Search Count: 302
All
Selected
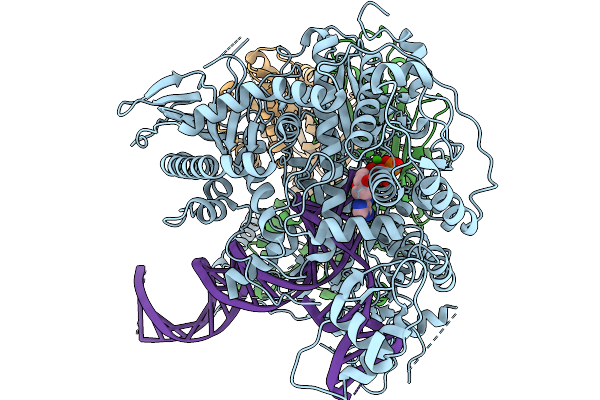 |
Strand Displacement State I Of Human Mitochondrial Dna Polymerase Gamma Ternary Complex
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.23 Å Release Date: 2026-02-18 Classification: TRANSFERASE/DNA Ligands: CA, DTP |
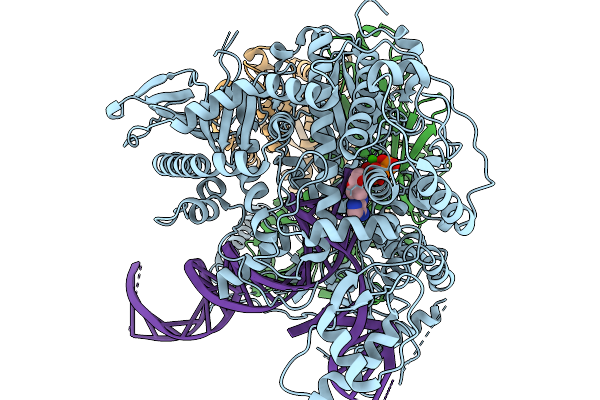 |
Strand Displacement State Ii Of Human Mitochondrial Dna Polymerase Gamma Ternary Complex
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: TRANSFERASE/DNA Ligands: CA, DTP |
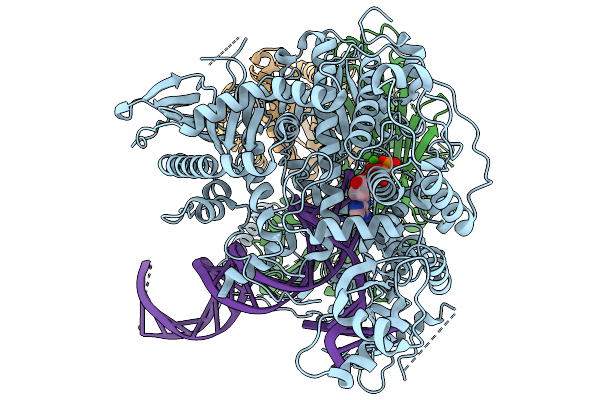 |
Strand Displacement State Iii Of Human Mitochondrial Dna Polymerase Gamma Ternary Complex
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: TRANSFERASE/DNA Ligands: CA, DTP |
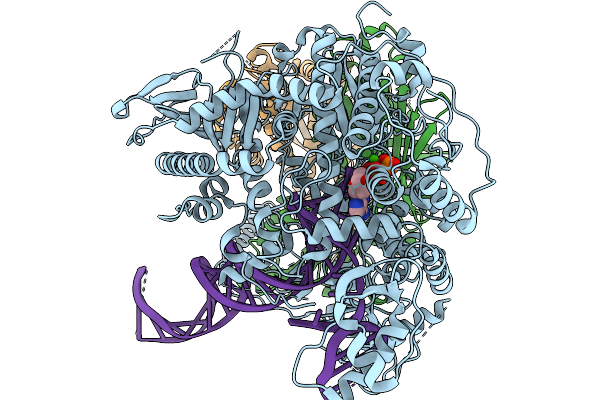 |
Strand Displacement State Iv Of Human Mitochondrial Dna Polymerase Gamma Ternary Complex
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: TRANSFERASE/DNA Ligands: CA, DTP |
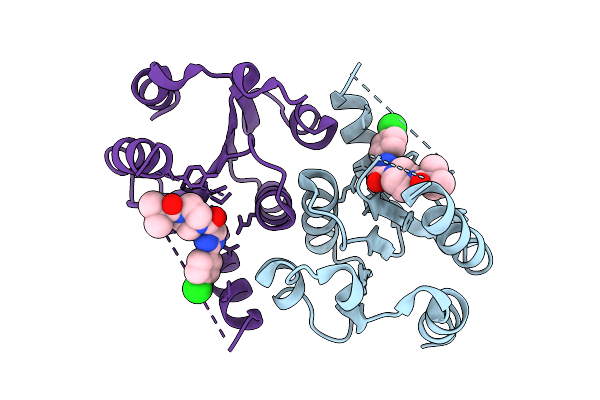 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2025-12-24 Classification: HYDROLASE Ligands: A1JH5 |
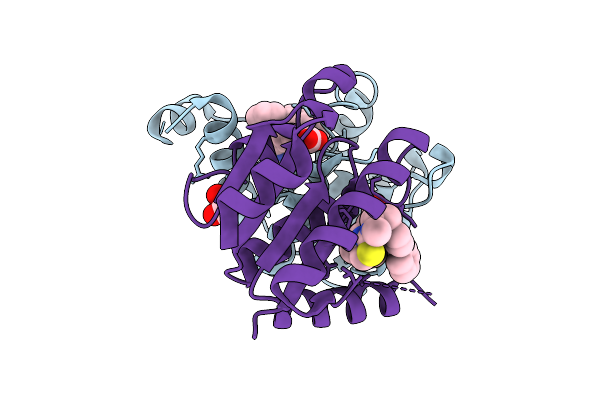 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.51 Å Release Date: 2025-12-24 Classification: HYDROLASE Ligands: A1JKT, EDO |
