Search Count: 68
All
Selected
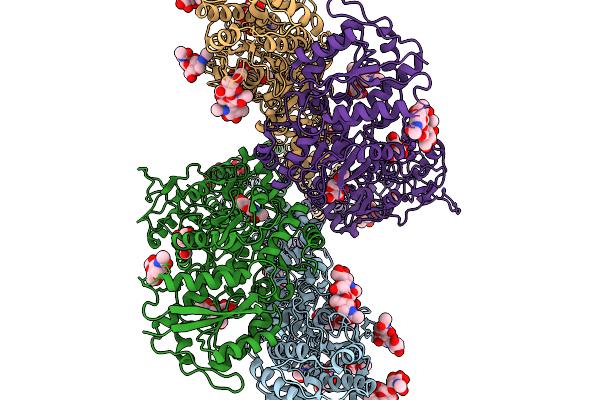 |
Cryoem Structure Of Gluk2 Bound To Glutamate In The Deep Desensitized State 3
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Resolution:3.83 Å Release Date: 2026-02-04 Classification: STRUCTURAL PROTEIN Ligands: NAG |
 |
Cryoem Structure Of Gluk2 Bound To Glutamate In The Deep Desensitized State 2
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Resolution:3.74 Å Release Date: 2026-02-04 Classification: STRUCTURAL PROTEIN Ligands: NAG |
 |
Cryoem Structure Of Gluk2 Bound To Glutamate In The Deep Desensitized State 1
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Resolution:3.81 Å Release Date: 2026-02-04 Classification: STRUCTURAL PROTEIN Ligands: NAG |
 |
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Resolution:4.02 Å Release Date: 2026-02-04 Classification: STRUCTURAL PROTEIN Ligands: NAG |
 |
Cryoem Structure Of Gluk2 Bound To Glutamate In The Shallow Desensitized State, Consensus Map
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Resolution:3.86 Å Release Date: 2026-02-04 Classification: STRUCTURAL PROTEIN Ligands: NAG, GLU |
 |
Cryoem Structure Of Gluk2 Bound To Glutamate In The Shallow Desensitized State, Composite Map
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Resolution:3.80 Å Release Date: 2026-02-04 Classification: STRUCTURAL PROTEIN Ligands: NAG, GLU |
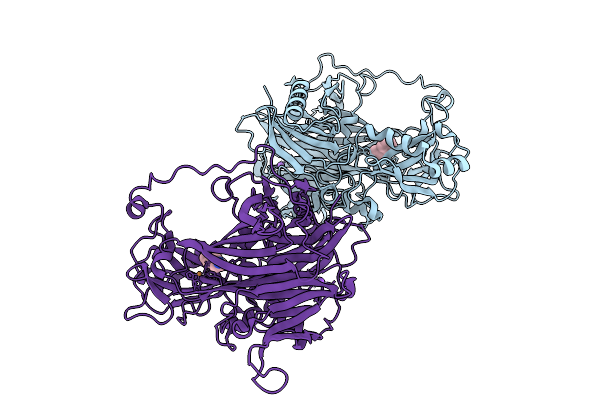 |
Reductive-Half Reaction Intermediate Of Copper Amine Oxidase From Arthrobacter Globiformis Captured By Mix-And-Inject Serial Crystallography At 100-Ms Time Delay
Organism: Arthrobacter globiformis
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-12-31 Classification: OXIDOREDUCTASE Ligands: PEA, NA, CU |
 |
Reductive-Half Reaction Intermediate Of Copper Amine Oxidase From Arthrobacter Globiformis Captured By Mix-And-Inject Serial Crystallography At 200-Ms Time Delay
Organism: Arthrobacter globiformis
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2025-12-31 Classification: OXIDOREDUCTASE Ligands: PEA, NA, CU |
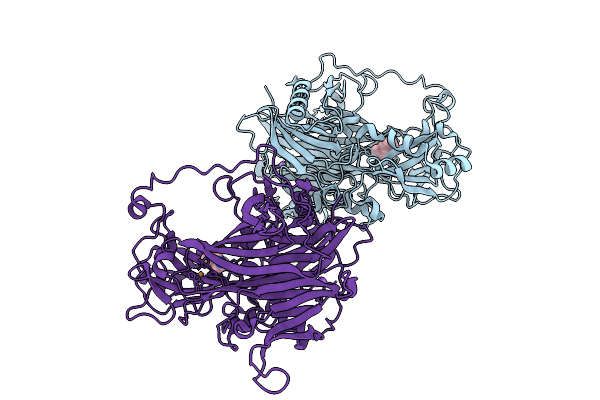 |
Reductive-Half Reaction Intermediate Of Copper Amine Oxidase From Arthrobacter Globiformis Captured By Mix-And-Inject Serial Crystallography At 300-Ms Time Delay
Organism: Arthrobacter globiformis
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2025-12-31 Classification: OXIDOREDUCTASE |
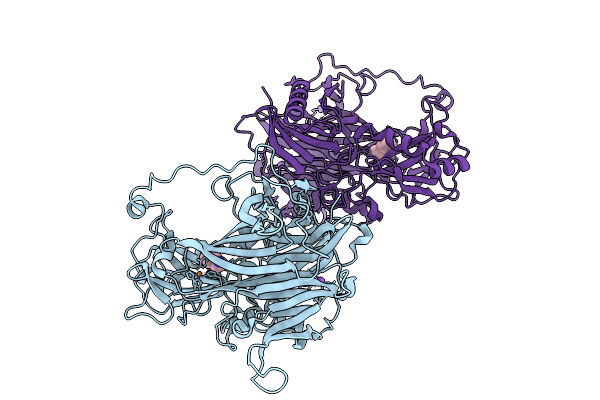 |
Reductive-Half Reaction Intermediate Of Copper Amine Oxidase From Arthrobacter Globiformis Captured By Mix-And-Inject Serial Crystallography At 400-Ms Time Delay
Organism: Arthrobacter globiformis
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2025-12-31 Classification: OXIDOREDUCTASE Ligands: PEA, NA, CU |
 |
Reductive-Half Reaction Intermediate Of Copper Amine Oxidase From Arthrobacter Globiformis Captured With Short-A-Axis Diffraction Data By Mix-And-Inject Serial Crystallography At 22-Ms Time Delay
