Search Count: 61
All
Selected
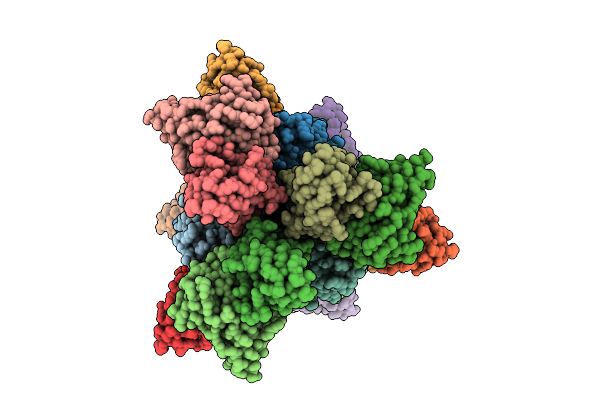 |
Cryo-Em Structure Of Sudan Ebolavirus Gp Bound By Three Neutralizing Antibodies 545S, 523S And 294S
Organism: Sudan ebolavirus, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
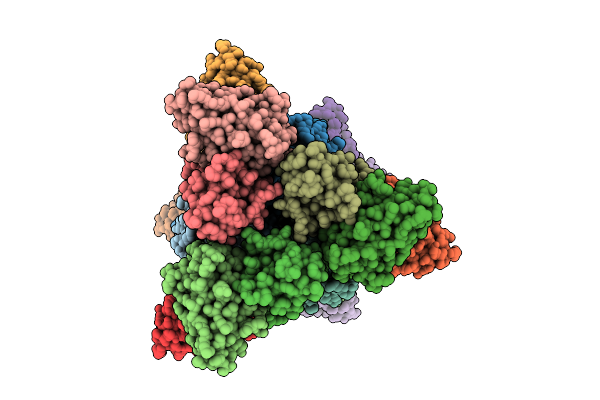 |
Cryo-Em Structure Of Sudan Ebolavirus Gp Bound By Three Neutralizing Antibodies 316L, 523S And 294S
Organism: Sudan ebolavirus, Macaca mulatta, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
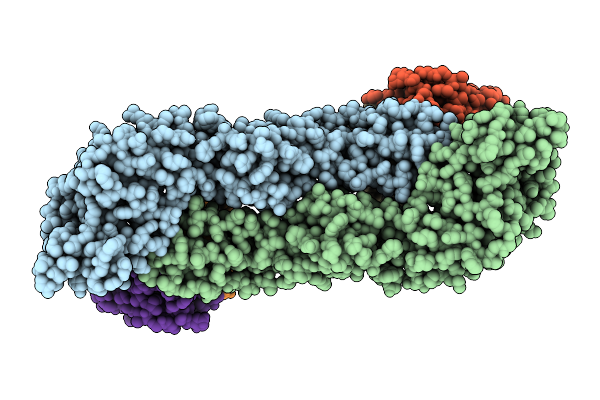 |
Cryo-Em Structure Of Modified Zika Virus E Protein Dimer Complexed With A Neutralizing Antibody Smzab2 Fab
Organism: Homo sapiens, Zika virus
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
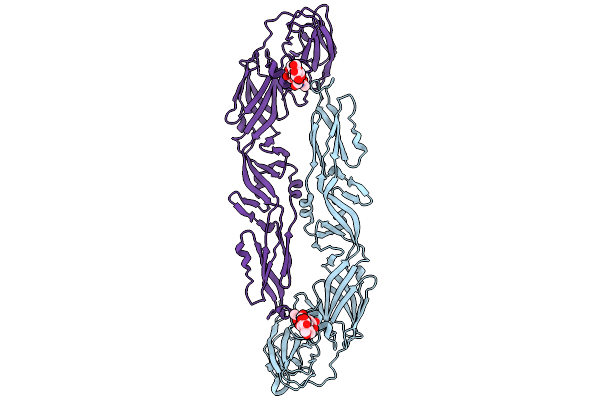 |
Organism: Japanese encephalitis virus
Method: ELECTRON MICROSCOPY Resolution:4.18 Å Release Date: 2026-01-21 Classification: VIRAL PROTEIN |
 |
Cryo-Em Structure Of Modified Zika Virus E Protein Dimer Complexed With A Neutralizing Antibody Oz-D4 Fab
Organism: Zika virus, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: VIRAL PROTEIN |
 |
Cryo-Em Structure Of J601-1B2 Fab In Complex With Hiv-1 Bg505 Ds-Sosip Env Trimer
Organism: Human immunodeficiency virus 1, Macaca mulatta
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG, MAN |
 |
Cryo-Em Structure Of J601-A6 Fab In Complex With Hiv-1 Bg505 Ds-Sosip Env Trimer
Organism: Human immunodeficiency virus 1, Macaca mulatta
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Cryo-Em Structure Of K001-A1 Fab In Complex With Hiv-1 459C-Opt Rns Ds-Sosip Env Trimer
Organism: Macaca mulatta, Human immunodeficiency virus 1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Organism: Human immunodeficiency virus 1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Organism: Human immunodeficiency virus 1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Cryo-Em Structure Of Neutralizing Murine Antibody Ws.Hsv-1.24 In Complex With Hsv-1 Glycoprotein B Trimer Gb-Ecto.516P.531E
Organism: Human alphaherpesvirus 1, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Cryo-Em Structure Of Neutralizing Human Antibody D48 In Complex With Hsv-1 Glycoprotein B Trimer Gb-Ecto.516P.531E.Ds
Organism: Human alphaherpesvirus 1, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Cryo-Em Structure Of Neutralizing Human Antibody D48 In Complex With Hsv-1 Glycoprotein B Trimer Gb-Ecto.516P
Organism: Human alphaherpesvirus 1, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: VIRAL PROTEIN Ligands: NAG |
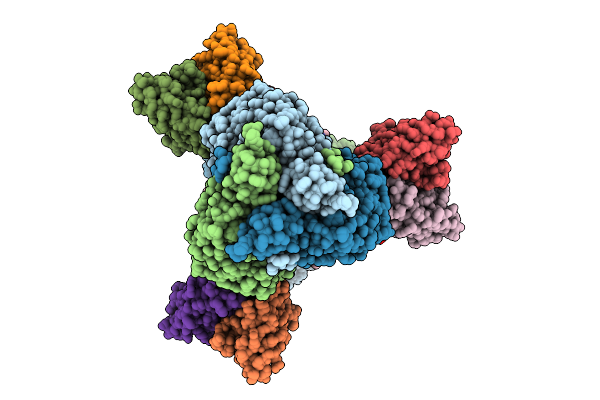 |
Cryo-Em Structure Of Neutralizing Human Antibody D48 In Complex With Hsv-1 Glycoprotein B Trimer Gb-Ecto.516P.531E
Organism: Human alphaherpesvirus 1, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: VIRAL PROTEIN Ligands: NAG |
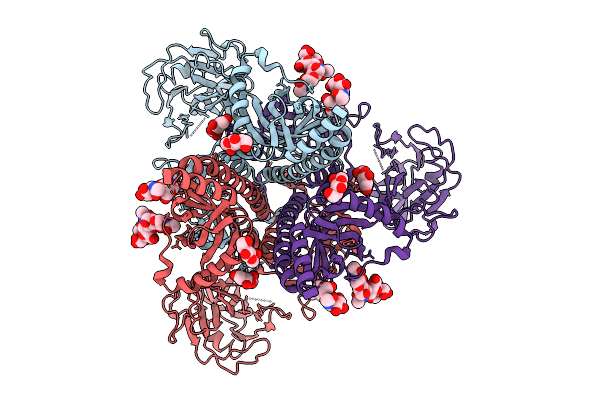 |
Cryo-Em Structure Of Gb-Ecto.516P.531E.Ds, A Prefusion-Stabilized Hsv-1 Glycoprotein B Extracellular Domain
Organism: Human alphaherpesvirus 1
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: VIRAL PROTEIN Ligands: NAG |
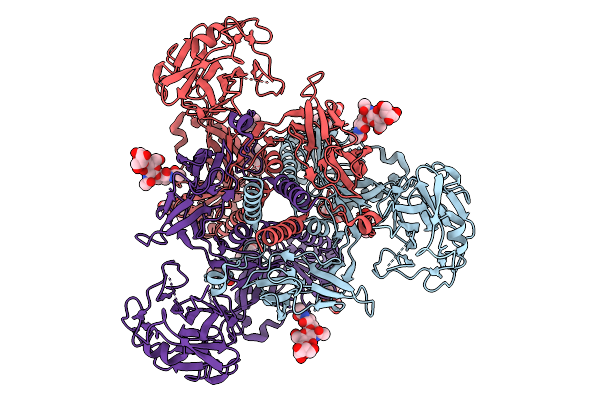 |
Cryo-Em Structure Of Gb-Ecto.516P.531E, A Prefusion-Stabilized Hsv-1 Glycoprotein B Extracellular Domain
Organism: Human alphaherpesvirus 1
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: VIRAL PROTEIN Ligands: NAG |
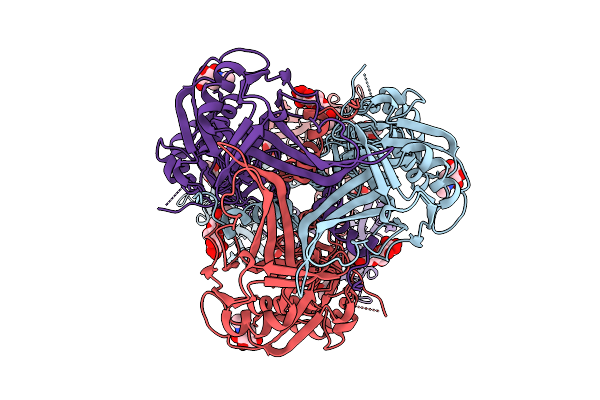 |
Cryo-Em Structure Of Gb-Ecto.516P, An Hsv-1 Glycoprotein B Extracellular Domain
Organism: Human alphaherpesvirus 1
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Cryoem Structure Of Nc99 Hemagglutinin Trimer In Complex With Fab Bb798E 3-C07
Organism: Influenza a virus, Macaca fascicularis
Method: ELECTRON MICROSCOPY Release Date: 2025-07-09 Classification: VIRAL PROTEIN |
 |
Cryoem Structure Of Nc99 Hemagglutinin Trimer In Complex With Fab T009 3-E04
Organism: Influenza a virus, Macaca fascicularis
Method: ELECTRON MICROSCOPY Release Date: 2025-07-09 Classification: VIRAL PROTEIN |
 |
Cryo-Em Structure Of Rhesus Antibody T646-A.01 In Complex With Hiv-1 Env Trimer Q23.17 Md39
Organism: Human immunodeficiency virus 1, Macaca mulatta
Method: ELECTRON MICROSCOPY Release Date: 2025-07-02 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |

