Search Count: 1,233
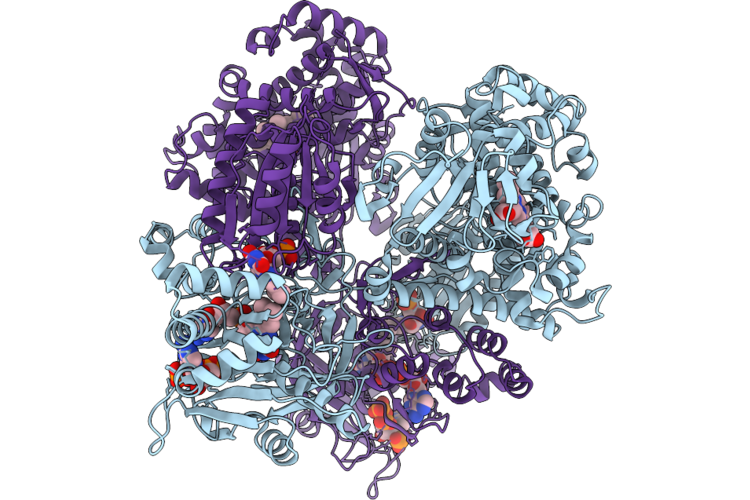 |
Cryo-Em Structure Of Full-Length Self-Sufficient P450 In Complex With Nadph From Shimazuella Soli
Organism: Shimazuella soli
Method: ELECTRON MICROSCOPY Resolution:2.68 Å Release Date: 2026-05-27 Classification: OXIDOREDUCTASE Ligands: HEM, FMN, FAD, NDP |
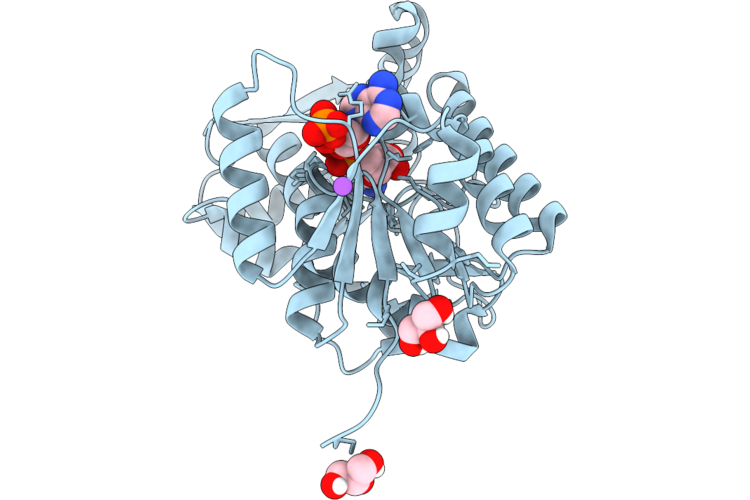 |
Organism: Illicium verum
Method: X-RAY DIFFRACTION Resolution:2.33 Å Release Date: 2026-05-20 Classification: PLANT PROTEIN Ligands: GOL, NDP, NA |
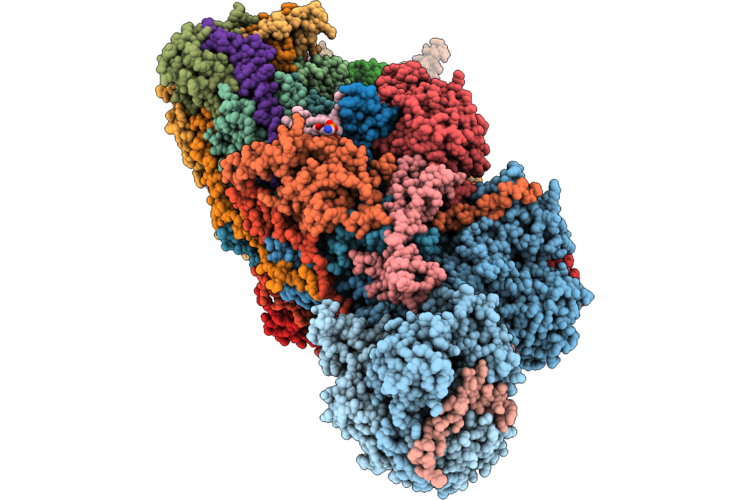 |
Organism: Ovis aries
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: PROTON TRANSPORT Ligands: 3PE, PC1, DGT, MG, MYR, ZMP, SF4, FMN, NAI, FES, K, A1JBT, ZN, NDP |
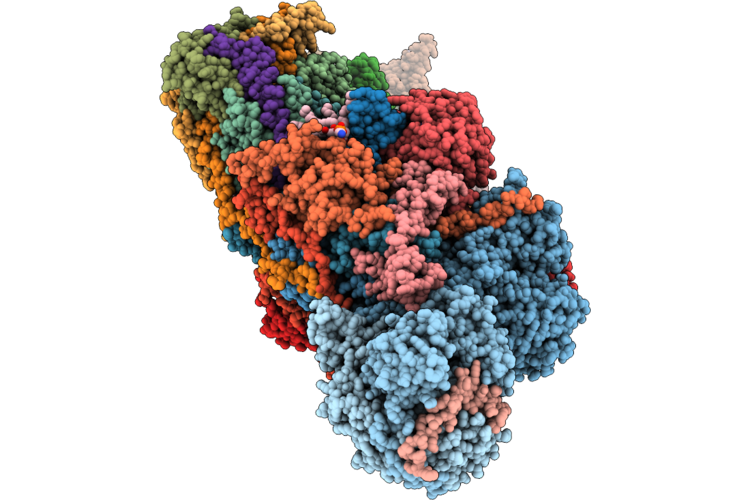 |
Organism: Ovis aries
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: PROTON TRANSPORT Ligands: 3PE, PC1, A1JBT, SF4, DGT, MG, MYR, ZMP, FMN, NAI, FES, K, ZN, NDP |
 |
Crystal Structure Of An Nta622L Variant In Complex With Nadph And Nicotinic Acid N-Glucoside
Organism: Nicotiana tabacum
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-05-06 Classification: PLANT PROTEIN Ligands: A1JBN, NDP |
 |
Organism: Plasmodium falciparum hb3
Method: X-RAY DIFFRACTION Resolution:1.69 Å Release Date: 2026-05-06 Classification: ISOMERASE Ligands: NDP, MN, A1L91, CA |
 |
Organism: Plasmodium falciparum hb3
