Search Count: 30
All
Selected
 |
Endo-Deglycosylated Hydroxynitrile Lyase Isozyme 5 Mutant L331A From Prunus Communis Complexed With 2,2-Dimethyl-4H-Benzo[D][1,3]Dioxine-6-Carbaldehyde From The Cyanohydrin Cleavage
Organism: Prunus dulcis
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2024-06-12 Classification: LYASE Ligands: NAG, FQ3, FAD, PGE, PEG, BCN |
 |
Endo-Deglycosylated Hydroxynitrile Lyase Isozyme 5 Mutant From Prunus Communis Mutant L331A Complexed With 2,2-Dimethyl-4H-Benzo[D][1,3]Dioxine-6-Carbaldehyde (Form A)
Organism: Prunus dulcis
Method: X-RAY DIFFRACTION Resolution:2.31 Å Release Date: 2024-06-12 Classification: LYASE Ligands: NAG, FQ3, FAD, EDO, PEG, BCN |
 |
Endo-Deglycosylated Hydroxynitrile Lyase Isozyme 5 Mutant L331A From Prunus Communis Complexed With 2,2-Dimethyl-4H-Benzo[D][1,3]Dioxine-6-Carbaldehyde (Form B)
Organism: Prunus dulcis
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2024-06-12 Classification: LYASE Ligands: NAG, FQ3, PGE, FAD, PEG, EDO |
 |
Endo-Deglycosylated Hydroxynitrile Lyase Isozyme 5 Mutant L331A From Prunus Communis Complexed With 4H-Benzo[D][1,3]Dioxine-6-Carbaldehyde
Organism: Prunus dulcis
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2024-06-12 Classification: LYASE Ligands: NAG, FQ6, FAD, PEG, PO4 |
 |
Endo-Deglycosylated Hydroxynitrile Lyase Isozyme 5 Mutant L331A From Prunus Communis Complexed With 2-Methyl-4H-Benzo[D][1,3]Dioxine-6-Carbaldehyde
Organism: Prunus dulcis
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2024-06-05 Classification: LYASE Ligands: FQ9, FAD, NAG, PEG, DMS |
 |
Endo-Deglycosylated Hydroxynitrile Lyase Isozyme 5 Mutant L331A/S333V/P340L From Prunus Communis
Organism: Prunus dulcis
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2024-06-05 Classification: LYASE Ligands: FAD, PEG, NAG, EDO |
 |
Endo-Deglycosylated Hydroxynitrile Lyase Isozyme 5 Mutant L331A/S333V/P340L From Prunus Communis Complexed With 2,2-Dimethyl-4H-Benzo[D][1,3]Dioxine-6-Carbaldehyde (Catalytic Conformation)
Organism: Prunus dulcis
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2024-06-05 Classification: LYASE Ligands: FQ3, FAD, NAG, PO4 |
 |
Endo-Deglycosylated Hydroxynitrile Lyase Isozyme 5 Mutant L331A/S333V/P340L From Prunus Communis Complexed With 2,2-Dimethyl-4H-Benzo[D][1,3]Dioxine-6-Carbaldehyde (Noncatalytic Conformation)
Organism: Prunus dulcis
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2024-06-05 Classification: LYASE Ligands: FQ3, FAD, NAG, PEG, PO4 |
 |
