Search Count: 288
All
Selected
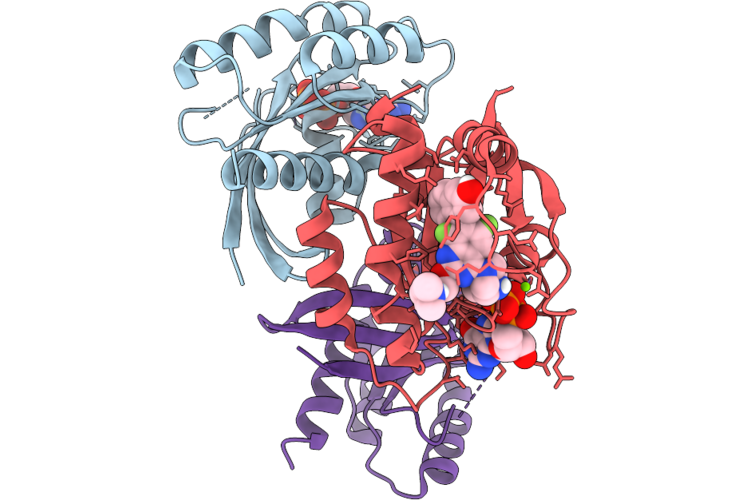 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-04-22 Classification: SIGNALING PROTEIN Ligands: MG, GDP, A1JU5 |
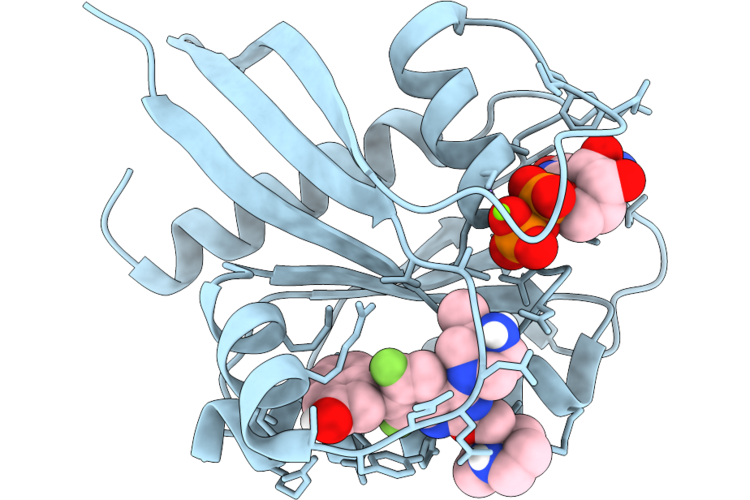 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-04-22 Classification: SIGNALING PROTEIN Ligands: GDP, MG, A1JU6 |
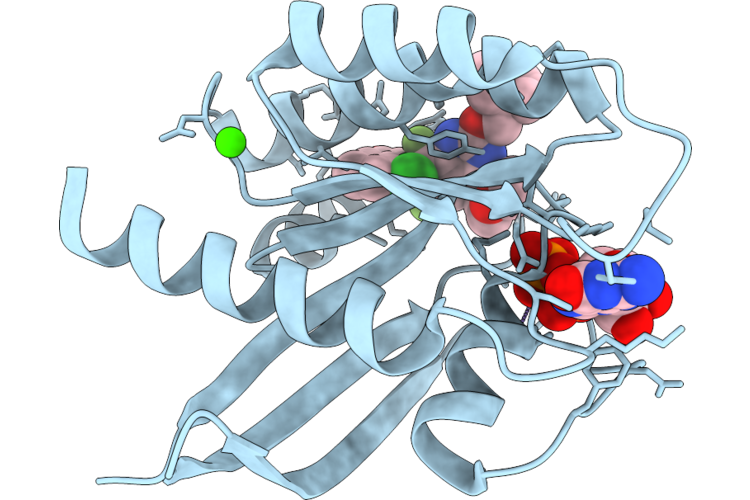 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.54 Å Release Date: 2026-04-22 Classification: SIGNALING PROTEIN Ligands: MG, GDP, CA, A1JU7 |
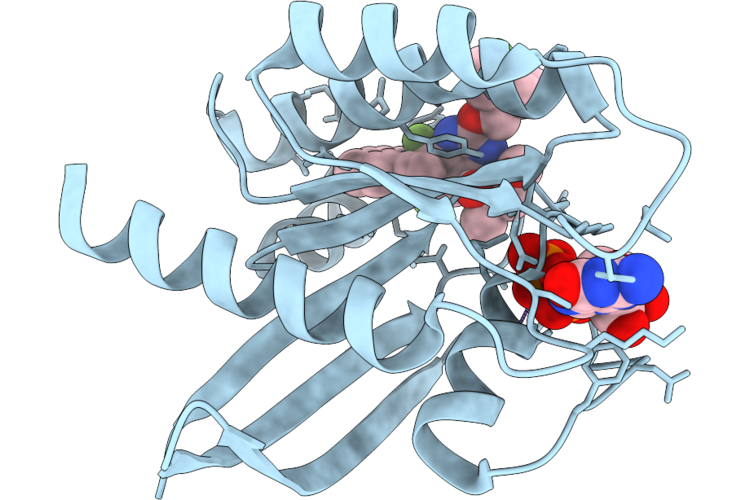 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-04-22 Classification: SIGNALING PROTEIN Ligands: MG, GDP, A1JU8 |
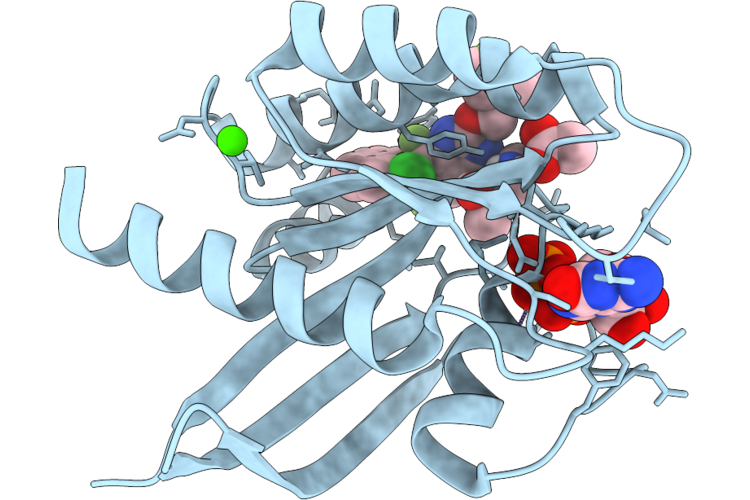 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2026-04-22 Classification: SIGNALING PROTEIN Ligands: GDP, MG, CA, ACT, A1JU3 |
