Search Count: 25
All
Selected
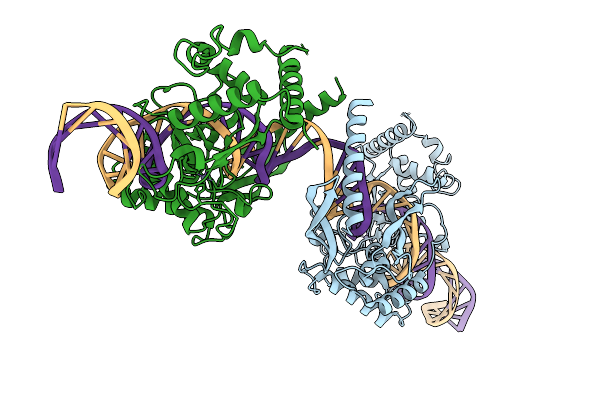 |
Organism: Streptomyces phage phi-c31, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN Ligands: ZN |
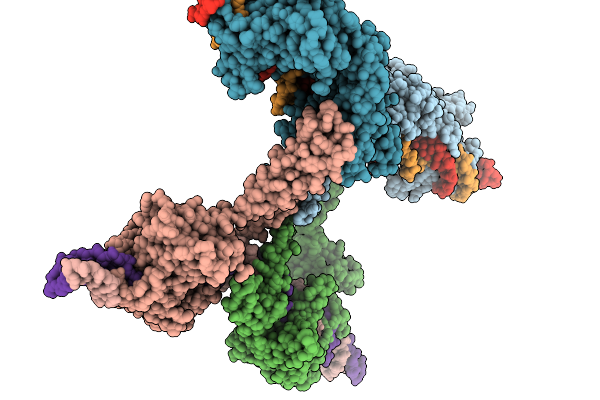 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: DNA BINDING PROTEIN Ligands: ZN |
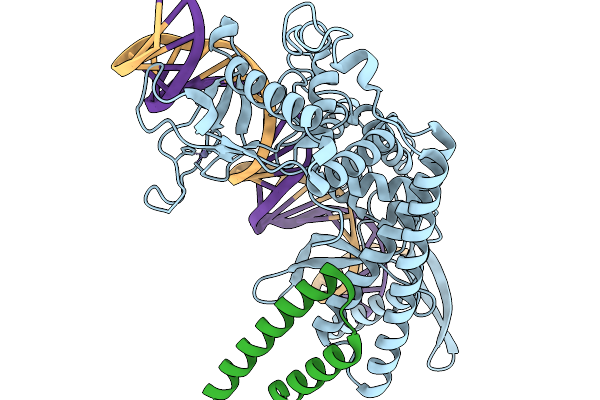 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: DNA BINDING PROTEIN Ligands: ZN |
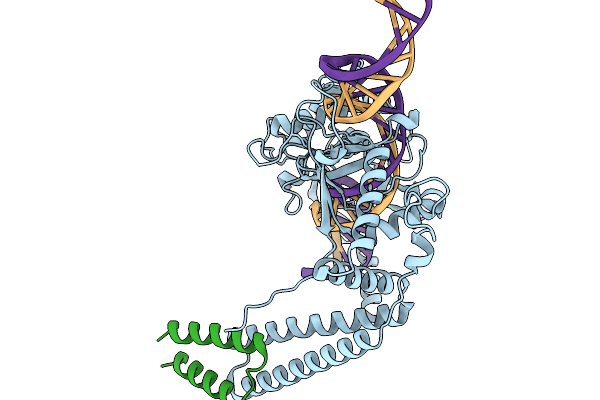 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: DNA BINDING PROTEIN Ligands: ZN |
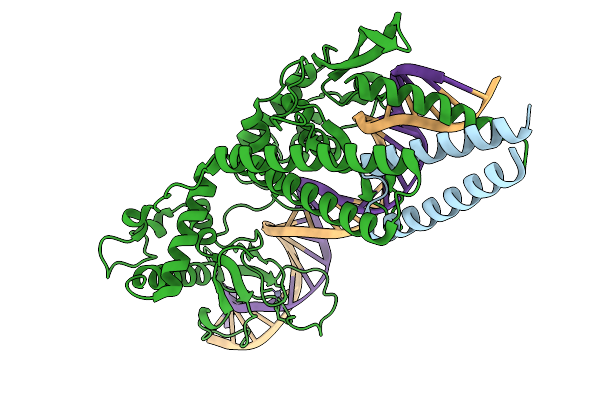 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: DNA BINDING PROTEIN Ligands: ZN |
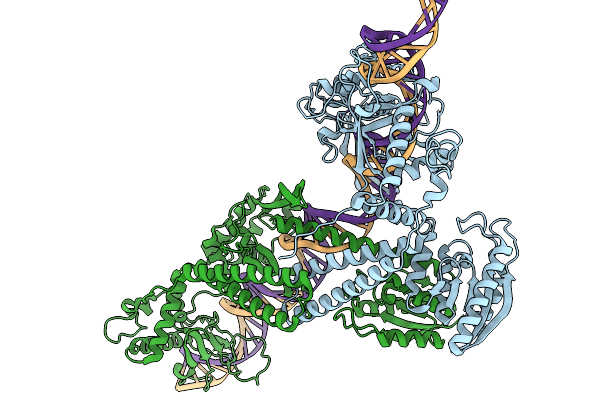 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: DNA BINDING PROTEIN Ligands: ZN |
 |
Phic31 Excisive Synapse; Attl X Attl With An Integrases12A-Rdf Protein Fusion
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: RECOMBINATION Ligands: ZN |
