Search Count: 24
All
Selected
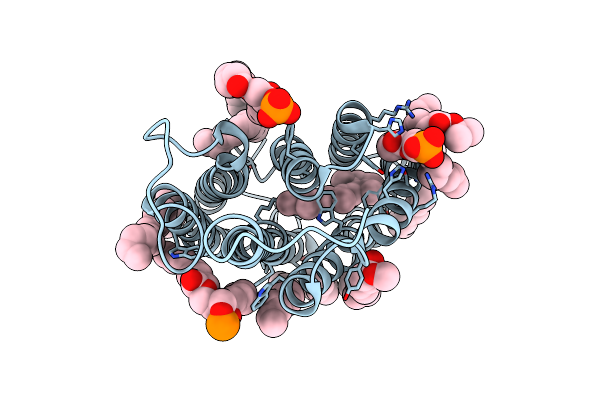 |
Cryo-Em Structure Of The Potassium-Selective Channelrhodopsin Hckcr1 In Lipid Nanodisc
Organism: Hyphochytrium catenoides
Method: ELECTRON MICROSCOPY Resolution:2.56 Å Release Date: 2023-09-06 Classification: MEMBRANE PROTEIN Ligands: RET, PSC, PLM |
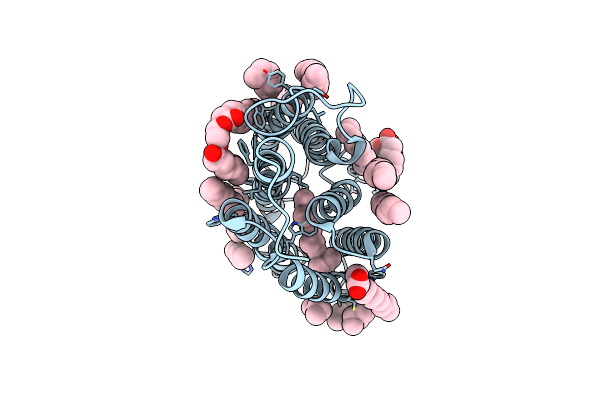 |
Cryo-Em Structure Of The Potassium-Selective Channelrhodopsin Hckcr2 In Lipid Nanodisc
Organism: Hyphochytrium catenoides
Method: ELECTRON MICROSCOPY Resolution:2.53 Å Release Date: 2023-09-06 Classification: MEMBRANE PROTEIN Ligands: RET, PSC, PLM |
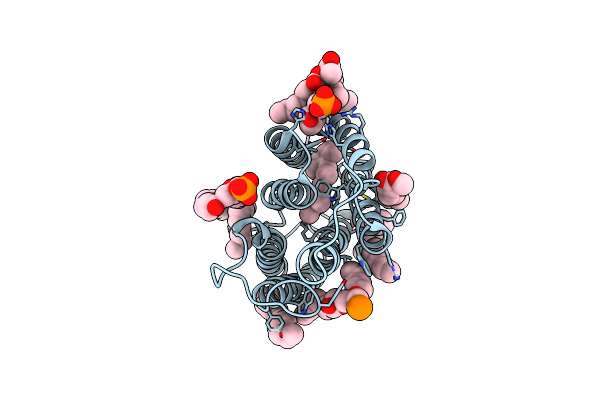 |
Cryo-Em Structure Of The Potassium-Selective Channelrhodopsin Hckcr1 H225F Mutant In Lipid Nanodisc
Organism: Hyphochytrium catenoides
Method: ELECTRON MICROSCOPY Resolution:2.66 Å Release Date: 2023-09-06 Classification: MEMBRANE PROTEIN Ligands: RET, PSC, PLM |
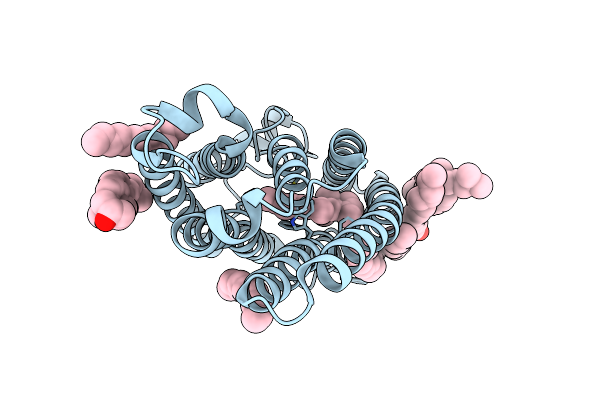 |
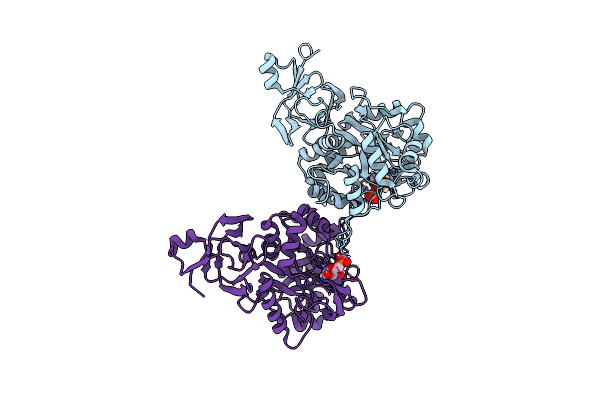
Organism: Rhodomonas lens
Method: ELECTRON MICROSCOPY Resolution:2.02 Å Release Date: 2022-02-02 Classification: MEMBRANE PROTEIN Ligands: RET, CLR, PLM |
 |
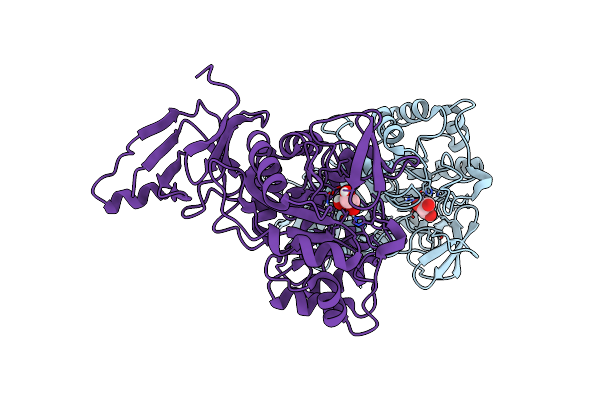
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2021-04-28 Classification: HYDROLASE Ligands: GLP, ZN |
