Search Count: 553
All
Selected
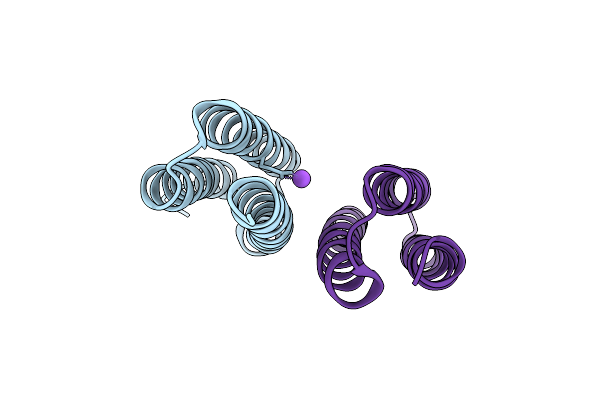 |
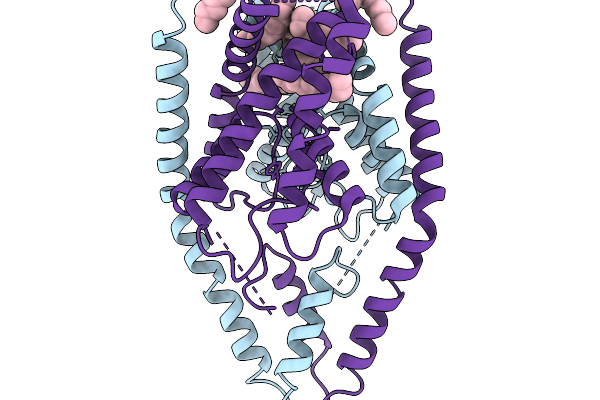
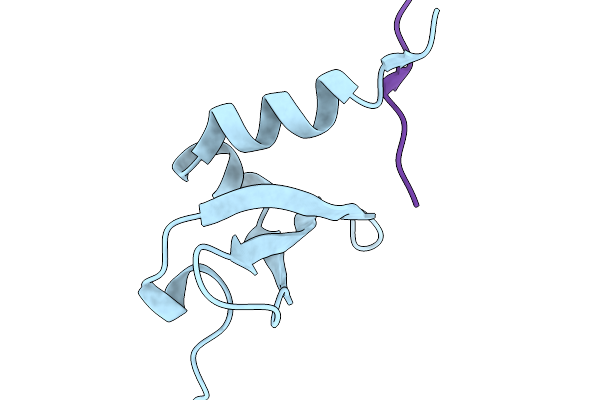
Mini-Protein Binder Of N-Terminal Domain Of Nucleocapsid Protein Of Sars-Cov2
Organism: Unidentified
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-02-25 Classification: DE NOVO PROTEIN |
 |
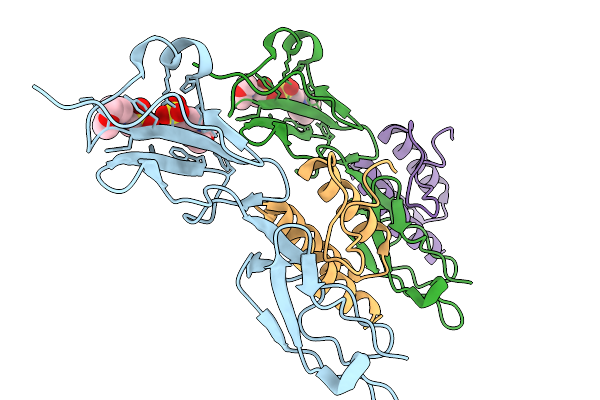
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.67 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN Ligands: D10, K |
 |
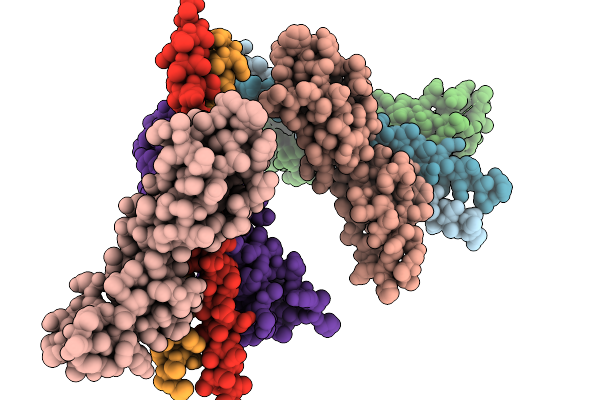
Organism: Homo sapiens, Streptococcus pyogenes
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2026-02-04 Classification: IMMUNE SYSTEM Ligands: MES |
 |
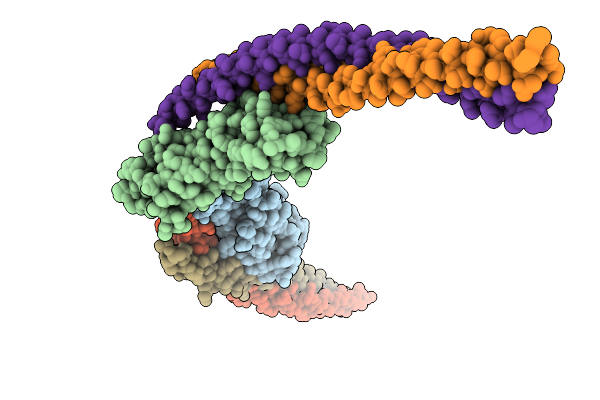
Organism: Streptococcus pyogenes str. manfredo, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2026-02-04 Classification: IMMUNE SYSTEM |
 |
Organism: Homo sapiens, Streptococcus pyogenes jrs4
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-02-04 Classification: IMMUNE SYSTEM Ligands: EDO |
 |
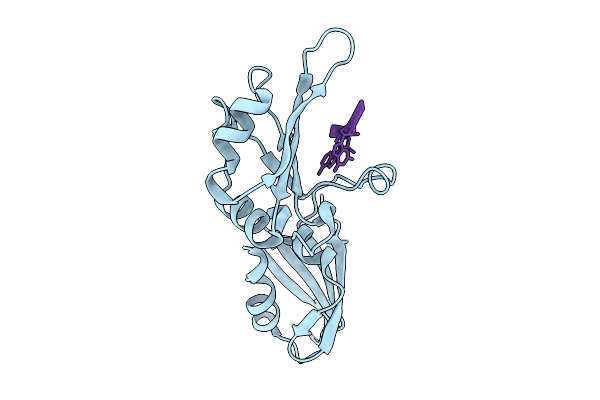
Crystal Structure Of Hp1Gamma Chromoshadow Domain In Complex With Kap1 Peptide
