Search Count: 112
All
Selected
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.46 Å Release Date: 2026-02-18 Classification: DE NOVO PROTEIN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-02-18 Classification: DE NOVO PROTEIN |
 |
Organism: Streptomyces klenkii
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2026-01-21 Classification: HYDROLASE |
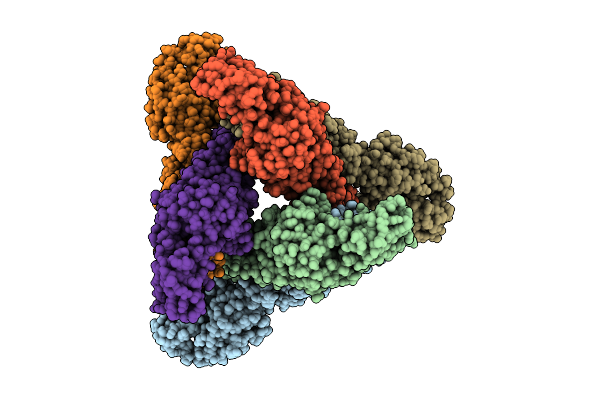 |
Organism: Western equine encephalitis virus
Method: ELECTRON MICROSCOPY Resolution:3.05 Å Release Date: 2025-11-26 Classification: VIRAL PROTEIN |
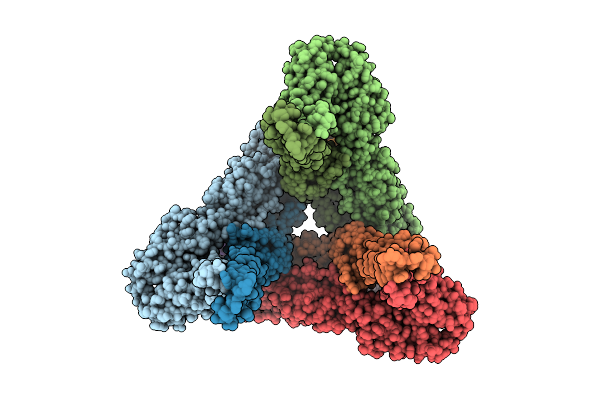 |
Cryoem Structure Of Western Equine Encephalitis Virus California Vlp In Complex With Vldlr-La1-3
Organism: Western equine encephalitis virus, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.03 Å Release Date: 2025-11-26 Classification: VIRAL PROTEIN Ligands: CA |
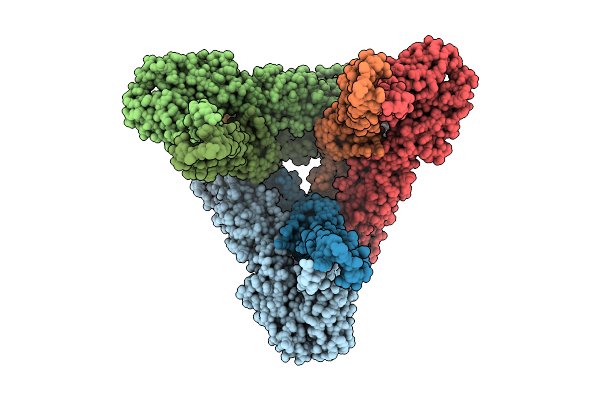 |
Cryoem Structure Of Western Equine Encephalitis Virus California Vlp In Complex With Vldlr-Lbd
Organism: Western equine encephalitis virus, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.27 Å Release Date: 2025-11-26 Classification: VIRAL PROTEIN Ligands: CA |
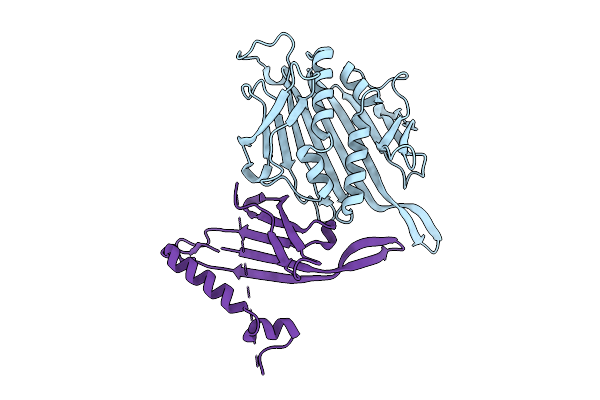 |
Organism: Escherichia phage ms2
Method: ELECTRON MICROSCOPY Resolution:2.62 Å Release Date: 2025-11-05 Classification: VIRUS LIKE PARTICLE |
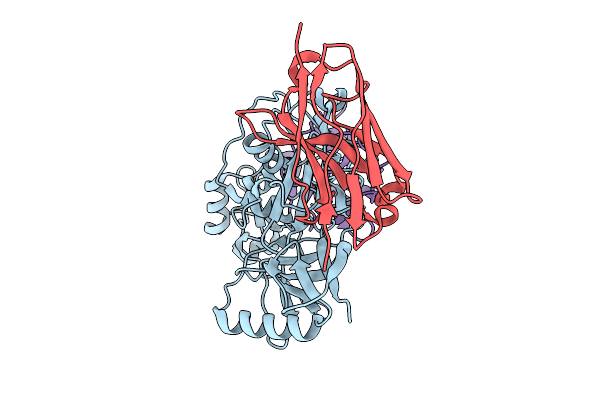 |
Organism: Severe fever with thrombocytopenia syndrome virus, Vicugna pacos
Method: ELECTRON MICROSCOPY Release Date: 2025-10-29 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2025-09-24 Classification: TRANSPORT PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: TRANSPORT PROTEIN Ligands: A1EIM |
 |
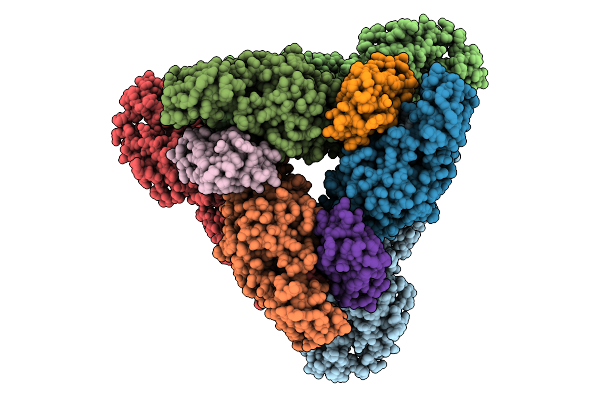
Organism: Western equine encephalitis virus
Method: ELECTRON MICROSCOPY Resolution:2.53 Å Release Date: 2025-08-27 Classification: VIRAL PROTEIN |
 |
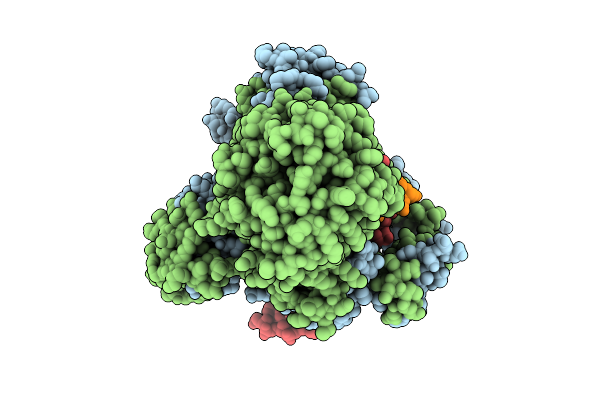
Structure Of Weev Strain 71V1658 Virus-Like Particles (Vlps) In Complex With Human Pcdh10 Extracellular Cadherin Repeats 1-2 (Ec1-Ec2)(3-Fold Region)
Organism: Western equine encephalitis virus, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-08-27 Classification: VIRAL PROTEIN |
 |
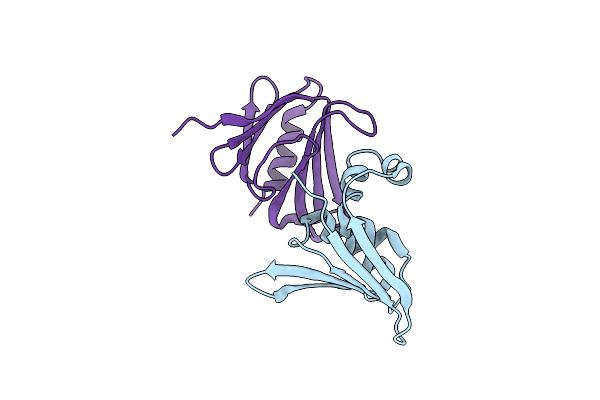
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-09 Classification: DNA BINDING PROTEIN/DNA Ligands: IHP, 2OP |
 |
Organism: Taylorella equigenitalis (strain mce9)
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2024-11-27 Classification: HYDROLASE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2024-10-16 Classification: SIGNALING PROTEIN Ligands: BA |
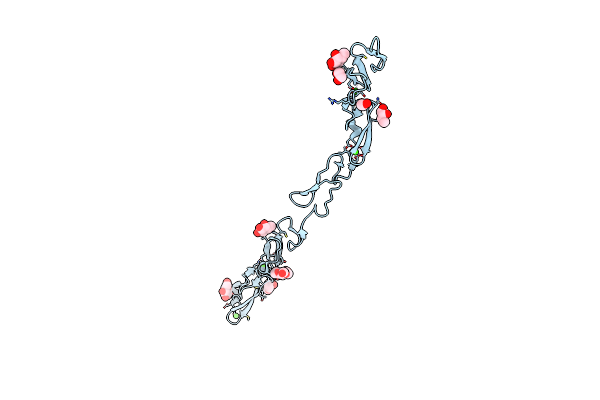 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2024-10-16 Classification: SIGNALING PROTEIN Ligands: BGC, FUC, CA, EDO |
 |
Organism: Homo sapiens, Oryctolagus cuniculus
Method: ELECTRON MICROSCOPY Release Date: 2024-05-22 Classification: IMMUNE SYSTEM Ligands: POV, HEM, NAG, LBN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2023-08-02 Classification: HYDROLASE/INHIBITOR Ligands: CL, NA, V2O, EDO |
 |
Organism: Papaver somniferum
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2023-03-01 Classification: BIOSYNTHETIC PROTEIN |
 |
Organism: Papaver somniferum
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2023-03-01 Classification: BIOSYNTHETIC PROTEIN Ligands: EV1 |