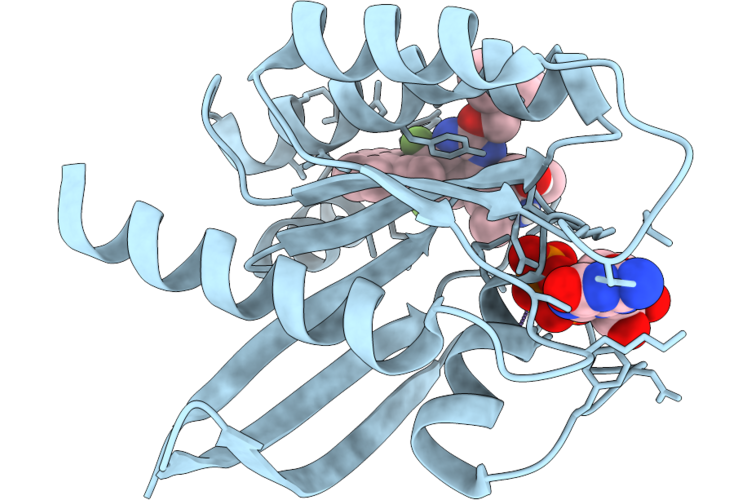 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2026-04-22 Classification: SIGNALING PROTEIN Ligands: MG, GDP, A1JU9 |
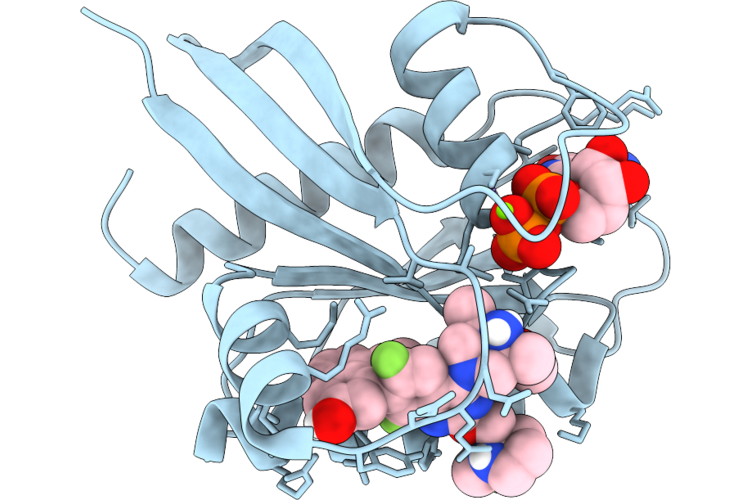 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2026-04-22 Classification: SIGNALING PROTEIN Ligands: GDP, MG, A1JU2 |
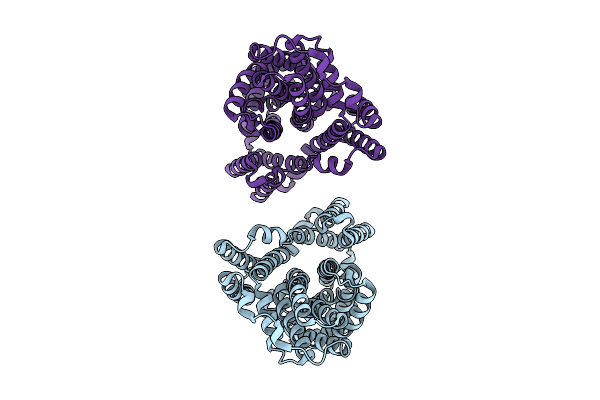 |
Organism: Leptospirillum ferriphilum
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: MEMBRANE PROTEIN |
 |
Organism: Leptospirillum ferriphilum
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: MEMBRANE PROTEIN |
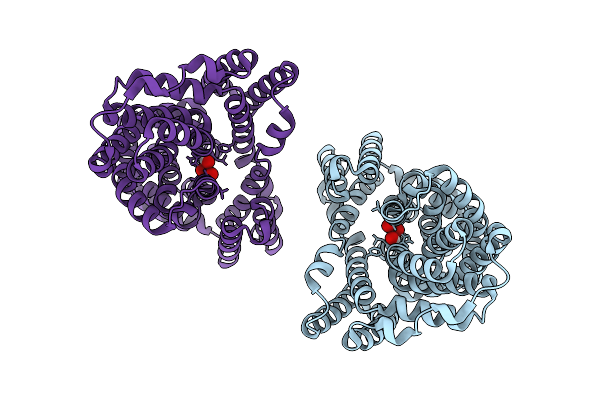 |
Arsb From L. Ferriphilum Bound To Arsenite In Inward-Facing State (Parallel Dimer)
Organism: Leptospirillum ferriphilum
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-03-11 Classification: MEMBRANE PROTEIN Ligands: TAS |
