Search Count: 31
All
Selected
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: RNA Ligands: MG |
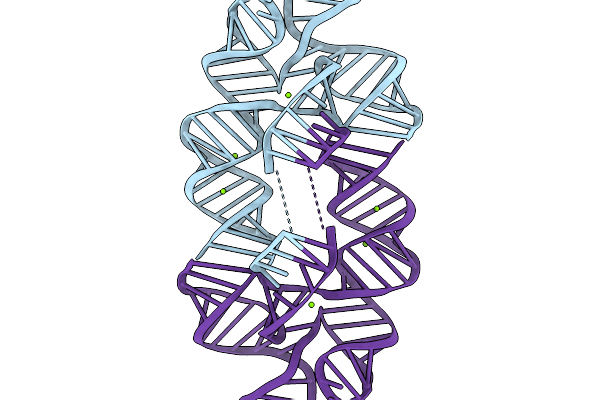 |
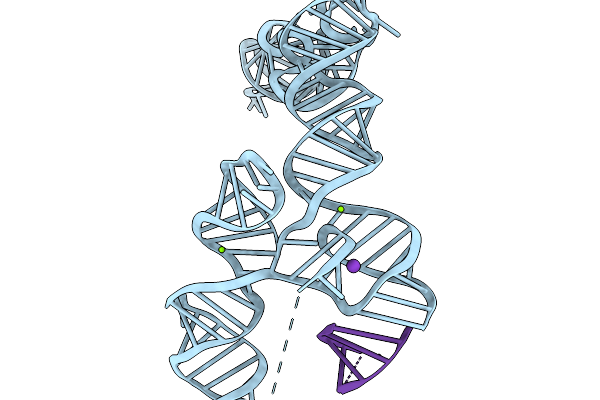
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.56 Å Release Date: 2026-03-04 Classification: RNA Ligands: MG |
 |
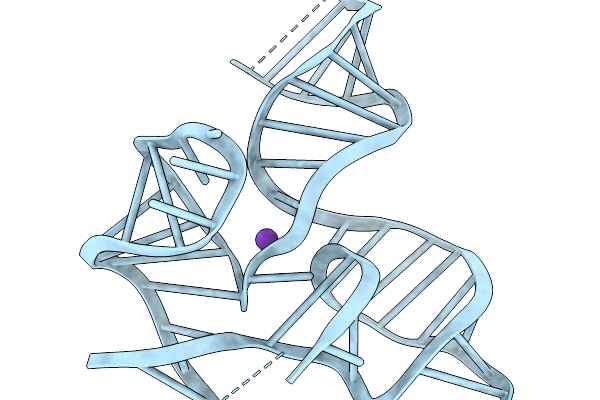
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.67 Å Release Date: 2026-03-04 Classification: RNA Ligands: MG, K |
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.87 Å Release Date: 2026-03-04 Classification: RNA Ligands: K |
 |
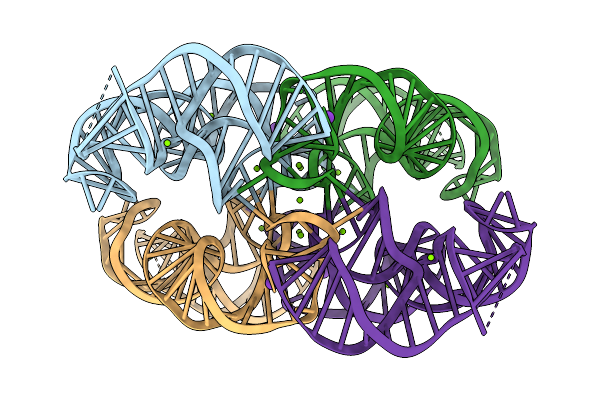
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.01 Å Release Date: 2026-03-04 Classification: RNA Ligands: EKJ |
 |
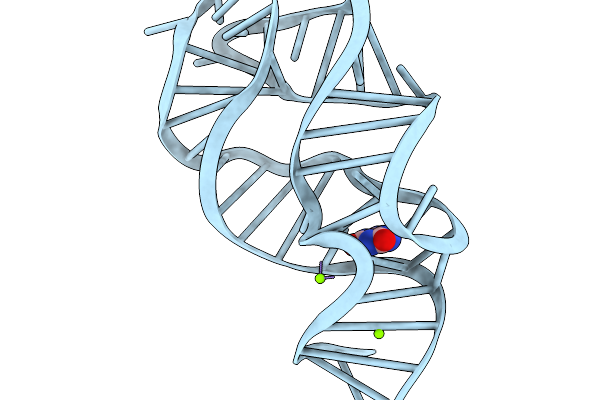
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: RNA Ligands: K, MG |
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.87 Å Release Date: 2026-03-04 Classification: RNA Ligands: MG, OXG |
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: RNA Ligands: MG |
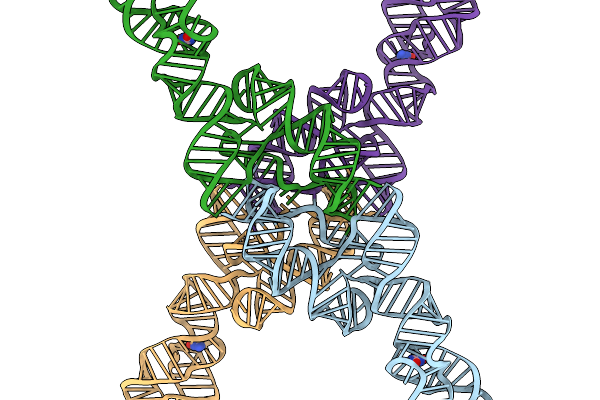 |
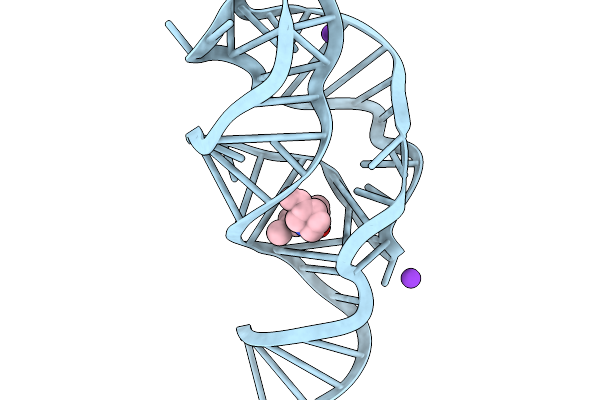
Rna Scaffold Attached To 8-Oxoguanine Riboswitch Aptamer, Combined Core Plus Aptamer
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.67 Å Release Date: 2026-03-04 Classification: RNA Ligands: OXG |
 |
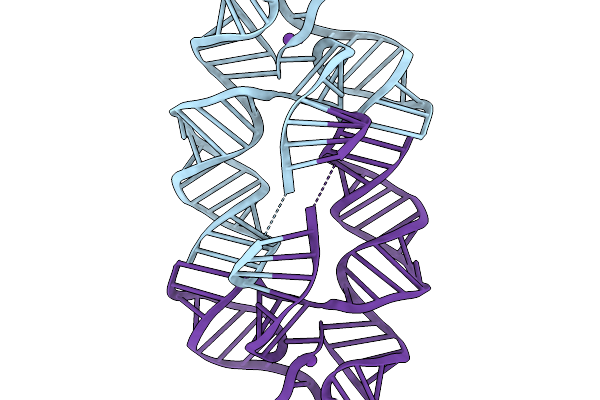