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2026-02-04 Classification: TRANSCRIPTION |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-01-28 Classification: RNA BINDING PROTEIN/RNA |
 |
Organism: Hepatitis c virus (isolate 1)
Method: SOLUTION NMR Release Date: 2026-01-28 Classification: RNA |
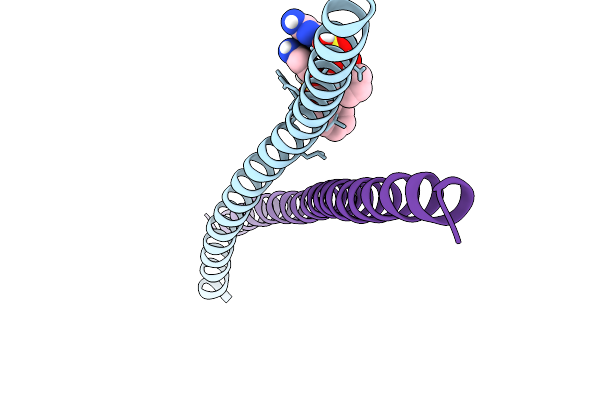 |
Transcription Factor Deltafosb/Jund Bzip Domain In Complex With An Effector Molecule
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-01-07 Classification: CELL ADHESION Ligands: A1CAO, CL |
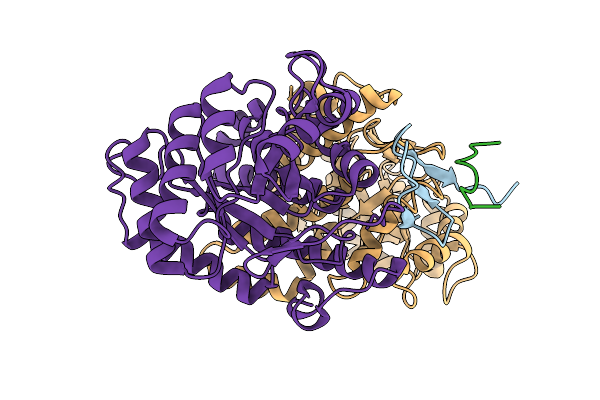 |
Structure Of The Energy Converting Methyltransferase (Mtr) Of Methanosarcina Mazei In Complex With A Novel Protein Binder
Organism: Methanosarcina mazei go1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN |
 |
Structure Of The Energy Converting Methyltransferase (Mtr) Of Methanosarcina Mazei In Complex With A Novel Protein Binder
Organism: Methanosarcina mazei go1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: A1JAP, L1P, A1JAO, NA, A1JAN |
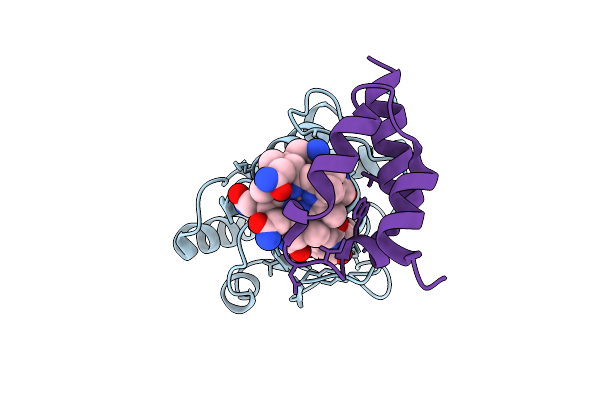 |