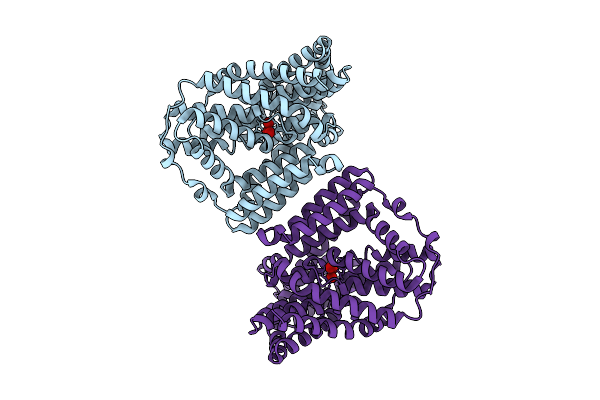 |
Arsb From L. Ferriphilum Bound To Antimonite In Inward-Facing State (Antiparallel Dimer)
Organism: Leptospirillum ferriphilum
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-03-11 Classification: MEMBRANE PROTEIN Ligands: SBO |
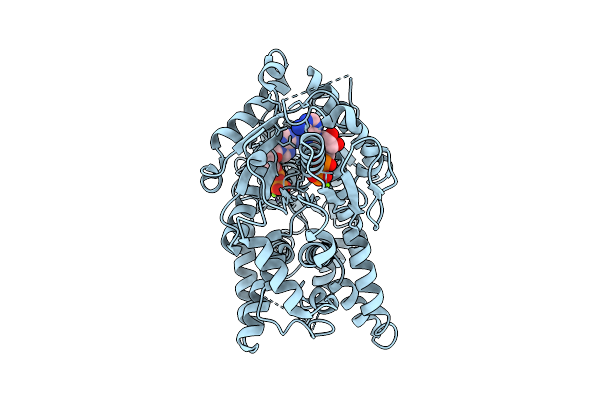 |
Open Conformation Of Arsa From L. Ferriphilum In Complex With Mgadp Determined In The Presence Of Arsenite
Organism: Leptospirillum ferriphilum
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: HYDROLASE Ligands: ADP, MG |
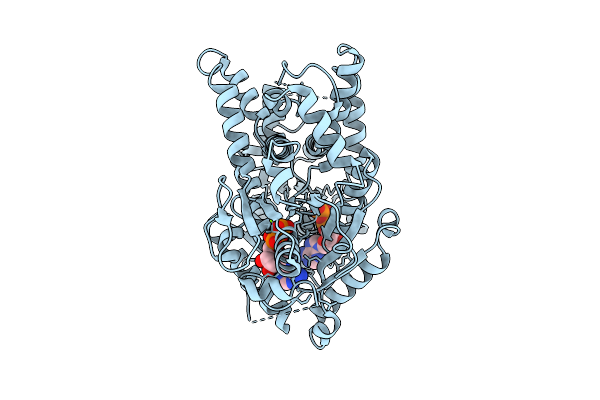 |
Open Conformation Of Arsa From L. Ferriphilum In Complex With Mgadp Determined In Absence Of Arsenite
Organism: Leptospirillum ferriphilum
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: HYDROLASE Ligands: ADP, MG |
 |
Closed Conformation Of Arsa From L. Ferriphilum In Complex With Mgatp And Arsenite
