Search Count: 329
All
Selected
 |
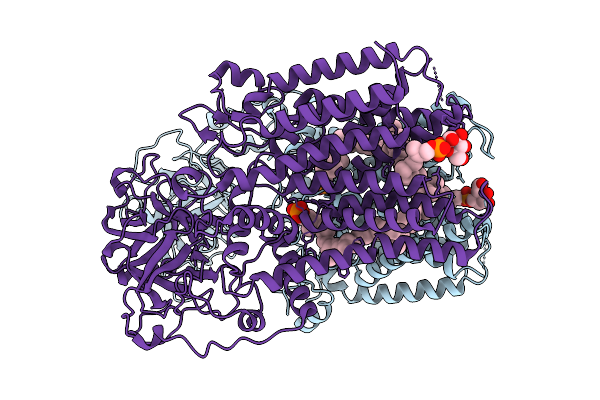
Organism: Thermochaetoides thermophila
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: A1I0N, A1I0M |
 |
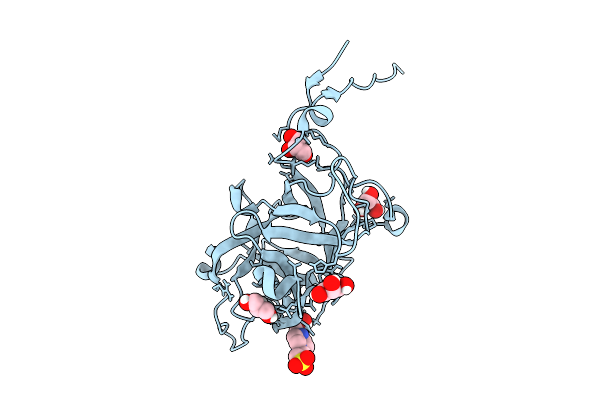
Organism: Thermochaetoides thermophila
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: A1I0N, A1I0M |
 |
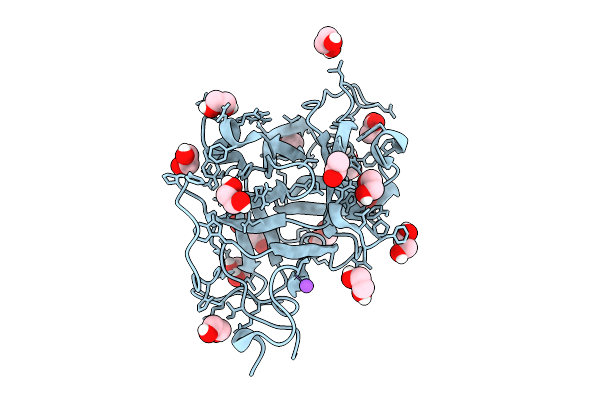
Structure Of The Saccharomyces Cerevisiae Pmt4-Mir Domain With Bound Ligands
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2025-11-26 Classification: PEPTIDE BINDING PROTEIN Ligands: EPE, GOL |
 |
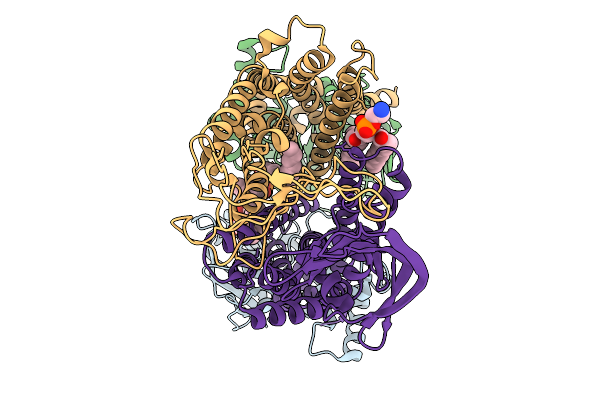
Structure Of The Chaetomium Thermophilum Pmt4-Mir Domain With Bound Ligands
Organism: Chaetomium
Method: X-RAY DIFFRACTION Resolution:1.22 Å Release Date: 2025-11-26 Classification: PEPTIDE BINDING PROTEIN Ligands: EDO, ACT, NA |
 |
Organism: Pseudomonas aeruginosa
Method: ELECTRON MICROSCOPY Release Date: 2025-11-19 Classification: LIPID TRANSPORT Ligands: 3PE |
 |
Organism: Pseudomonas aeruginosa
Method: ELECTRON MICROSCOPY Release Date: 2025-11-19 Classification: LIPID TRANSPORT Ligands: 3PE |
 |
Organism: Pseudomonas aeruginosa
Method: ELECTRON MICROSCOPY Release Date: 2025-11-19 Classification: LIPID TRANSPORT Ligands: 3PE |
 |
Organism: Pseudomonas aeruginosa
Method: ELECTRON MICROSCOPY Release Date: 2025-11-19 Classification: LIPID TRANSPORT Ligands: 3PE |
 |
Organism: Pseudomonas aeruginosa
Method: ELECTRON MICROSCOPY Release Date: 2025-11-19 Classification: LIPID TRANSPORT |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-04-30 Classification: MEMBRANE PROTEIN Ligands: ADP, MG |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-04-30 Classification: MEMBRANE PROTEIN Ligands: ADP, MG |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-04-23 Classification: MEMBRANE PROTEIN Ligands: ADP, MG |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-04-23 Classification: MEMBRANE PROTEIN Ligands: ADP, MG |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-04-23 Classification: MEMBRANE PROTEIN Ligands: ADP, MG |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-04-23 Classification: MEMBRANE PROTEIN Ligands: ADP, MG |
 |
Crystal Structure Of Orphan G Protein-Coupled Receptor 6 With Bound Inverse Agonist 3H
Organism: Homo sapiens, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2024-12-04 Classification: MEMBRANE PROTEIN Ligands: XY8, NA |
 |
Organism: Homo sapiens, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:3.49 Å Release Date: 2024-12-04 Classification: MEMBRANE PROTEIN Ligands: X7T |
 |
Organism: Homo sapiens, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2024-12-04 Classification: MEMBRANE PROTEIN Ligands: OLC, P15, OLA, CLR |
 |
Organism: Homo sapiens, Lama glama
Method: ELECTRON MICROSCOPY Release Date: 2024-12-04 Classification: MEMBRANE PROTEIN |
 |
Cvhsp (Hspb7) C131S Alpha-Crystallin Domain - Filamin C (Flnc) Domain 24 Complex
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2024-10-16 Classification: CHAPERONE |

