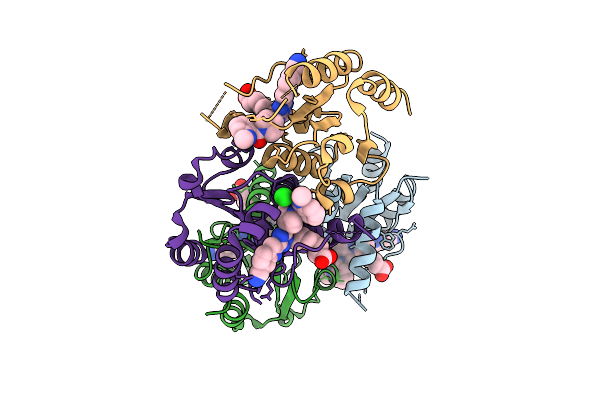 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: CL, A1I1U |
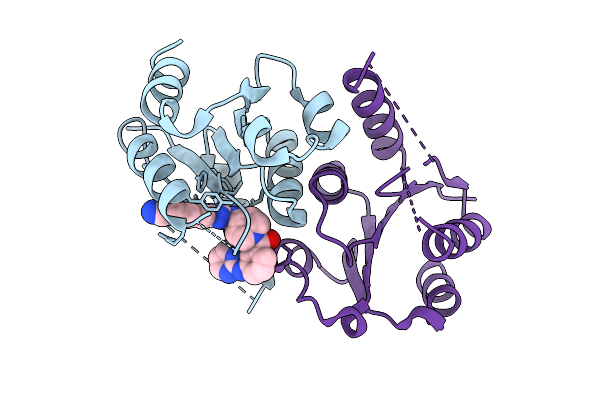 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.07 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: A1JFX |