Organism: Leptospirillum ferriphilum
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: HYDROLASE Ligands: ATP, MG, ARS |
 |
Closed Conformation Of Arsa From L. Ferriphilum In Complex With Mgatp And Arsenite At 1.5 Minute Time Point
Organism: Leptospirillum ferriphilum ml-04
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: HYDROLASE Ligands: ATP, MG, ARS |
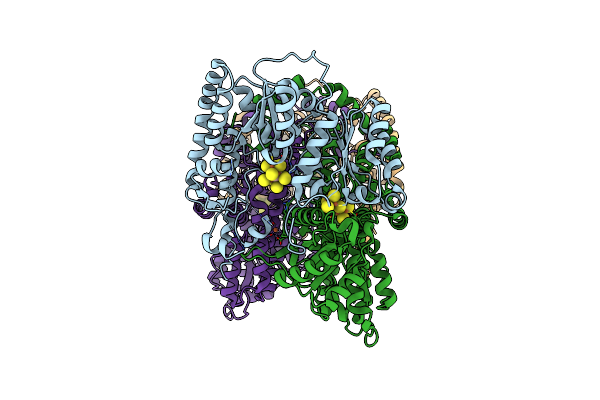 |
Cryoem Structure Of Nitrogenase Mofe-Protein 60 Minute Time Point Under Alkaline Turnover
Organism: Azotobacter vinelandii
Method: ELECTRON MICROSCOPY Release Date: 2024-12-18 Classification: METAL BINDING PROTEIN Ligands: ICS, CLF, FE |
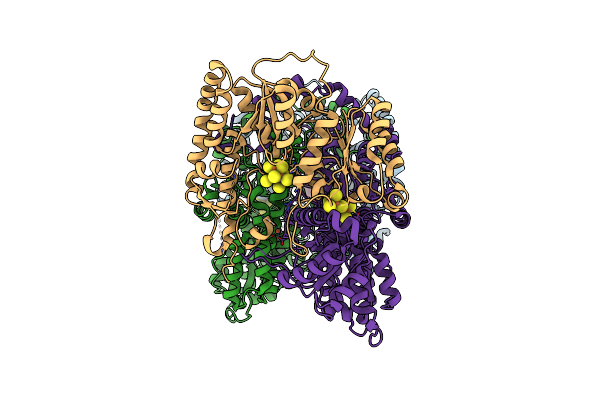 |
Cryoem Structure Of Nitrogenase Mofe-Protein 20 Minute Time Point Under Alkaline Turnover
Organism: Azotobacter vinelandii