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2026-05-06 Classification: ISOMERASE Ligands: NDP, MN, A1L92, CA |
 |
Crystal Structure Of An Nta622L Variant In Complex With Nadph And Trigonelline
Organism: Nicotiana tabacum
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-04-29 Classification: PLANT PROTEIN Ligands: NDP, A1JBP |
 |
Organism: Aspergillus fumigatus af293
Method: ELECTRON MICROSCOPY Resolution:3.36 Å Release Date: 2026-04-08 Classification: BIOSYNTHETIC PROTEIN Ligands: COZ, NDP |
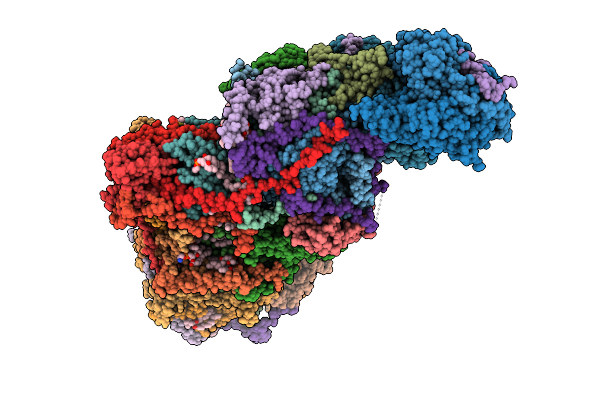 |
Cryo-Em Structure Of Complex I On The Bovine Heart Submitochondrial Particles, Open
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: FME, PC1, SF4, FES, FMN, K, 3PE, CDL, CHD, GTP, MG, NDP, ZN, MYR |
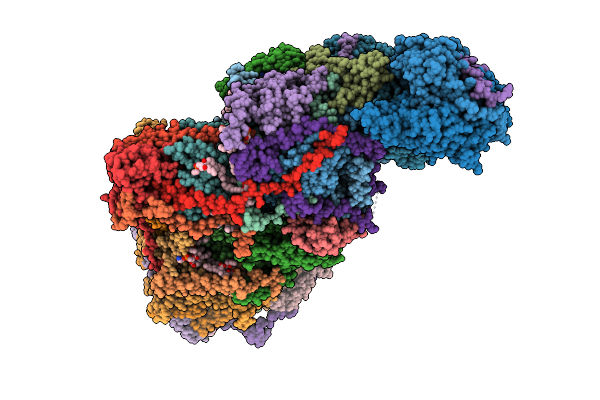 |
Cryo-Em Structure Of Complex I On The Bovine Heart Submitochondrial Particles, Closed
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: FME, 3PE, PC1, SF4, FES, FMN, K, CDL, MG, GTP, NDP, ZN |
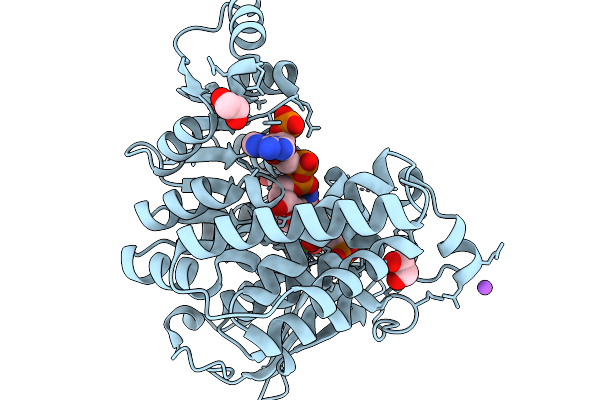 |
1-Deoxy-D-Xylulose 5-Phosphate Reductoisomerase (Ispc) From Acinetobacter Baumannii In Complex With A Fosmidomycin Analog, Nadph, And A Magnesium Ion
Organism: Acinetobacter baumannii
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: EDO, NDP, A1CFA, MG, CL, NA |
 |
1-Deoxy-D-Xylulose 5-Phosphate Reductoisomerase (Ispc) From Acinetobacter Baumannii In Complex With A Fosmidomycin Analog, Nadph, And A Magnesium Ion