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: RNA Ligands: QI9, MG, K |
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: RNA Ligands: K |
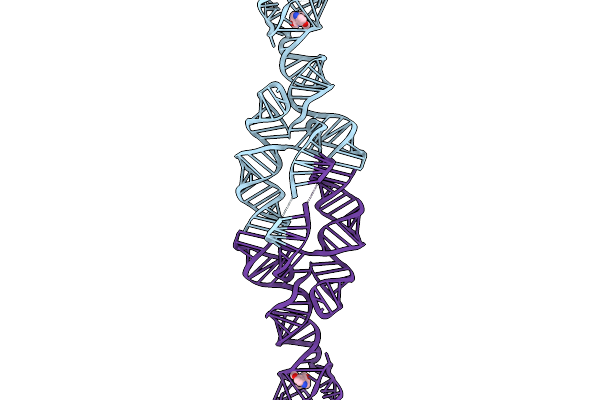 |
Scaffold Attached To Quinine-I Aptamer (Tonic) Refinement Of Aptamer And Core
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.94 Å Release Date: 2026-03-04 Classification: RNA Ligands: QI9 |
 |
Organism: Oceanobacillus iheyensis
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: RNA |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:3.51 Å Release Date: 2025-11-05 Classification: RNA Ligands: CS, MG |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:3.33 Å Release Date: 2025-07-09 Classification: RNA Ligands: MG |
 |
Organism: Schaalia odontolytica
Method: X-RAY DIFFRACTION Resolution:2.43 Å Release Date: 2025-06-11 Classification: RNA Ligands: MG, UG4 |
 |
Organism: Schaalia odontolytica
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2025-06-11 Classification: RNA Ligands: MG, A1AD3 |
 |
Organism: Rous sarcoma virus - prague c
Method: X-RAY DIFFRACTION Resolution:2.68 Å Release Date: 2025-04-09 Classification: RNA Ligands: IRI, MG, K |
 |
Organism: Rous sarcoma virus - prague c
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2025-04-09 Classification: RNA Ligands: IRI, K, TAM |
 |
Organism: Rous sarcoma virus - prague c
Method: ELECTRON MICROSCOPY Release Date: 2025-04-09 Classification: RNA Ligands: K, MG |