Method: ELECTRON MICROSCOPY Release Date: 2024-12-18 Classification: METAL BINDING PROTEIN Ligands: ICS, HCA, CLF, FE |
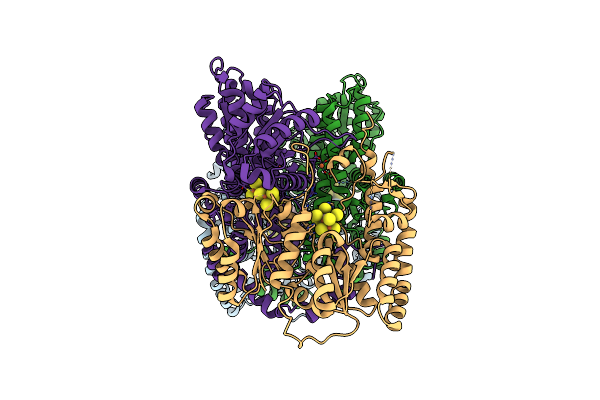 |
Cryoem Structure Of Nitrogenase Mofe-Protein 5 Minute Time Point Under Alkaline Turnover
Organism: Azotobacter vinelandii
Method: ELECTRON MICROSCOPY Release Date: 2024-12-18 Classification: METAL BINDING PROTEIN Ligands: ICS, HCA, CLF, FE |
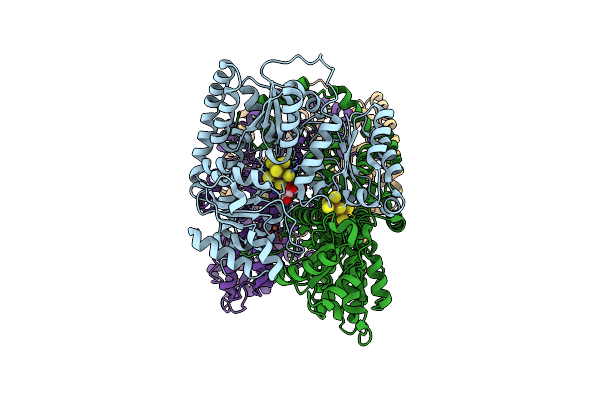 |
Cryoem Structure Of Nitrogenase Mofe-Protein 20 Second Time Point Under Alkaline Turnover
Organism: Azotobacter vinelandii
Method: ELECTRON MICROSCOPY Release Date: 2024-12-18 Classification: METAL BINDING PROTEIN Ligands: ICS, HCA, CLF, FE |
 |
Cryoem Structure Of Alkaline-Inactivated Nitrogenase Mofe-Protein In Complex With Naft
Organism: Azotobacter vinelandii
Method: ELECTRON MICROSCOPY Release Date: 2024-12-18 Classification: METAL BINDING PROTEIN Ligands: ICS, CLF, FE, 1N7 |