Organism: Acinetobacter baumannii
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: NDP, A1CFB, EDO, MG, CL |
 |
1-Deoxy-D-Xylulose 5-Phosphate Reductoisomerase (Ispc) From Acinetobacter Baumannii In Complex With A Fosmidomycin Analog, Nadph, And A Magnesium Ion
Organism: Acinetobacter baumannii
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: NDP, A1CE6, EDO, MG |
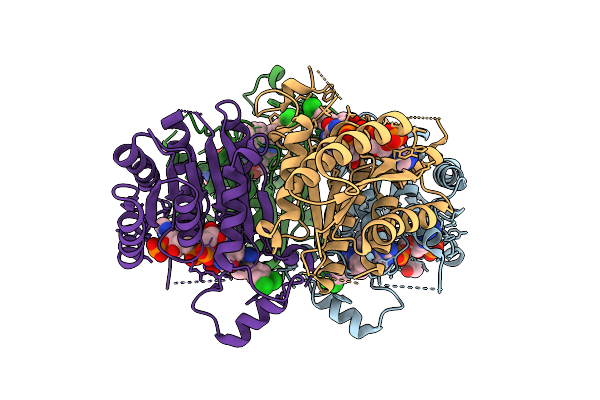 |
Organism: Trypanosoma brucei brucei
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: NDP, QY8, AX8, ACT, GOL |
 |
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: ELECTRON TRANSPORT Ligands: 3PE, PC1, SF4, U10, FES, FMN, NAI, K, CDL, DGT, MG, NDP, ZN, EHZ, CHD, MYR |
 |
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: ELECTRON TRANSPORT Ligands: PC1, SF4, FES, FMN, NAI, K, U10, 3PE, CDL, DGT, MG, NDP, ZN, EHZ, CHD, MYR |
 |
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: ELECTRON TRANSPORT Ligands: 3PE, PC1, SF4, FES, NAI, FMN, K, LMT, CDL, DGT, MG, NDP, ZN, EHZ, CHD, MYR |
 |
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: ELECTRON TRANSPORT Ligands: 3PE, PC1, SF4, LMT, FES, FMN, NAI, K, CDL, CHD, DGT, MG, NDP, ZN, EHZ, MYR |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: MEMBRANE PROTEIN Ligands: SF4, FMN, PEE, 8Q1, NDP, FES, PLX, ZN, CDL, DGT, MG |