Organism: Arthrobacter globiformis
Method: X-RAY DIFFRACTION Resolution:2.22 Å Release Date: 2025-12-31 Classification: OXIDOREDUCTASE Ligands: CU, NA |
 |
Reductive-Half Reaction Intermediate Of Copper Amine Oxidase From Arthrobacter Globiformis Captured With Long-A-Axis Diffraction Data By Mix-And-Inject Serial Crystallography At 22-Ms Time Delay
Organism: Arthrobacter globiformis
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-12-31 Classification: OXIDOREDUCTASE Ligands: CU, NA |
 |
Reductive-Half Reaction Intermediate Of Copper Amine Oxidase From Arthrobacter Globiformis Captured With Short-A-Axis Diffraction Data By Mix-And-Inject Serial Crystallography At 25-Ms Time Delay
Organism: Arthrobacter globiformis
Method: X-RAY DIFFRACTION Resolution:2.13 Å Release Date: 2025-12-31 Classification: OXIDOREDUCTASE Ligands: CU |
 |
Reductive-Half Reaction Intermediate Of Copper Amine Oxidase From Arthrobacter Globiformis Captured With Long-A-Axis Diffraction Data By Mix-And-Inject Serial Crystallography At 25-Ms Time Delay
Organism: Arthrobacter globiformis
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2025-12-31 Classification: OXIDOREDUCTASE Ligands: CU, NA |
 |
Reductive-Half Reaction Intermediate Of Copper Amine Oxidase From Arthrobacter Globiformis Captured With Short-A-Axis Diffraction Data By Mix-And-Inject Serial Crystallography At 50-Ms Time Delay
Organism: Arthrobacter globiformis
Method: X-RAY DIFFRACTION Resolution:1.92 Å Release Date: 2025-12-31 Classification: OXIDOREDUCTASE Ligands: CU |
 |
Reductive-Half Reaction Intermediate Of Copper Amine Oxidase From Arthrobacter Globiformis Captured With Long-A-Axis Diffraction Data By Mix-And-Inject Serial Crystallography At 50-Ms Time Delay
Organism: Arthrobacter globiformis
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2025-12-31 Classification: OXIDOREDUCTASE Ligands: CU, NA |
 |
Oxidative Form Of Copper Amine Oxidase From Arthrobacter Globiformis Determined At Ph 9.0 By Single-Flow Serial Crystallography
Organism: Arthrobacter globiformis
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-12-31 Classification: OXIDOREDUCTASE Ligands: CU, NA |
 |
Reductive-Half Reaction Intermediate Of Copper Amine Oxidase From Arthrobacter Globiformis Captured By Mix-And-Inject Serial Crystallography At 500-Ms Time Delay
Organism: Arthrobacter globiformis
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2025-12-31 Classification: OXIDOREDUCTASE Ligands: CU, NA, PEA |
 |
Reductive-Half Reaction Intermediate Of Copper Amine Oxidase From Arthrobacter Globiformis Captured By Mix-And-Inject Serial Crystallography At 1000-Ms Time Delay
Organism: Arthrobacter globiformis
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2025-12-31 Classification: OXIDOREDUCTASE Ligands: CU, NA, PEA |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2025-07-30 Classification: NUCLEAR PROTEIN Ligands: A1MAM, EDO, ACT |

