Search Count: 49,196
All
Selected
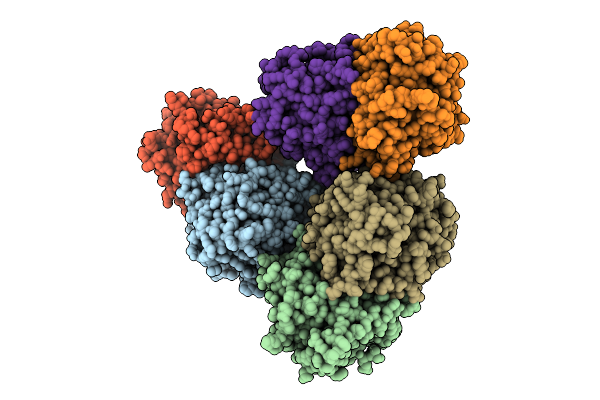 |
Sesterterpene Synthase From Streptomyces Mobaraensis (Sestermobaraene Synthase, Smts1) In Complex With Pyrophosphate (Ppi)
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-08 Classification: LYASE Ligands: MG, POP, PO4 |
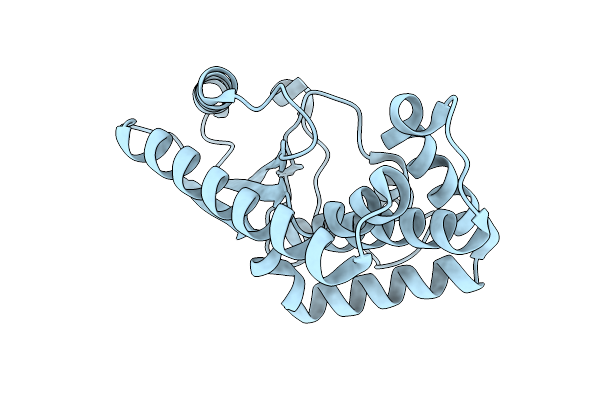 |
Structure Of A Putative Cyclodipeptide Synthase From Porphyromonas Gingivalis
Organism: Porphyromonas gingivalis
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2026-04-08 Classification: BIOSYNTHETIC PROTEIN |
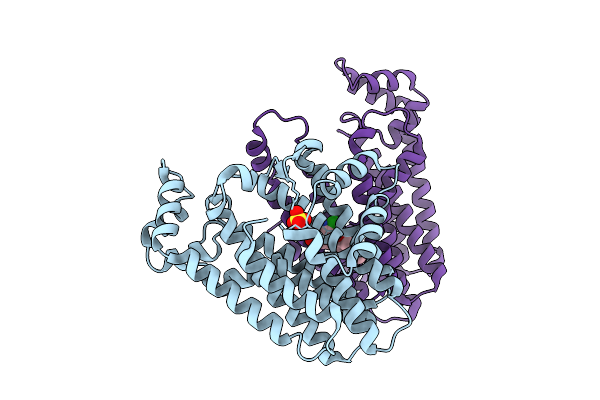 |
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: SO4, A1EC1 |
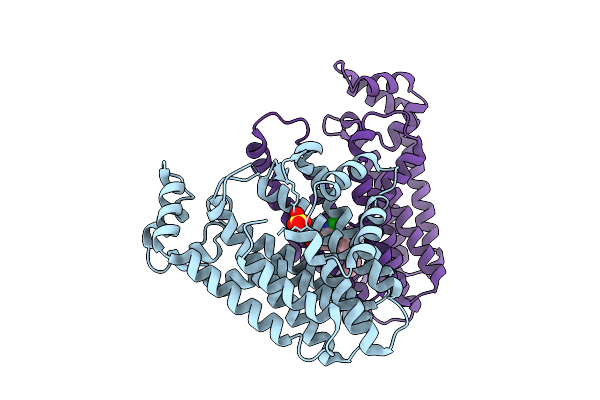 |
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: SO4, A1EC2 |
 |
Crystal Structure Of The Post-Reactive State Of Porcine Oas1 In Complex With Dsrna And Products 25A2 And Ppi Bound To The Catalytic Center.
Organism: Sus scrofa, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-04-08 Classification: TRANSFERASE/RNA Ligands: MG, MN, 25L, POP, EDO |
 |
Crystal Structure Of The Post-Reactive State Of Porcine Oas1 In Complex With Dsrna, Catalytic Center Bound Ppi, And Dissociated 25A2.