 |
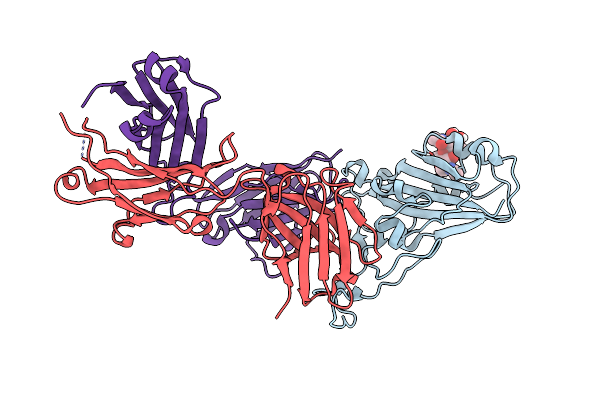
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2021-04-28 Classification: HYDROLASE Ligands: ZN, GOL |
 |
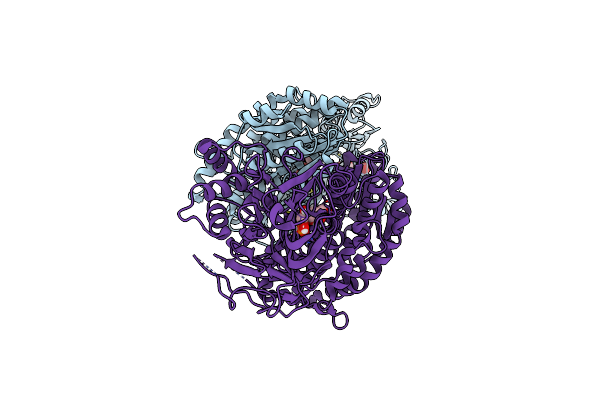
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2020-12-16 Classification: ANTIVIRAL PROTEIN |
 |
Crystal Structure Of Human Gfat-1 In Complex With Glucose-6-Phosphate And L-Glu
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2020-01-15 Classification: TRANSFERASE Ligands: G6Q, GLU |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2020-01-15 Classification: TRANSFERASE Ligands: G6Q |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2020-01-15 Classification: TRANSFERASE Ligands: UD1, MG, G6Q |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.24 Å Release Date: 2020-01-15 Classification: TRANSFERASE Ligands: G6Q, GLU |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.59 Å Release Date: 2020-01-15 Classification: TRANSFERASE Ligands: G6Q, GLU |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.42 Å Release Date: 2020-01-15 Classification: TRANSFERASE Ligands: UD1, MG, G6Q, GLU |
 |
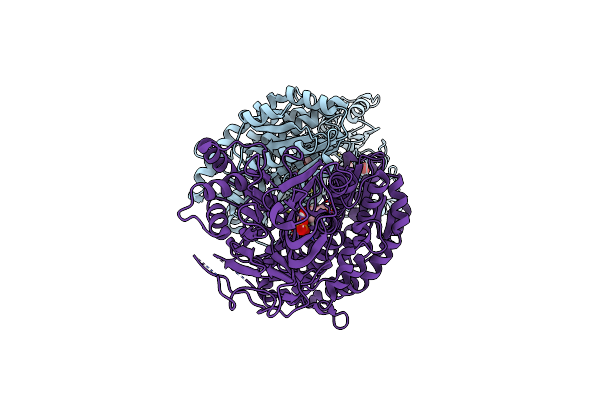
Crystal Structure Of Human Gfat-1 In Complex With Glucose-6-Phosphate, L-Glu, And Udp-Galnac