Endo-Deglycosylated Hydroxynitrile Lyase Isozyme 5 Mutant L331A From Prunus Communis Complexed With (R)-2-(2,2-Dimethyl-4H-Benzo[D][1,3]Dioxin-6-Yl)-2-Hydroxyacetonitrile
Organism: Prunus dulcis
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2024-06-05 Classification: LYASE Ligands: E2Y, NAG, FAD, PEG, EDO |
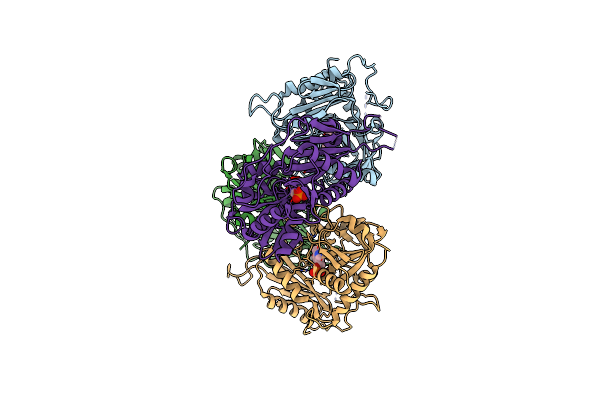 |
Organism: Rhodobacter sp. 140a
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2023-02-08 Classification: TRANSFERASE Ligands: PLP |
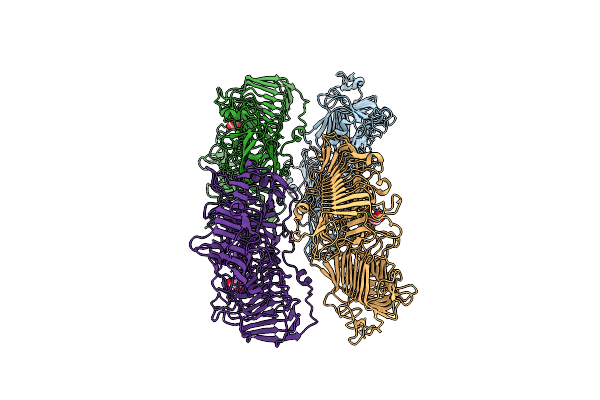 |
Organism: Bacteroides clarus
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2021-10-06 Classification: LYASE Ligands: CA, MLI |
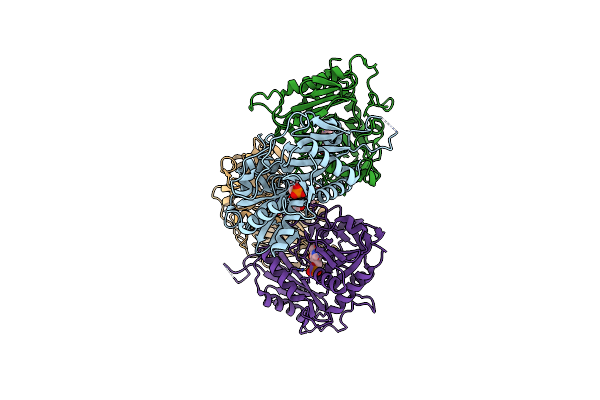 |
Organism: Rhodobacter sp. 140a
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2021-09-01 Classification: TRANSFERASE Ligands: PLP, EDO |
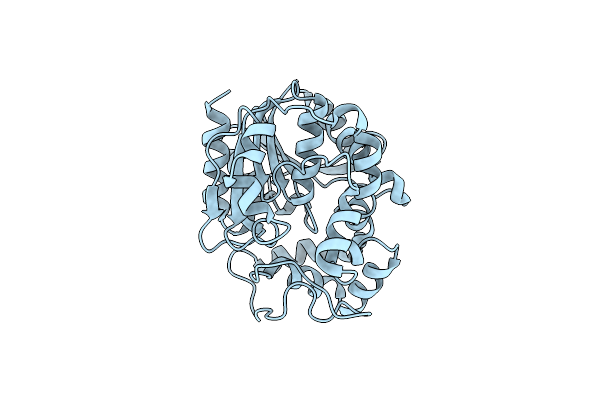 |
Organism: Vigna radiata
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2021-07-07 Classification: HYDROLASE |
 |
Organism: Vigna radiata
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2021-07-07 Classification: HYDROLASE |
 |
Crystal Complex Of Endo-Deglycosylated Hydroxynitrile Lyase Isozyme 5 Of Prunus Communis With Benzaldehyde
Organism: Prunus dulcis
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2020-05-13 Classification: LYASE Ligands: NAG, FAD, HBX |
 |
Mutant L331A Of Deglycosylated Hydroxynitrile Lyase Isozyme 5 From Prunus Communis
Organism: Prunus dulcis
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2020-05-13 Classification: LYASE Ligands: NAG, FAD |
 |
Organism: Prunus dulcis
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2020-05-13 Classification: LYASE Ligands: MXN, NAG, FAD, PO4, PEG |
 |
Crystal Structure Of Endo-Deglycosylated Hydroxynitrile Lyase Isozyme 5 Of Prunus Communis
Organism: Prunus dulcis
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2020-04-01 Classification: LYASE Ligands: NAG, FAD |
 |
Crystal Structure Of The Catalytic Domain Of A Multi-Domain Alginate Lyase Dp0100 From Thermophilic Bacterium Defluviitalea Phaphyphila
Organism: Defluviitalea phaphyphila
Method: X-RAY DIFFRACTION Resolution:2.07 Å Release Date: 2019-10-30 Classification: LYASE Ligands: MN, MG, CA, ACT, EDO, CAC |
 |
Crystal Structure Of The Catalytic Domain Of A Multi-Domain Alginate Lyase Dp0100 From Thermophilic Bacterium Defluviitalea Phaphyphila
Organism: Defluviitalea phaphyphila
Method: X-RAY DIFFRACTION Resolution:2.76 Å Release Date: 2019-10-30 Classification: LYASE Ligands: MN, CA, ACT, MG |