Structure Of The Energy Converting Methyltransferase (Mtr) Of Methanosarcina Mazei In Complex With A Novel Protein Binder
Organism: Methanosarcina mazei go1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: B13 |
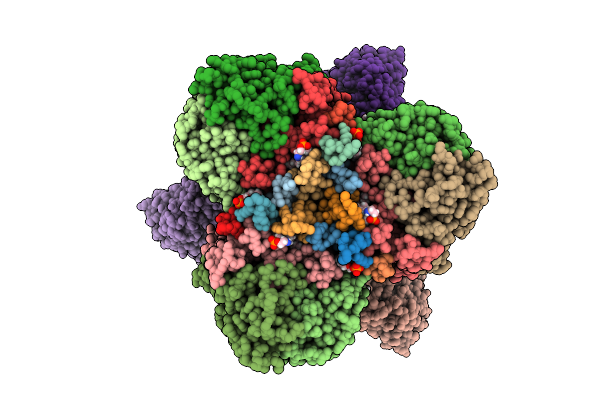 |
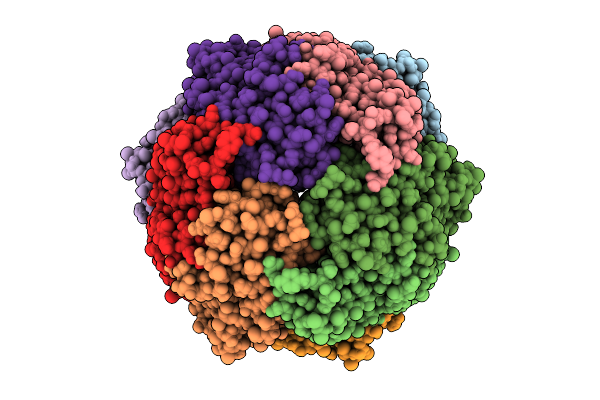
Structure Of The Energy Converting Methyltransferase (Mtr) Of Methanosarcina Mazei In Complex With A Novel Protein Binder
Organism: Methanosarcina mazei go1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: B13, A1JAP, L1P, A1JAO, NA, A1JAN |
 |
Structure Of E. Coli Dna Protection During Starvation Protein (Dps) From Single Particle Cryoem
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:1.30 Å Release Date: 2025-12-24 Classification: DNA BINDING PROTEIN Ligands: FE |
 |
Crystal Structure Of Proline Dipeptidase From Pyrococcus Furiosus Complexed With Co
Organism: Pyrococcus furiosus dsm 3638
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-12-10 Classification: HYDROLASE |
 |
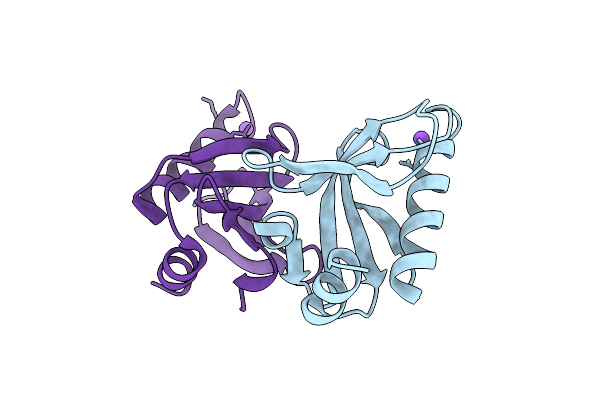
Crystal Structure Of Proline Dipeptidase From Pyrococcus Furiosus Complexed With Mn
Organism: Pyrococcus furiosus dsm 3638