Organism: Sus scrofa, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.74 Å Release Date: 2026-04-08 Classification: TRANSFERASE/RNA Ligands: MG, POP, 25L |
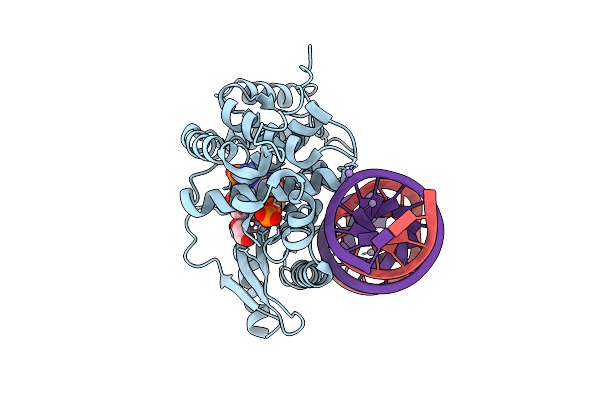 |
Crystal Structure Of The Pre-Reactive State Of Porcine Oas1 In Complex With Dsrna, Two Apcpp Substrate Analogs, Three Catalytic Mn2+ Ions.
Organism: Sus scrofa, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-04-08 Classification: TRANSFERASE/RNA Ligands: APC, MN, EDO |
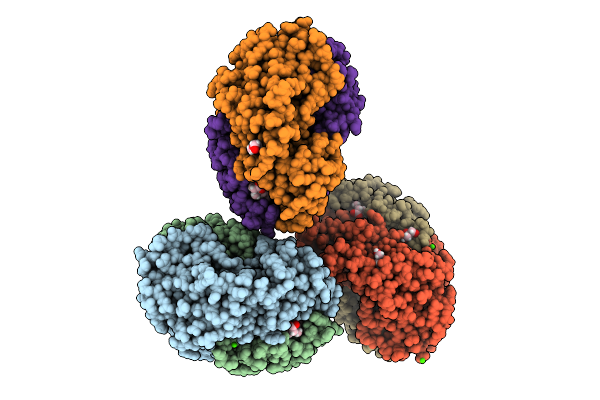 |
Organism: Streptomyces sp. b93
Method: X-RAY DIFFRACTION Resolution:2.14 Å Release Date: 2026-04-08 Classification: LYASE Ligands: HEM, GOL, CA, PEG |
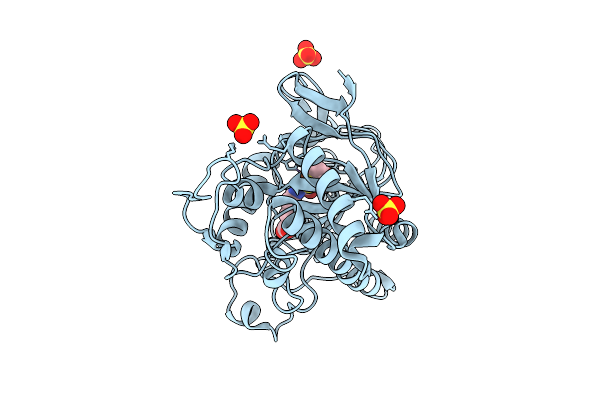 |
Organism: Aspergillus nidulans fgsc a4
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: FE, SO4, IP1 |