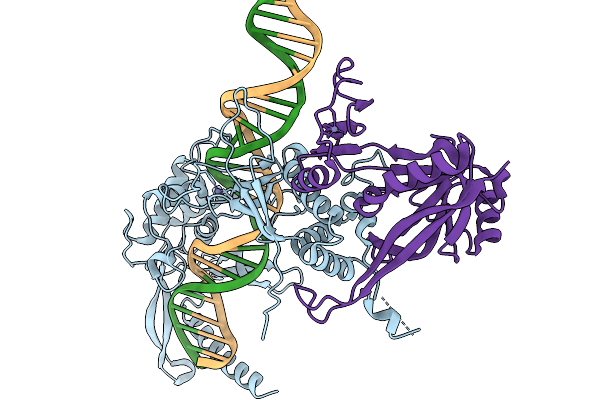 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: RECOMBINATION Ligands: ZN |
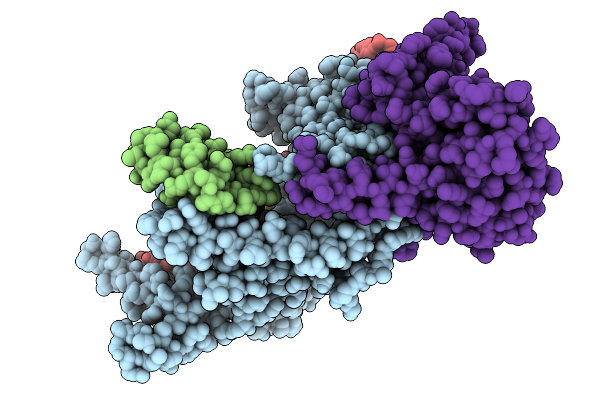 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: RECOMBINATION Ligands: ZN |
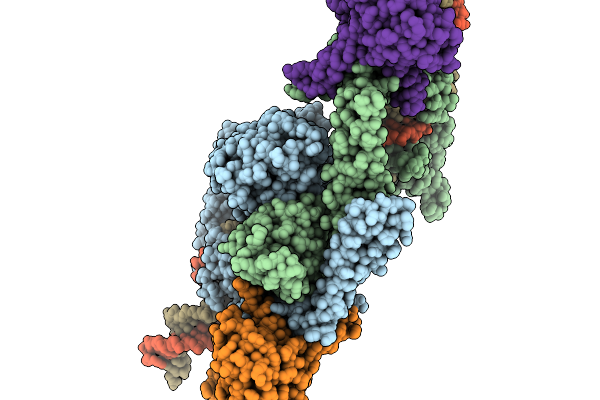 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: RECOMBINATION Ligands: ZN |
 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: RECOMBINATION Ligands: ZN |
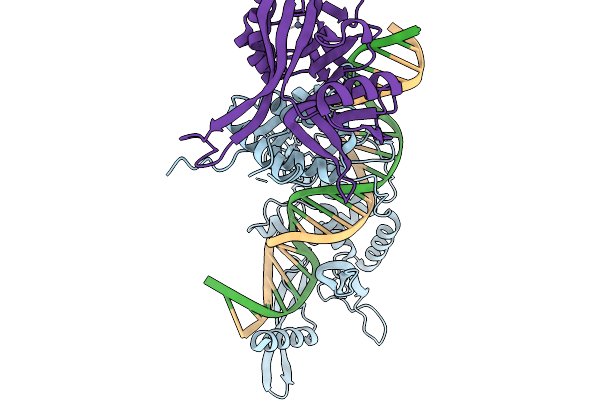 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: RECOMBINATION Ligands: ZN |
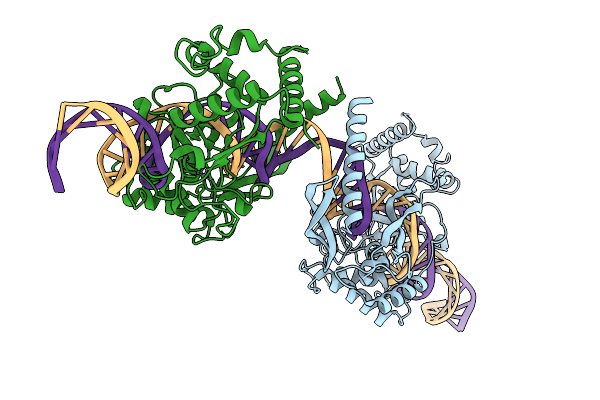 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: RECOMBINATION Ligands: ZN |
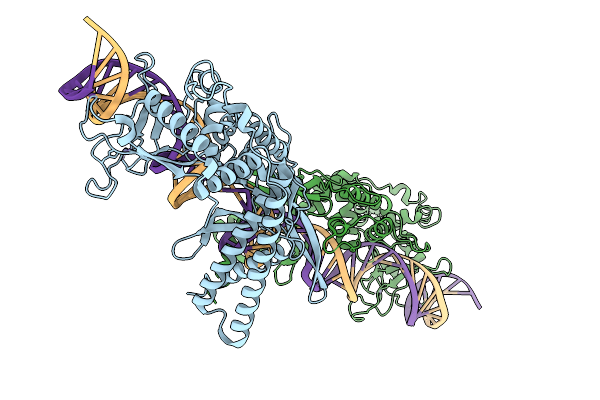 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: RECOMBINATION Ligands: ZN |
 |
Organism: Saccharolobus solfataricus
Method: ELECTRON MICROSCOPY Release Date: 2023-03-01 Classification: RNA BINDING PROTEIN |
 |
Organism: Microbacterium ketosireducens
