Search Count: 18
All
Selected
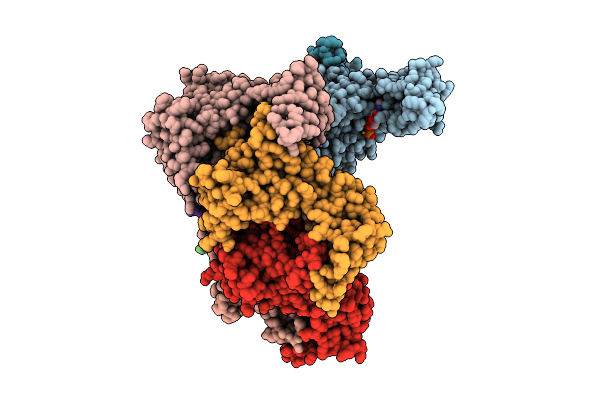 |
Heptamer Msp1 From S.Cerevisiae (With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
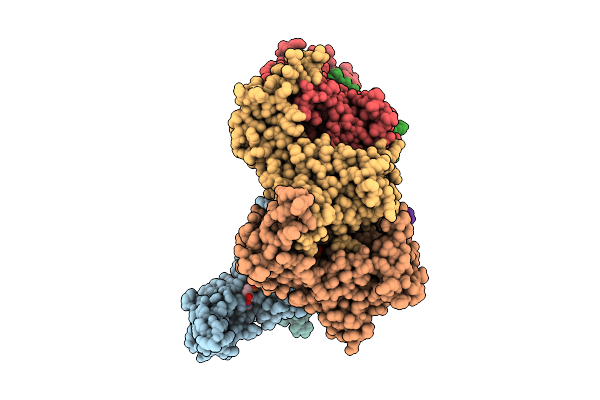 |
Hexamer Msp1 From S.Cerevisiae (With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
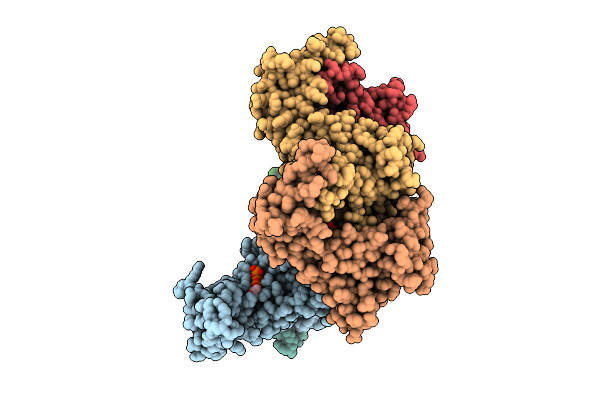 |
Hexamer Msp1 From S.Cerevisiae (With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
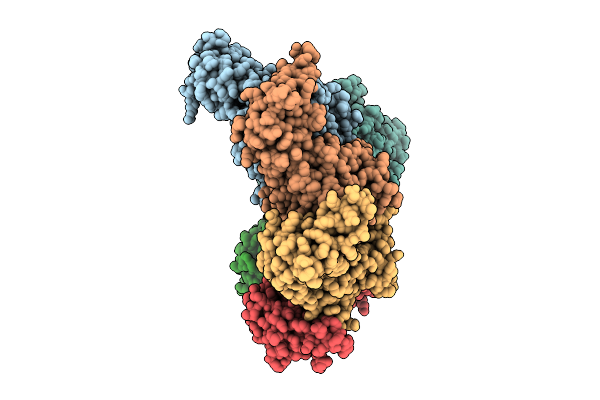 |
Hexamer Msp1 From S.Cerevisiae (With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
 |
Octamer Msp1 From S.Cerevisiae (With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
 |
Nonamer Msp1 From S.Cerevisiae (With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
 |
Decamer Msp1 From S.Cerevisiae(With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
 |
Pentamer Msp1 From S.Cerevisiae (With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate State2
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
 |
Sixteen Polymer Msp1 From S.Cerevisiae (With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
 |
Twenty-Two Polymer Msp1 From S.Cerevisiae(With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: MEMBRANE PROTEIN Ligands: MG, ATP |
 |
Hexamer Msp1 From S.Cerevisiae(With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: MEMBRANE PROTEIN Ligands: MG, ATP |
 |
Hexamer Msp1 From S.Cerevisiae (With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Resolution:2.87 Å Release Date: 2026-01-14 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
 |
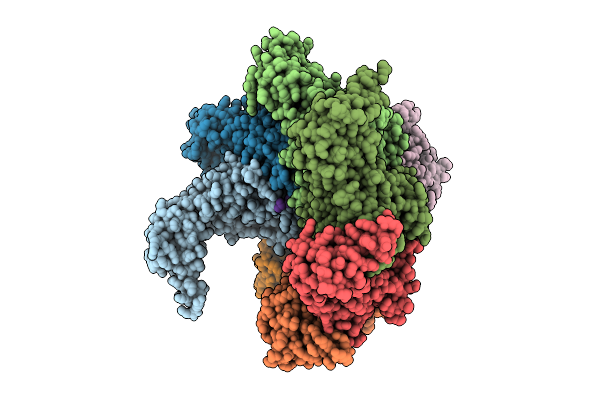
Hexamer Msp1 From S.Cerevisiae(With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c), Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2025-12-31 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
 |
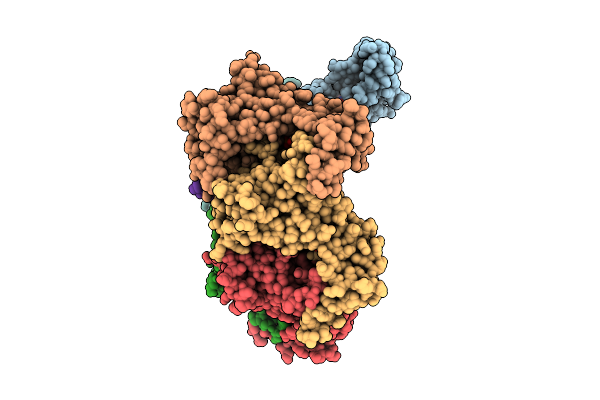
Octamer Msp1 From S.Cerevisiae(With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Resolution:3.17 Å Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: MG, ATP |
 |
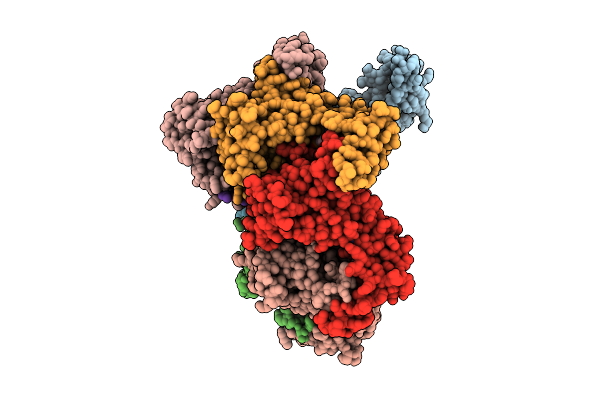
Hexamer Msp1 From S.Cerevisiae (With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Escherichia coli bl21(de3), Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
 |
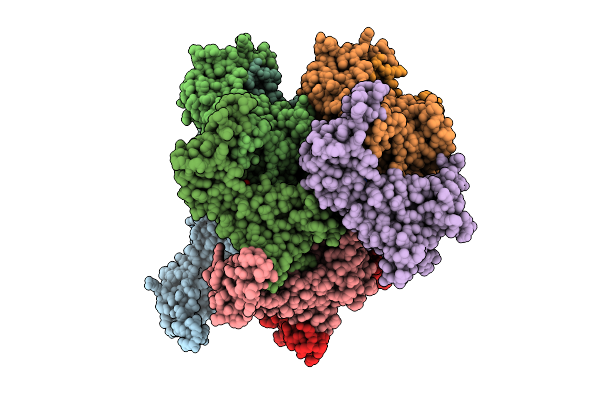
Heptamer Msp1 From S.Cerevisiae(With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Resolution:2.96 Å Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
 |
Eleven Polymer Msp1 From S.Cerevisiae (With A Catalytic Dead Mutaion) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Resolution:2.98 Å Release Date: 2025-12-10 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
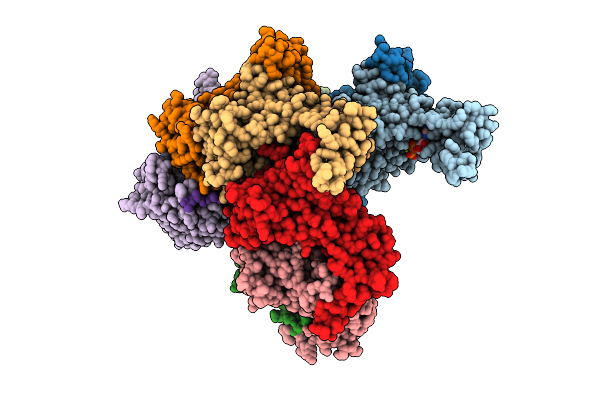 |
Nanomer Msp1 From S.Cerevisiae (With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2025-11-26 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |