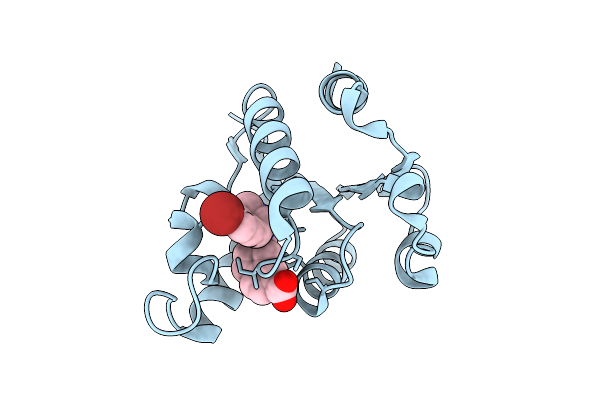 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-12-24 Classification: HYDROLASE Ligands: A1JKZ |
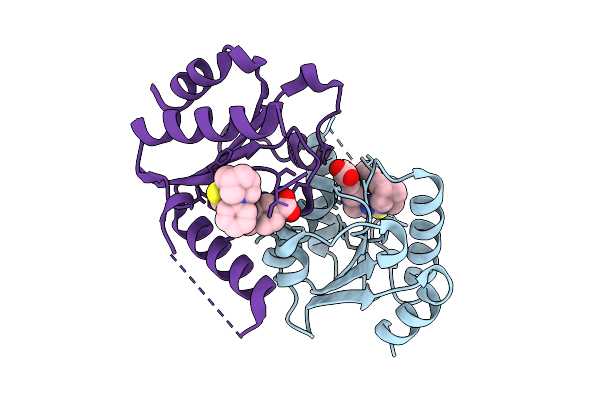 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2025-12-24 Classification: HYDROLASE Ligands: CL, A1JK6 |
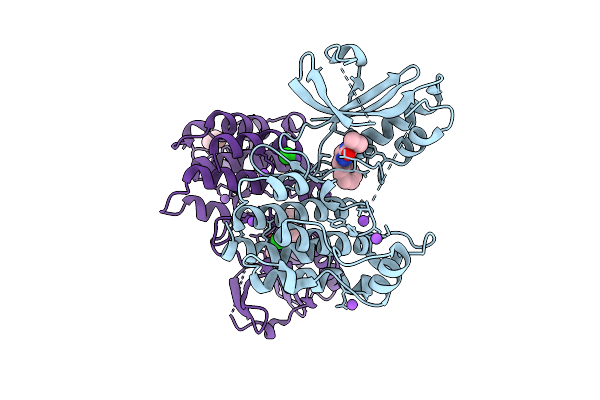 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2025-11-12 Classification: TRANSFERASE/Inhibitor Ligands: A1CNU, BR, CL, NA, TME |
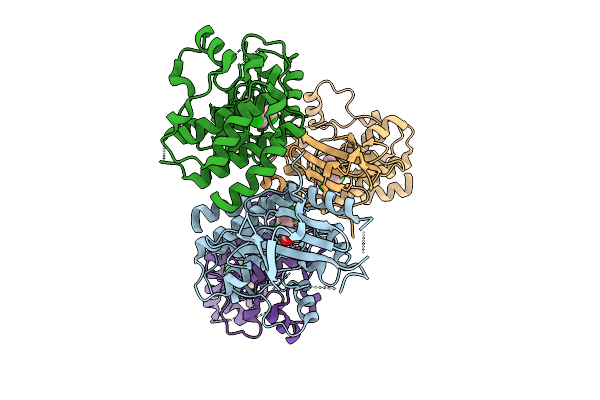 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.49 Å Release Date: 2025-11-12 Classification: Transferase/Inhibitor Ligands: A1CNV |
 |
Cryo-Em Structure Of The Human Mitochondrial Rna Polymerase Transcription Initiation Complex (Polrmt/Tfam/Tfb2M/Dna/Rna) With A 2-Mer Rna (Pppgpa) And Gtp Poised For Catalysis (Pre-Ic3)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-07-30 Classification: TRANSCRIPTION Ligands: GTP, MG |
 |
Cryo-Em Structure Of The Human Mitochondrial Rna Polymerase Transcription Initiation Complex (Polrmt/Tfb2M/Dna/Rna) Without Tfam; And With A 2-Mer Rna (Pppgpa) And Gtp Poised For Catalysis (Pre-Ic3-Tfam)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-07-30 Classification: TRANSCRIPTION Ligands: GTP, MG |