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.48 Å Release Date: 2020-01-15 Classification: TRANSFERASE Ligands: G6Q, GLU, UD2, MG |
 |
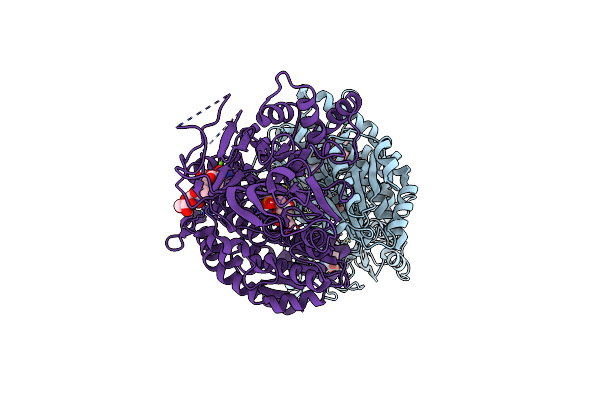
Crystal Structure Of Human Gfat-1 In Complex With Glucosamine-6-Phosphate And L-Glu
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.33 Å Release Date: 2020-01-15 Classification: TRANSFERASE Ligands: AGP, GLU |
 |
Crystal Structure Of Human Gfat-1 In Complex With Glucose-6-Phosphate, L-Glu, And Udp-Glcnac
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.53 Å Release Date: 2020-01-15 Classification: TRANSFERASE Ligands: UD1, MG, G6Q, GLU |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.72 Å Release Date: 2020-01-15 Classification: TRANSFERASE Ligands: G6Q, GLU |
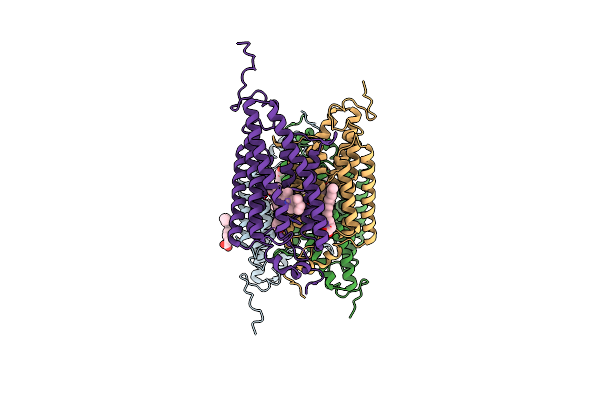 |
Organism: Guillardia theta ccmp2712
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2018-09-05 Classification: MEMBRANE PROTEIN Ligands: RET, OLA |
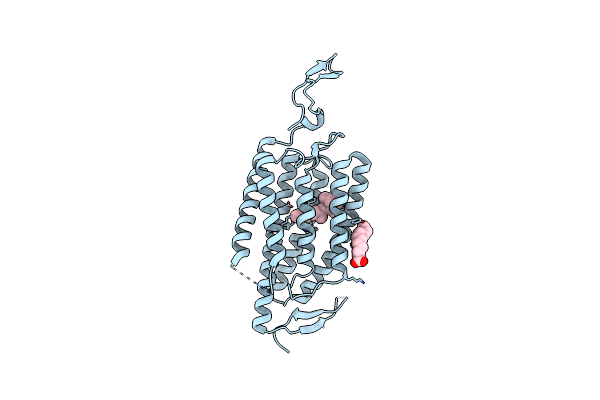 |
Organism: Chlamydomonas reinhardtii
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2018-09-05 Classification: MEMBRANE PROTEIN Ligands: RET, OLA, CL |
 |
Organism: Chlamydomonas reinhardtii
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2018-09-05 Classification: MEMBRANE PROTEIN Ligands: RET, OLA |