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-12-10 Classification: HYDROLASE Ligands: MN, NA, PEG |
 |
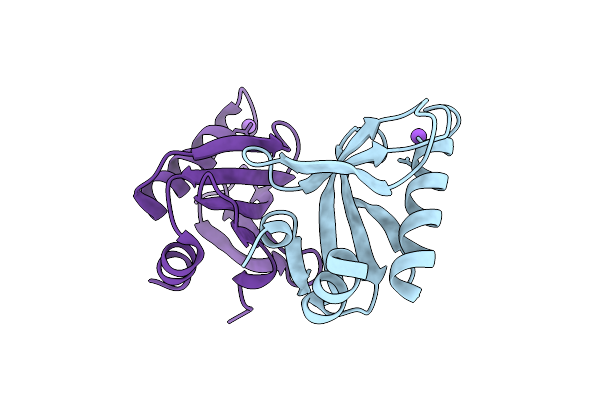
Crystal Structure Of Upf0235 Protein Pf1765 From Pyrococcus Furiosus Crystallized With Sodium Acetate At Ph 8
Organism: Pyrococcus furiosus dsm 3638
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2025-11-26 Classification: UNKNOWN FUNCTION |
 |
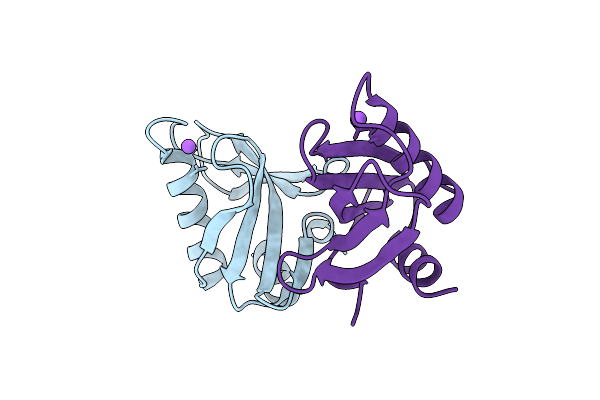
Crystal Structure Of Upf0235 Protein Pf1765 From Pyrococcus Furiosus At Ph 6
Organism: Pyrococcus furiosus dsm 3638
Method: X-RAY DIFFRACTION Resolution:1.13 Å Release Date: 2025-11-26 Classification: UNKNOWN FUNCTION |
 |
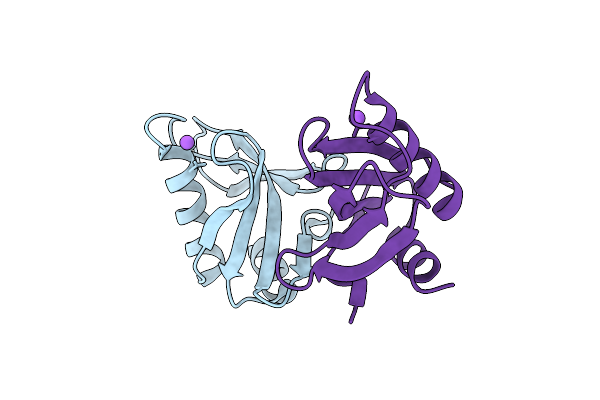
Crystal Structure Of Upf0235 Protein Pf1765 From Pyrococcus Furiosus At Ph 9
Organism: Pyrococcus furiosus dsm 3638
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2025-11-26 Classification: UNKNOWN FUNCTION |
 |
Crystal Structure Of Upf0235 Protein Pf1765 From Pyrococcus Furiosus Crystallized With Sodium Citrate At Ph 8.0
Organism: Pyrococcus furiosus dsm 3638
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2025-11-26 Classification: UNKNOWN FUNCTION |