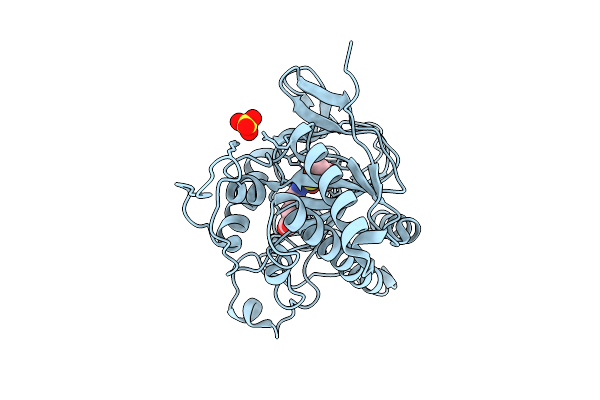 |
Organism: Aspergillus nidulans fgsc a4
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: FE, SO4, IP1 |
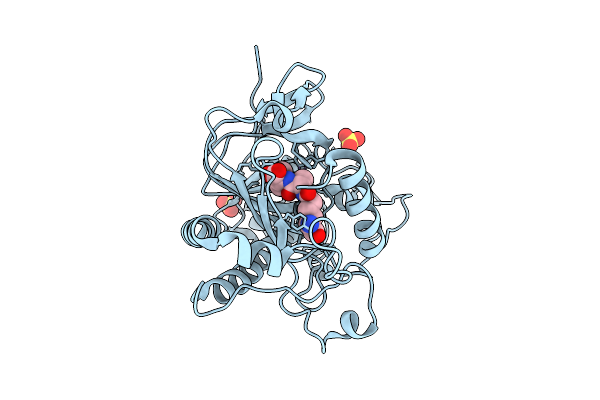 |
Organism: Aspergillus nidulans fgsc a4
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: SO4, FE, ACV |
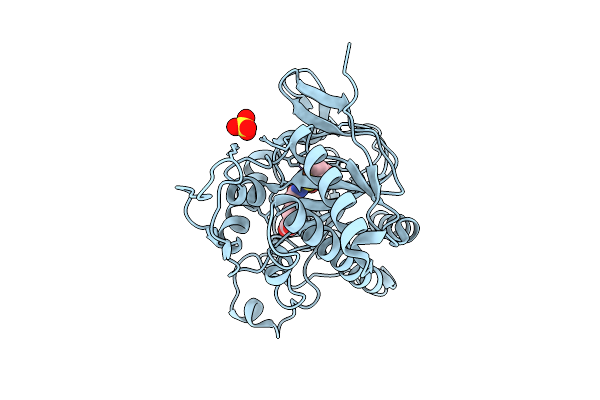 |
Organism: Aspergillus nidulans fgsc a4
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: FE, SO4, IP1 |
 |
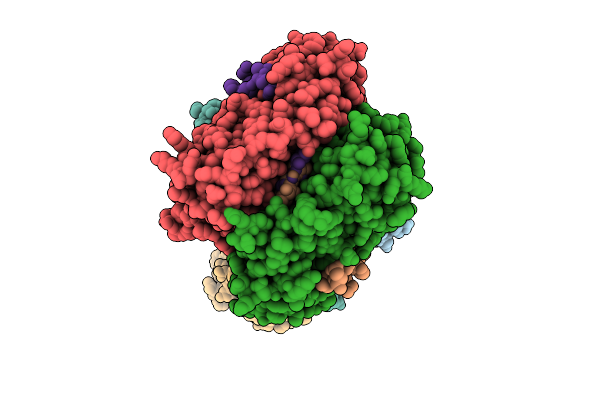
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
 |
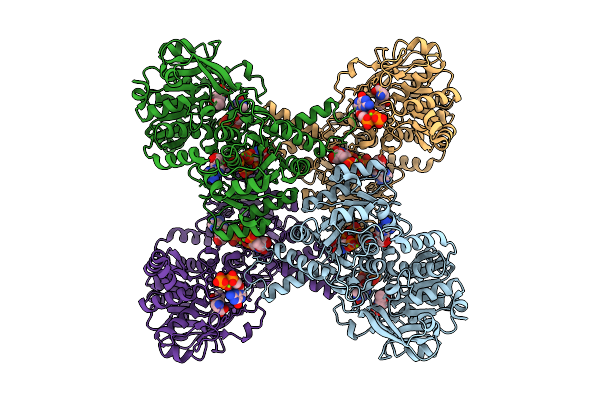
Cryoem Structure Of Human Mata2 In Complex With Mat2B Isoform V1 At 2.6 A Resolution
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
 |
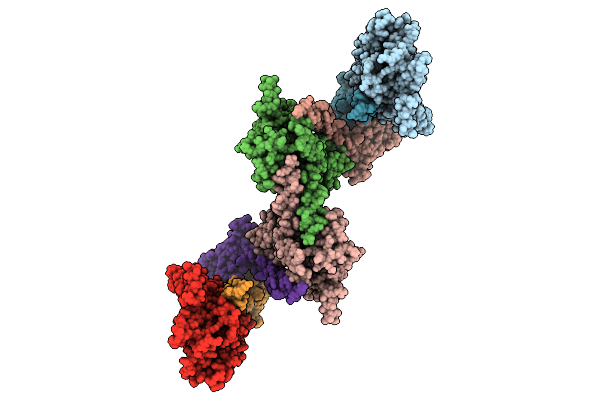
Organism: Drosophila melanogaster
