Search Count: 23
All
Selected
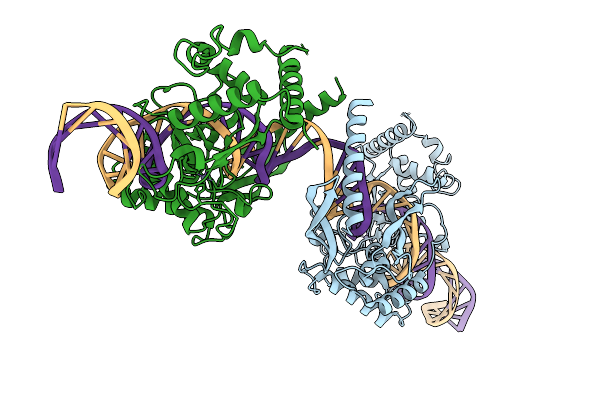 |
Organism: Streptomyces phage phi-c31, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN Ligands: ZN |
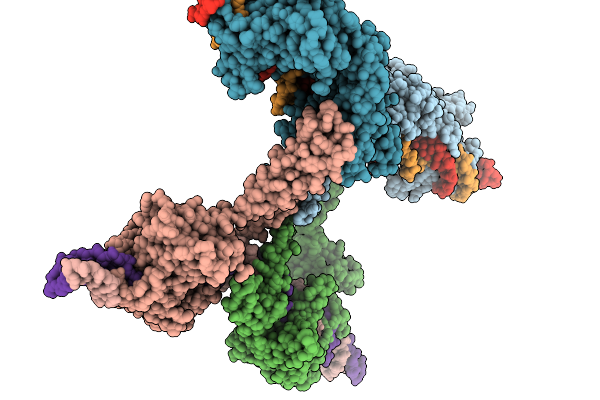 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: DNA BINDING PROTEIN Ligands: ZN |
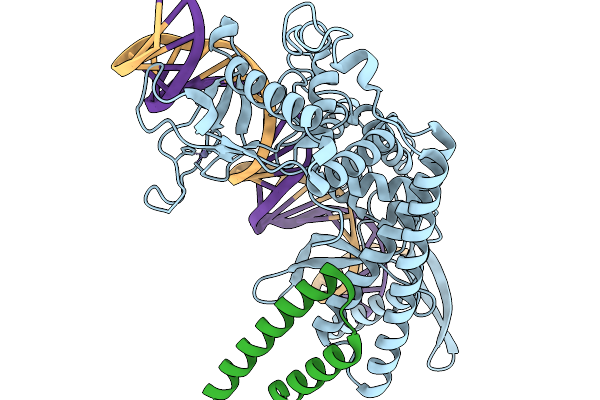 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: DNA BINDING PROTEIN Ligands: ZN |
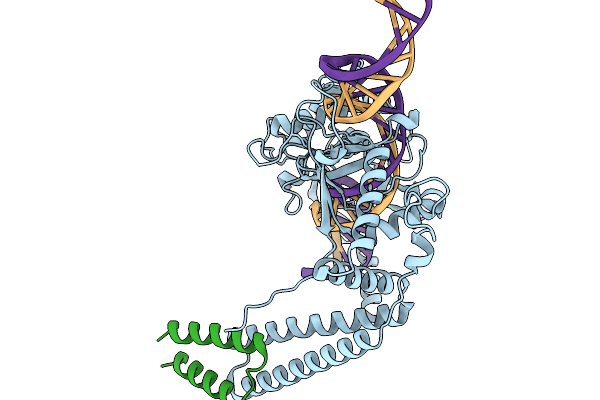 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: DNA BINDING PROTEIN Ligands: ZN |
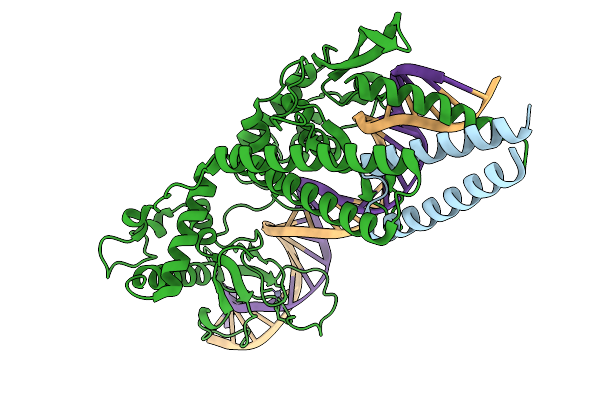 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: DNA BINDING PROTEIN Ligands: ZN |
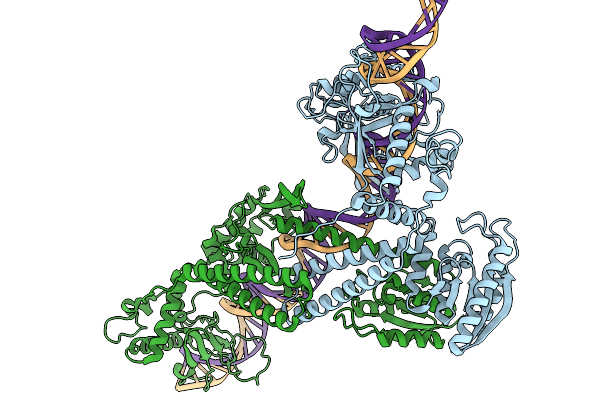 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: DNA BINDING PROTEIN Ligands: ZN |
 |
Phic31 Excisive Synapse; Attl X Attl With An Integrases12A-Rdf Protein Fusion
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: RECOMBINATION Ligands: ZN |
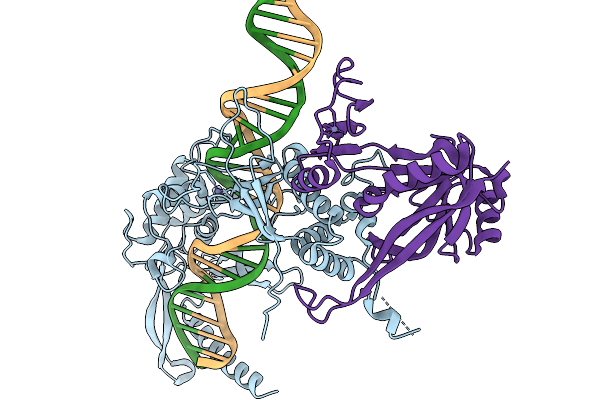 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: RECOMBINATION Ligands: ZN |
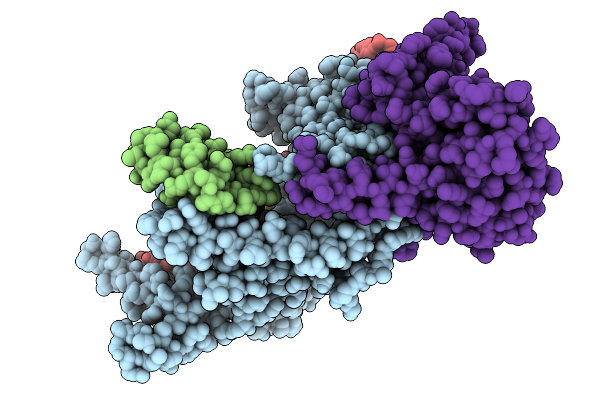 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: RECOMBINATION Ligands: ZN |
 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: RECOMBINATION Ligands: ZN |
 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: RECOMBINATION Ligands: ZN |
 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: RECOMBINATION Ligands: ZN |
 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: RECOMBINATION Ligands: ZN |
 |
Organism: Lomovskayavirus c31
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: RECOMBINATION Ligands: ZN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:5.60 Å Release Date: 2018-12-19 Classification: IMMUNE SYSTEM Ligands: NAG |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:5.60 Å Release Date: 2018-12-19 Classification: IMMUNE SYSTEM Ligands: NAG |
 |
Organism: Homo sapiens, Bos taurus, Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2017-10-25 Classification: TRANSPORT PROTEIN Ligands: ZN, MG, GTP, GDP, TA1 |
 |
Organism: Homo sapiens, Bos taurus, Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2017-10-25 Classification: TRANSPORT PROTEIN Ligands: MG, ANP, ZN, GTP, GDP, TA1 |
 |
Organism: Homo sapiens, Bos taurus, Sus scrofa
Method: ELECTRON MICROSCOPY Resolution:4.80 Å Release Date: 2017-10-25 Classification: TRANSPORT PROTEIN Ligands: 9V5, ZN, MG, GTP, GDP, TA1 |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2017-10-04 Classification: STRUCTURAL PROTEIN |