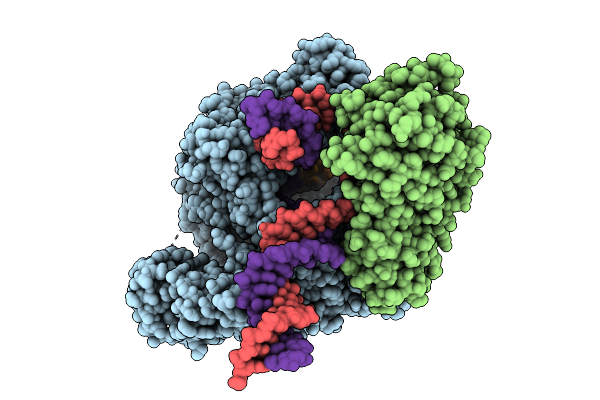 |
Cryo-Em Structure Of The Human Mitochondrial Rna Polymerase Transcription Initiation Complex (Polrmt/Tfb2M/Dna/Rna) Without Tfam; And With A Slipped 3-Mer Rna, Pppgpgpa (Slipped Ic3-Tfam)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-07-30 Classification: TRANSCRIPTION |
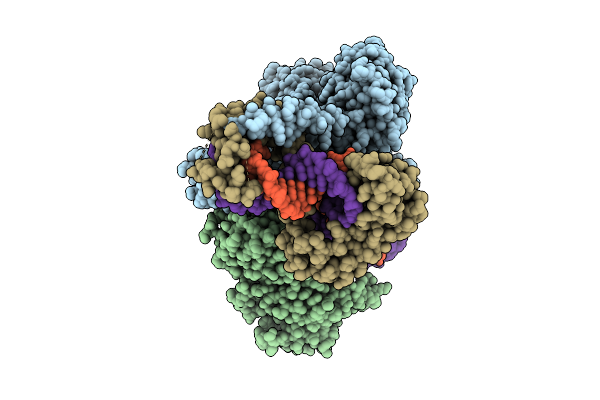 |
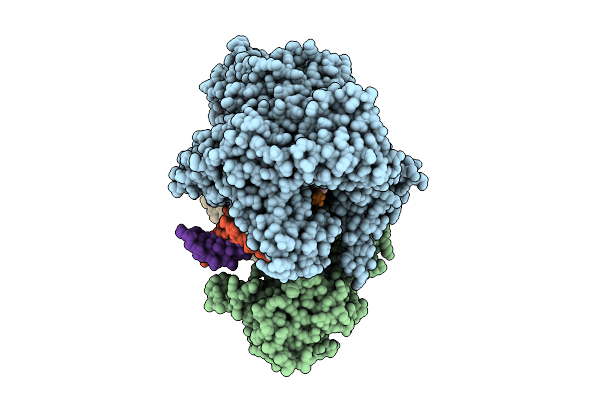
Cryo-Em Structure Of The Human Mitochondrial Rna Polymerase Transcription Initiation Complex (Polrmt/Tfam/Tfb2M/Dna/Rna) With A Slipped 3-Mer Rna, Pppgpgpa (Slipped Ic3)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-07-30 Classification: TRANSCRIPTION |
 |
Cryo-Em Structure Of The Human Mitochondrial Rna Polymerase Transcription Initiation Complex (Polrmt/Tfam/Tfb2M/Dna/Rna) With A Slipped 3-Mer Rna (Pppgpgpa) And Gtp Poised For Catalysis (Slipped Pre-Ic4)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-07-30 Classification: TRANSCRIPTION Ligands: GTP, MG |
 |
Organism: Oscillatoria sp. n9dm
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-04-16 Classification: PHOTOSYNTHESIS Ligands: CYC, GOL |