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: LIGASE Ligands: ADP, 5ZL, GTP, ONL, MG |
 |
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: TRANSFERASE |
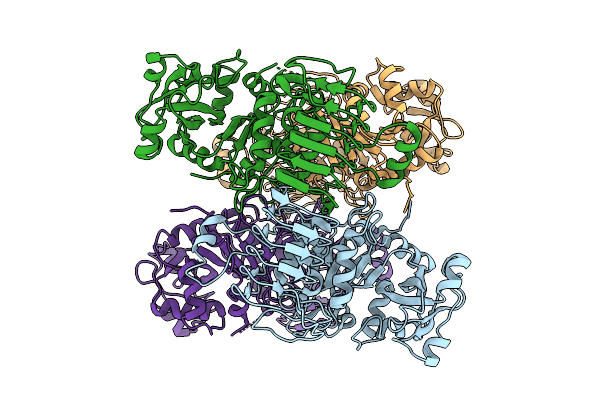 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:2.95 Å Release Date: 2026-04-08 Classification: PLANT PROTEIN |
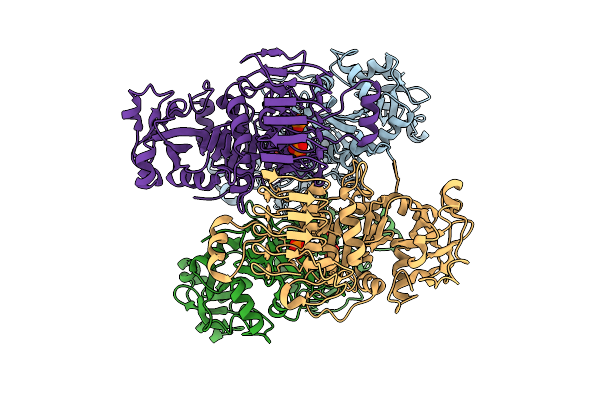 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:2.81 Å Release Date: 2026-04-08 Classification: PLANT PROTEIN Ligands: PO4 |
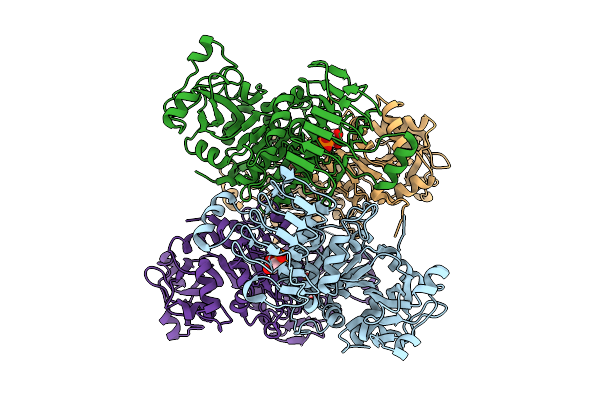 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:2.81 Å Release Date: 2026-04-08 Classification: PLANT PROTEIN Ligands: 3PG |
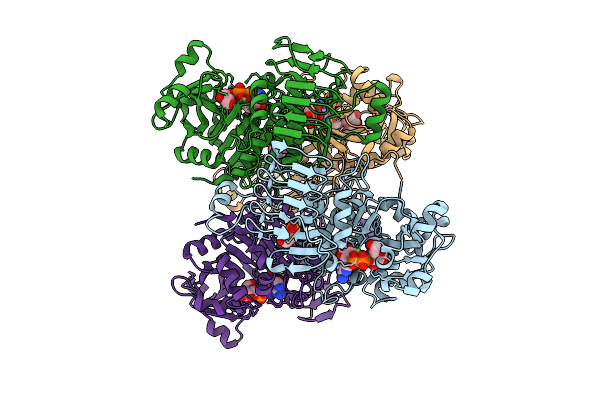 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:2.49 Å Release Date: 2026-04-08 Classification: PLANT PROTEIN Ligands: 3PG, ADQ, MG |