Method: ELECTRON MICROSCOPY Release Date: 2022-06-15 Classification: SIGNALING PROTEIN |
 |
Crystal Structure Of Malonyl-Coa Decarboxylase (Rmet_2797) From Cupriavidus Metallidurans, Northeast Structural Genomics Consortium Target Crr76
Organism: Cupriavidus metallidurans
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2013-06-19 Classification: LYASE Ligands: MG |
 |
Crystal Structure Of Malonyl-Coa Decarboxylase From Rhodopseudomonas Palustris, Northeast Structural Genomics Consortium Target Rpr127
Organism: Rhodopseudomonas palustris
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2013-06-19 Classification: LYASE Ligands: MG |
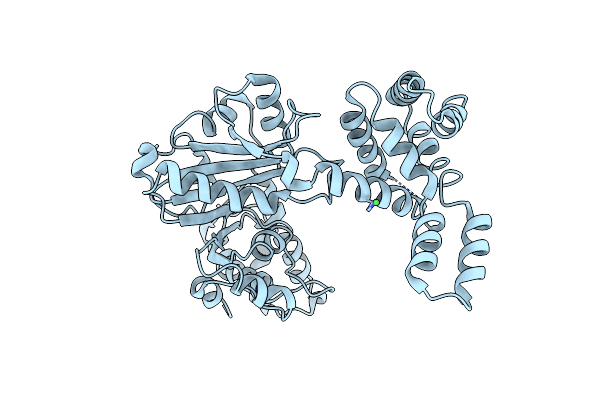 |
Crystal Structure Of Malonyl-Coa Decarboxylase From Agrobacterium Vitis, Northeast Structural Genomics Consortium Target Rir35
Organism: Agrobacterium vitis
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2013-06-19 Classification: LYASE Ligands: NI, CL |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2012-02-15 Classification: LYASE Ligands: UNX, EDO |