 |
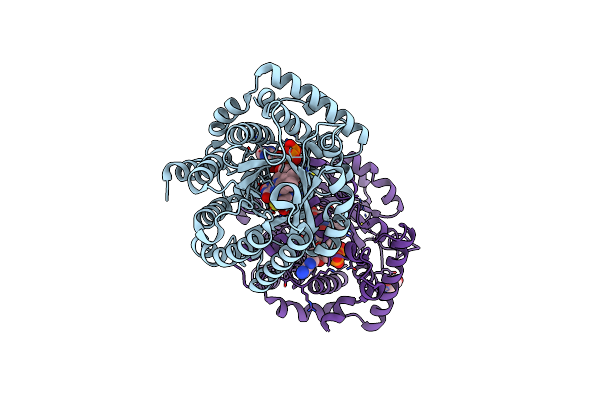
Minimal Puta Proline Dehydrogenase Domain (Design #2) Complexed With (Prop-2-Ynylthio)Acetic Acid
Organism: Sinorhizobium meliloti sm11
Method: X-RAY DIFFRACTION Resolution:1.38 Å Release Date: 2024-10-30 Classification: OXIDOREDUCTASE Ligands: FAD, PEG, X79 |
 |
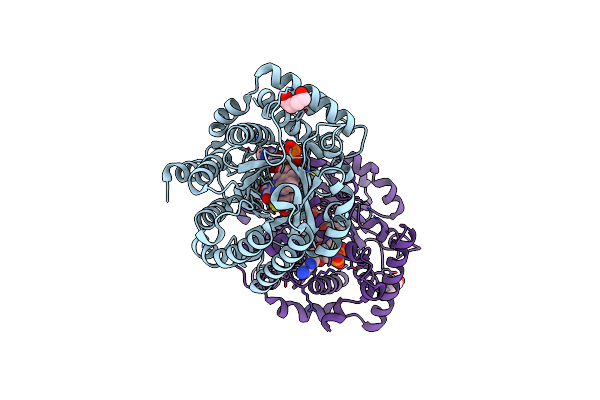
Minimal Puta Proline Dehydrogenase Domain (Design #2) Complexed With 2-(Cyanomethylthio)Acetic Acid
Organism: Sinorhizobium meliloti sm11
Method: X-RAY DIFFRACTION Resolution:1.43 Å Release Date: 2024-10-30 Classification: OXIDOREDUCTASE Ligands: FAD, X7K, PEG |
 |
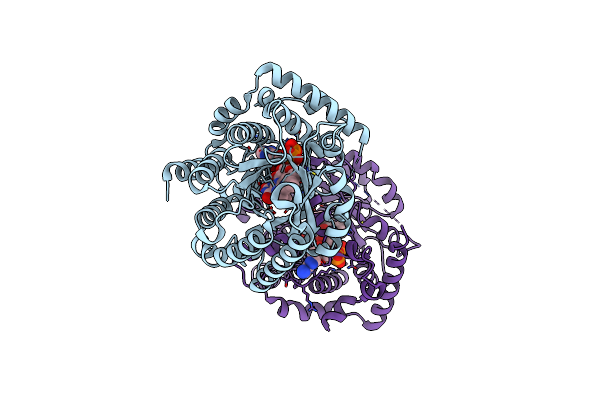
Minimal Puta Proline Dehydrogenase Domain (Design #2) Complexed With (Allylthio)Acetic Acid
Organism: Sinorhizobium meliloti sm11
Method: X-RAY DIFFRACTION Resolution:1.41 Å Release Date: 2024-10-30 Classification: OXIDOREDUCTASE Ligands: FAD, PEG, X7Q |
 |
Minimal Puta Proline Dehydrogenase Domain (Design #2) With The Fad N5 Modified With Propanal Resulting From Inactivation With N-Allylglycine(Replicate #1)
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.54 Å Release Date: 2024-10-30 Classification: OXIDOREDUCTASE Ligands: FDA, FAD, CBG |

